Dataset Viewer
code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 09 Strain Gage
#
# This is one of the most commonly used sensor. It is used in many transducers. Its fundamental operating principle is fairly easy to understand and it will be the purpose of this lecture.
#
# A strain gage is essentially a thin wire that is wrapped on film of plastic.
# <img src="img/StrainGage.png" width="200">
# The strain gage is then mounted (glued) on the part for which the strain must be measured.
# <img src="img/Strain_gauge_2.jpg" width="200">
#
# ## Stress, Strain
# When a beam is under axial load, the axial stress, $\sigma_a$, is defined as:
# \begin{align*}
# \sigma_a = \frac{F}{A}
# \end{align*}
# with $F$ the axial load, and $A$ the cross sectional area of the beam under axial load.
#
# <img src="img/BeamUnderStrain.png" width="200">
#
# Under the load, the beam of length $L$ will extend by $dL$, giving rise to the definition of strain, $\epsilon_a$:
# \begin{align*}
# \epsilon_a = \frac{dL}{L}
# \end{align*}
# The beam will also contract laterally: the cross sectional area is reduced by $dA$. This results in a transverval strain $\epsilon_t$. The transversal and axial strains are related by the Poisson's ratio:
# \begin{align*}
# \nu = - \frac{\epsilon_t }{\epsilon_a}
# \end{align*}
# For a metal the Poission's ratio is typically $\nu = 0.3$, for an incompressible material, such as rubber (or water), $\nu = 0.5$.
#
# Within the elastic limit, the axial stress and axial strain are related through Hooke's law by the Young's modulus, $E$:
# \begin{align*}
# \sigma_a = E \epsilon_a
# \end{align*}
#
# <img src="img/ElasticRegime.png" width="200">
# ## Resistance of a wire
#
# The electrical resistance of a wire $R$ is related to its physical properties (the electrical resistiviy, $\rho$ in $\Omega$/m) and its geometry: length $L$ and cross sectional area $A$.
#
# \begin{align*}
# R = \frac{\rho L}{A}
# \end{align*}
#
# Mathematically, the change in wire dimension will result inchange in its electrical resistance. This can be derived from first principle:
# \begin{align}
# \frac{dR}{R} = \frac{d\rho}{\rho} + \frac{dL}{L} - \frac{dA}{A}
# \end{align}
# If the wire has a square cross section, then:
# \begin{align*}
# A & = L'^2 \\
# \frac{dA}{A} & = \frac{d(L'^2)}{L'^2} = \frac{2L'dL'}{L'^2} = 2 \frac{dL'}{L'}
# \end{align*}
# We have related the change in cross sectional area to the transversal strain.
# \begin{align*}
# \epsilon_t = \frac{dL'}{L'}
# \end{align*}
# Using the Poisson's ratio, we can relate then relate the change in cross-sectional area ($dA/A$) to axial strain $\epsilon_a = dL/L$.
# \begin{align*}
# \epsilon_t &= - \nu \epsilon_a \\
# \frac{dL'}{L'} &= - \nu \frac{dL}{L} \; \text{or}\\
# \frac{dA}{A} & = 2\frac{dL'}{L'} = -2 \nu \frac{dL}{L}
# \end{align*}
# Finally we can substitute express $dA/A$ in eq. for $dR/R$ and relate change in resistance to change of wire geometry, remembering that for a metal $\nu =0.3$:
# \begin{align}
# \frac{dR}{R} & = \frac{d\rho}{\rho} + \frac{dL}{L} - \frac{dA}{A} \\
# & = \frac{d\rho}{\rho} + \frac{dL}{L} - (-2\nu \frac{dL}{L}) \\
# & = \frac{d\rho}{\rho} + 1.6 \frac{dL}{L} = \frac{d\rho}{\rho} + 1.6 \epsilon_a
# \end{align}
# It also happens that for most metals, the resistivity increases with axial strain. In general, one can then related the change in resistance to axial strain by defining the strain gage factor:
# \begin{align}
# S = 1.6 + \frac{d\rho}{\rho}\cdot \frac{1}{\epsilon_a}
# \end{align}
# and finally, we have:
# \begin{align*}
# \frac{dR}{R} = S \epsilon_a
# \end{align*}
# $S$ is materials dependent and is typically equal to 2.0 for most commercially availabe strain gages. It is dimensionless.
#
# Strain gages are made of thin wire that is wraped in several loops, effectively increasing the length of the wire and therefore the sensitivity of the sensor.
#
# _Question:
#
# Explain why a longer wire is necessary to increase the sensitivity of the sensor_.
#
# Most commercially available strain gages have a nominal resistance (resistance under no load, $R_{ini}$) of 120 or 350 $\Omega$.
#
# Within the elastic regime, strain is typically within the range $10^{-6} - 10^{-3}$, in fact strain is expressed in unit of microstrain, with a 1 microstrain = $10^{-6}$. Therefore, changes in resistances will be of the same order. If one were to measure resistances, we will need a dynamic range of 120 dB, whih is typically very expensive. Instead, one uses the Wheatstone bridge to transform the change in resistance to a voltage, which is easier to measure and does not require such a large dynamic range.
# ## Wheatstone bridge:
# <img src="img/WheatstoneBridge.png" width="200">
#
# The output voltage is related to the difference in resistances in the bridge:
# \begin{align*}
# \frac{V_o}{V_s} = \frac{R_1R_3-R_2R_4}{(R_1+R_4)(R_2+R_3)}
# \end{align*}
#
# If the bridge is balanced, then $V_o = 0$, it implies: $R_1/R_2 = R_4/R_3$.
#
# In practice, finding a set of resistors that balances the bridge is challenging, and a potentiometer is used as one of the resistances to do minor adjustement to balance the bridge. If one did not do the adjustement (ie if we did not zero the bridge) then all the measurement will have an offset or bias that could be removed in a post-processing phase, as long as the bias stayed constant.
#
# If each resistance $R_i$ is made to vary slightly around its initial value, ie $R_i = R_{i,ini} + dR_i$. For simplicity, we will assume that the initial value of the four resistances are equal, ie $R_{1,ini} = R_{2,ini} = R_{3,ini} = R_{4,ini} = R_{ini}$. This implies that the bridge was initially balanced, then the output voltage would be:
#
# \begin{align*}
# \frac{V_o}{V_s} = \frac{1}{4} \left( \frac{dR_1}{R_{ini}} - \frac{dR_2}{R_{ini}} + \frac{dR_3}{R_{ini}} - \frac{dR_4}{R_{ini}} \right)
# \end{align*}
#
# Note here that the changes in $R_1$ and $R_3$ have a positive effect on $V_o$, while the changes in $R_2$ and $R_4$ have a negative effect on $V_o$. In practice, this means that is a beam is a in tension, then a strain gage mounted on the branch 1 or 3 of the Wheatstone bridge will produce a positive voltage, while a strain gage mounted on branch 2 or 4 will produce a negative voltage. One takes advantage of this to increase sensitivity to measure strain.
#
# ### Quarter bridge
# One uses only one quarter of the bridge, ie strain gages are only mounted on one branch of the bridge.
#
# \begin{align*}
# \frac{V_o}{V_s} = \pm \frac{1}{4} \epsilon_a S
# \end{align*}
# Sensitivity, $G$:
# \begin{align*}
# G = \frac{V_o}{\epsilon_a} = \pm \frac{1}{4}S V_s
# \end{align*}
#
#
# ### Half bridge
# One uses half of the bridge, ie strain gages are mounted on two branches of the bridge.
#
# \begin{align*}
# \frac{V_o}{V_s} = \pm \frac{1}{2} \epsilon_a S
# \end{align*}
#
# ### Full bridge
#
# One uses of the branches of the bridge, ie strain gages are mounted on each branch.
#
# \begin{align*}
# \frac{V_o}{V_s} = \pm \epsilon_a S
# \end{align*}
#
# Therefore, as we increase the order of bridge, the sensitivity of the instrument increases. However, one should be carefull how we mount the strain gages as to not cancel out their measurement.
# _Exercise_
#
# 1- Wheatstone bridge
#
# <img src="img/WheatstoneBridge.png" width="200">
#
# > How important is it to know \& match the resistances of the resistors you employ to create your bridge?
# > How would you do that practically?
# > Assume $R_1=120\,\Omega$, $R_2=120\,\Omega$, $R_3=120\,\Omega$, $R_4=110\,\Omega$, $V_s=5.00\,\text{V}$. What is $V_\circ$?
Vs = 5.00
Vo = (120**2-120*110)/(230*240) * Vs
print('Vo = ',Vo, ' V')
# typical range in strain a strain gauge can measure
# 1 -1000 micro-Strain
AxialStrain = 1000*10**(-6) # axial strain
StrainGageFactor = 2
R_ini = 120 # Ohm
R_1 = R_ini+R_ini*StrainGageFactor*AxialStrain
print(R_1)
Vo = (120**2-120*(R_1))/((120+R_1)*240) * Vs
print('Vo = ', Vo, ' V')
# > How important is it to know \& match the resistances of the resistors you employ to create your bridge?
# > How would you do that practically?
# > Assume $R_1= R_2 =R_3=120\,\Omega$, $R_4=120.01\,\Omega$, $V_s=5.00\,\text{V}$. What is $V_\circ$?
Vs = 5.00
Vo = (120**2-120*120.01)/(240.01*240) * Vs
print(Vo)
# 2- Strain gage 1:
#
# One measures the strain on a bridge steel beam. The modulus of elasticity is $E=190$ GPa. Only one strain gage is mounted on the bottom of the beam; the strain gage factor is $S=2.02$.
#
# > a) What kind of electronic circuit will you use? Draw a sketch of it.
#
# > b) Assume all your resistors including the unloaded strain gage are balanced and measure $120\,\Omega$, and that the strain gage is at location $R_2$. The supply voltage is $5.00\,\text{VDC}$. Will $V_\circ$ be positive or negative when a downward load is added?
# In practice, we cannot have all resistances = 120 $\Omega$. at zero load, the bridge will be unbalanced (show $V_o \neq 0$). How could we balance our bridge?
#
# Use a potentiometer to balance bridge, for the load cell, we ''zero'' the instrument.
#
# Other option to zero-out our instrument? Take data at zero-load, record the voltage, $V_{o,noload}$. Substract $V_{o,noload}$ to my data.
# > c) For a loading in which $V_\circ = -1.25\,\text{mV}$, calculate the strain $\epsilon_a$ in units of microstrain.
# \begin{align*}
# \frac{V_o}{V_s} & = - \frac{1}{4} \epsilon_a S\\
# \epsilon_a & = -\frac{4}{S} \frac{V_o}{V_s}
# \end{align*}
S = 2.02
Vo = -0.00125
Vs = 5
eps_a = -1*(4/S)*(Vo/Vs)
print(eps_a)
# > d) Calculate the axial stress (in MPa) in the beam under this load.
# > e) You now want more sensitivity in your measurement, you install a second strain gage on to
# p of the beam. Which resistor should you use for this second active strain gage?
#
# > f) With this new setup and the same applied load than previously, what should be the output voltage?
# 3- Strain Gage with Long Lead Wires
#
# <img src="img/StrainGageLongWires.png" width="360">
#
# A quarter bridge strain gage Wheatstone bridge circuit is constructed with $120\,\Omega$ resistors and a $120\,\Omega$ strain gage. For this practical application, the strain gage is located very far away form the DAQ station and the lead wires to the strain gage are $10\,\text{m}$ long and the lead wire have a resistance of $0.080\,\Omega/\text{m}$. The lead wire resistance can lead to problems since $R_{lead}$ changes with temperature.
#
# > Design a modified circuit that will cancel out the effect of the lead wires.
# ## Homework
#
| Lectures/09_StrainGage.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#export
from fastai.basics import *
from fastai.tabular.core import *
from fastai.tabular.model import *
from fastai.tabular.data import *
#hide
from nbdev.showdoc import *
# +
#default_exp tabular.learner
# -
# # Tabular learner
#
# > The function to immediately get a `Learner` ready to train for tabular data
# The main function you probably want to use in this module is `tabular_learner`. It will automatically create a `TabulaModel` suitable for your data and infer the irght loss function. See the [tabular tutorial](http://docs.fast.ai/tutorial.tabular) for an example of use in context.
# ## Main functions
#export
@log_args(but_as=Learner.__init__)
class TabularLearner(Learner):
"`Learner` for tabular data"
def predict(self, row):
tst_to = self.dls.valid_ds.new(pd.DataFrame(row).T)
tst_to.process()
tst_to.conts = tst_to.conts.astype(np.float32)
dl = self.dls.valid.new(tst_to)
inp,preds,_,dec_preds = self.get_preds(dl=dl, with_input=True, with_decoded=True)
i = getattr(self.dls, 'n_inp', -1)
b = (*tuplify(inp),*tuplify(dec_preds))
full_dec = self.dls.decode((*tuplify(inp),*tuplify(dec_preds)))
return full_dec,dec_preds[0],preds[0]
show_doc(TabularLearner, title_level=3)
# It works exactly as a normal `Learner`, the only difference is that it implements a `predict` method specific to work on a row of data.
#export
@log_args(to_return=True, but_as=Learner.__init__)
@delegates(Learner.__init__)
def tabular_learner(dls, layers=None, emb_szs=None, config=None, n_out=None, y_range=None, **kwargs):
"Get a `Learner` using `dls`, with `metrics`, including a `TabularModel` created using the remaining params."
if config is None: config = tabular_config()
if layers is None: layers = [200,100]
to = dls.train_ds
emb_szs = get_emb_sz(dls.train_ds, {} if emb_szs is None else emb_szs)
if n_out is None: n_out = get_c(dls)
assert n_out, "`n_out` is not defined, and could not be infered from data, set `dls.c` or pass `n_out`"
if y_range is None and 'y_range' in config: y_range = config.pop('y_range')
model = TabularModel(emb_szs, len(dls.cont_names), n_out, layers, y_range=y_range, **config)
return TabularLearner(dls, model, **kwargs)
# If your data was built with fastai, you probably won't need to pass anything to `emb_szs` unless you want to change the default of the library (produced by `get_emb_sz`), same for `n_out` which should be automatically inferred. `layers` will default to `[200,100]` and is passed to `TabularModel` along with the `config`.
#
# Use `tabular_config` to create a `config` and cusotmize the model used. There is just easy access to `y_range` because this argument is often used.
#
# All the other arguments are passed to `Learner`.
path = untar_data(URLs.ADULT_SAMPLE)
df = pd.read_csv(path/'adult.csv')
cat_names = ['workclass', 'education', 'marital-status', 'occupation', 'relationship', 'race']
cont_names = ['age', 'fnlwgt', 'education-num']
procs = [Categorify, FillMissing, Normalize]
dls = TabularDataLoaders.from_df(df, path, procs=procs, cat_names=cat_names, cont_names=cont_names,
y_names="salary", valid_idx=list(range(800,1000)), bs=64)
learn = tabular_learner(dls)
#hide
tst = learn.predict(df.iloc[0])
# +
#hide
#test y_range is passed
learn = tabular_learner(dls, y_range=(0,32))
assert isinstance(learn.model.layers[-1], SigmoidRange)
test_eq(learn.model.layers[-1].low, 0)
test_eq(learn.model.layers[-1].high, 32)
learn = tabular_learner(dls, config = tabular_config(y_range=(0,32)))
assert isinstance(learn.model.layers[-1], SigmoidRange)
test_eq(learn.model.layers[-1].low, 0)
test_eq(learn.model.layers[-1].high, 32)
# -
#export
@typedispatch
def show_results(x:Tabular, y:Tabular, samples, outs, ctxs=None, max_n=10, **kwargs):
df = x.all_cols[:max_n]
for n in x.y_names: df[n+'_pred'] = y[n][:max_n].values
display_df(df)
# ## Export -
#hide
from nbdev.export import notebook2script
notebook2script()
| nbs/43_tabular.learner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table class="ee-notebook-buttons" align="left">
# <td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Algorithms/landsat_radiance.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
# <td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Algorithms/landsat_radiance.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Algorithms/landsat_radiance.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
# </table>
# ## Install Earth Engine API and geemap
# Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://geemap.org). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
# The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
# +
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('Installing geemap ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# -
import ee
import geemap
# ## Create an interactive map
# The default basemap is `Google Maps`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/basemaps.py) can be added using the `Map.add_basemap()` function.
Map = geemap.Map(center=[40,-100], zoom=4)
Map
# ## Add Earth Engine Python script
# +
# Add Earth Engine dataset
# Load a raw Landsat scene and display it.
raw = ee.Image('LANDSAT/LC08/C01/T1/LC08_044034_20140318')
Map.centerObject(raw, 10)
Map.addLayer(raw, {'bands': ['B4', 'B3', 'B2'], 'min': 6000, 'max': 12000}, 'raw')
# Convert the raw data to radiance.
radiance = ee.Algorithms.Landsat.calibratedRadiance(raw)
Map.addLayer(radiance, {'bands': ['B4', 'B3', 'B2'], 'max': 90}, 'radiance')
# Convert the raw data to top-of-atmosphere reflectance.
toa = ee.Algorithms.Landsat.TOA(raw)
Map.addLayer(toa, {'bands': ['B4', 'B3', 'B2'], 'max': 0.2}, 'toa reflectance')
# -
# ## Display Earth Engine data layers
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
| Algorithms/landsat_radiance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Copyright 2020 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# -
# <img src="http://developer.download.nvidia.com/compute/machine-learning/frameworks/nvidia_logo.png" style="width: 90px; float: right;">
#
# # Object Detection with TRTorch (SSD)
# ---
# ## Overview
#
#
# In PyTorch 1.0, TorchScript was introduced as a method to separate your PyTorch model from Python, make it portable and optimizable.
#
# TRTorch is a compiler that uses TensorRT (NVIDIA's Deep Learning Optimization SDK and Runtime) to optimize TorchScript code. It compiles standard TorchScript modules into ones that internally run with TensorRT optimizations.
#
# TensorRT can take models from any major framework and specifically tune them to perform better on specific target hardware in the NVIDIA family, and TRTorch enables us to continue to remain in the PyTorch ecosystem whilst doing so. This allows us to leverage the great features in PyTorch, including module composability, its flexible tensor implementation, data loaders and more. TRTorch is available to use with both PyTorch and LibTorch.
#
# To get more background information on this, we suggest the **lenet-getting-started** notebook as a primer for getting started with TRTorch.
# ### Learning objectives
#
# This notebook demonstrates the steps for compiling a TorchScript module with TRTorch on a pretrained SSD network, and running it to test the speedup obtained.
#
# ## Contents
# 1. [Requirements](#1)
# 2. [SSD Overview](#2)
# 3. [Creating TorchScript modules](#3)
# 4. [Compiling with TRTorch](#4)
# 5. [Running Inference](#5)
# 6. [Measuring Speedup](#6)
# 7. [Conclusion](#7)
# ---
# <a id="1"></a>
# ## 1. Requirements
#
# Follow the steps in `notebooks/README` to prepare a Docker container, within which you can run this demo notebook.
#
# In addition to that, run the following cell to obtain additional libraries specific to this demo.
# Known working versions
# !pip install numpy==1.21.2 scipy==1.5.2 Pillow==6.2.0 scikit-image==0.17.2 matplotlib==3.3.0
# ---
# <a id="2"></a>
# ## 2. SSD
#
# ### Single Shot MultiBox Detector model for object detection
#
# _ | _
# - | -
# 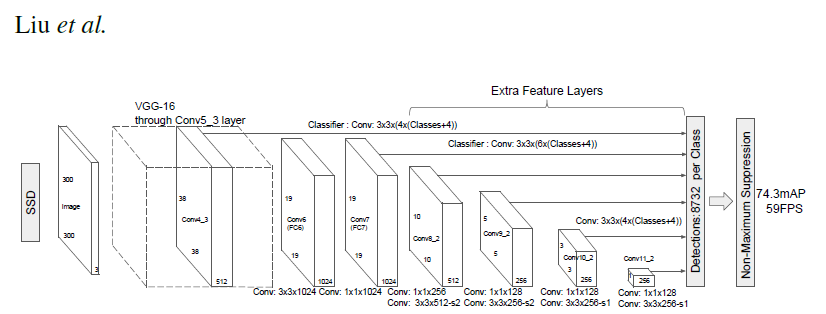 | 
# PyTorch has a model repository called the PyTorch Hub, which is a source for high quality implementations of common models. We can get our SSD model pretrained on [COCO](https://cocodataset.org/#home) from there.
#
# ### Model Description
#
# This SSD300 model is based on the
# [SSD: Single Shot MultiBox Detector](https://arxiv.org/abs/1512.02325) paper, which
# describes SSD as “a method for detecting objects in images using a single deep neural network".
# The input size is fixed to 300x300.
#
# The main difference between this model and the one described in the paper is in the backbone.
# Specifically, the VGG model is obsolete and is replaced by the ResNet-50 model.
#
# From the
# [Speed/accuracy trade-offs for modern convolutional object detectors](https://arxiv.org/abs/1611.10012)
# paper, the following enhancements were made to the backbone:
# * The conv5_x, avgpool, fc and softmax layers were removed from the original classification model.
# * All strides in conv4_x are set to 1x1.
#
# The backbone is followed by 5 additional convolutional layers.
# In addition to the convolutional layers, we attached 6 detection heads:
# * The first detection head is attached to the last conv4_x layer.
# * The other five detection heads are attached to the corresponding 5 additional layers.
#
# Detector heads are similar to the ones referenced in the paper, however,
# they are enhanced by additional BatchNorm layers after each convolution.
#
# More information about this SSD model is available at Nvidia's "DeepLearningExamples" Github [here](https://github.com/NVIDIA/DeepLearningExamples/tree/master/PyTorch/Detection/SSD).
import torch
torch.hub._validate_not_a_forked_repo=lambda a,b,c: True
# List of available models in PyTorch Hub from Nvidia/DeepLearningExamples
torch.hub.list('NVIDIA/DeepLearningExamples:torchhub')
# load SSD model pretrained on COCO from Torch Hub
precision = 'fp32'
ssd300 = torch.hub.load('NVIDIA/DeepLearningExamples:torchhub', 'nvidia_ssd', model_math=precision);
# Setting `precision="fp16"` will load a checkpoint trained with mixed precision
# into architecture enabling execution on Tensor Cores. Handling mixed precision data requires the Apex library.
# ### Sample Inference
# We can now run inference on the model. This is demonstrated below using sample images from the COCO 2017 Validation set.
# +
# Sample images from the COCO validation set
uris = [
'http://images.cocodataset.org/val2017/000000397133.jpg',
'http://images.cocodataset.org/val2017/000000037777.jpg',
'http://images.cocodataset.org/val2017/000000252219.jpg'
]
# For convenient and comprehensive formatting of input and output of the model, load a set of utility methods.
utils = torch.hub.load('NVIDIA/DeepLearningExamples:torchhub', 'nvidia_ssd_processing_utils')
# Format images to comply with the network input
inputs = [utils.prepare_input(uri) for uri in uris]
tensor = utils.prepare_tensor(inputs, False)
# The model was trained on COCO dataset, which we need to access in order to
# translate class IDs into object names.
classes_to_labels = utils.get_coco_object_dictionary()
# +
# Next, we run object detection
model = ssd300.eval().to("cuda")
detections_batch = model(tensor)
# By default, raw output from SSD network per input image contains 8732 boxes with
# localization and class probability distribution.
# Let’s filter this output to only get reasonable detections (confidence>40%) in a more comprehensive format.
results_per_input = utils.decode_results(detections_batch)
best_results_per_input = [utils.pick_best(results, 0.40) for results in results_per_input]
# -
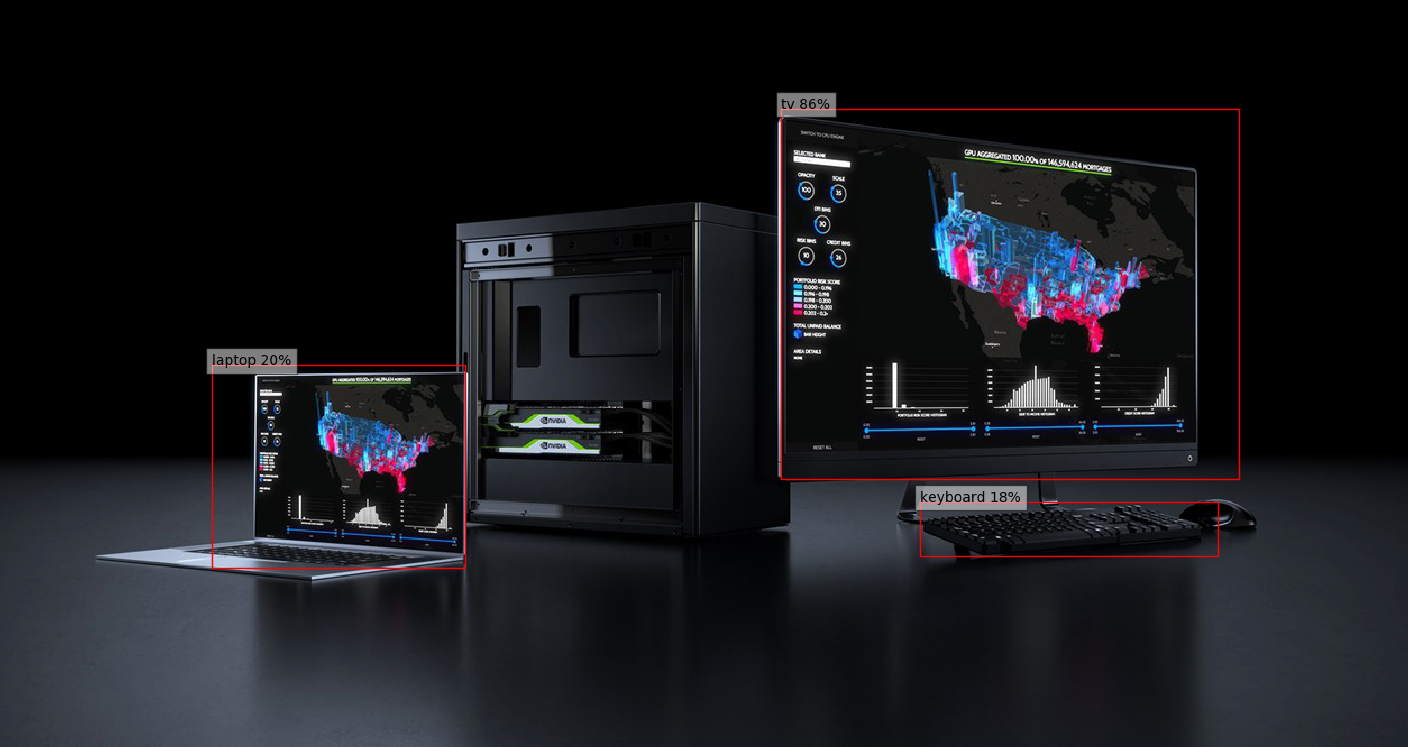
# ### Visualize results
# +
from matplotlib import pyplot as plt
import matplotlib.patches as patches
# The utility plots the images and predicted bounding boxes (with confidence scores).
def plot_results(best_results):
for image_idx in range(len(best_results)):
fig, ax = plt.subplots(1)
# Show original, denormalized image...
image = inputs[image_idx] / 2 + 0.5
ax.imshow(image)
# ...with detections
bboxes, classes, confidences = best_results[image_idx]
for idx in range(len(bboxes)):
left, bot, right, top = bboxes[idx]
x, y, w, h = [val * 300 for val in [left, bot, right - left, top - bot]]
rect = patches.Rectangle((x, y), w, h, linewidth=1, edgecolor='r', facecolor='none')
ax.add_patch(rect)
ax.text(x, y, "{} {:.0f}%".format(classes_to_labels[classes[idx] - 1], confidences[idx]*100), bbox=dict(facecolor='white', alpha=0.5))
plt.show()
# -
# Visualize results without TRTorch/TensorRT
plot_results(best_results_per_input)
# ### Benchmark utility
# +
import time
import numpy as np
import torch.backends.cudnn as cudnn
cudnn.benchmark = True
# Helper function to benchmark the model
def benchmark(model, input_shape=(1024, 1, 32, 32), dtype='fp32', nwarmup=50, nruns=1000):
input_data = torch.randn(input_shape)
input_data = input_data.to("cuda")
if dtype=='fp16':
input_data = input_data.half()
print("Warm up ...")
with torch.no_grad():
for _ in range(nwarmup):
features = model(input_data)
torch.cuda.synchronize()
print("Start timing ...")
timings = []
with torch.no_grad():
for i in range(1, nruns+1):
start_time = time.time()
pred_loc, pred_label = model(input_data)
torch.cuda.synchronize()
end_time = time.time()
timings.append(end_time - start_time)
if i%10==0:
print('Iteration %d/%d, avg batch time %.2f ms'%(i, nruns, np.mean(timings)*1000))
print("Input shape:", input_data.size())
print("Output location prediction size:", pred_loc.size())
print("Output label prediction size:", pred_label.size())
print('Average batch time: %.2f ms'%(np.mean(timings)*1000))
# -
# We check how well the model performs **before** we use TRTorch/TensorRT
# Model benchmark without TRTorch/TensorRT
model = ssd300.eval().to("cuda")
benchmark(model, input_shape=(128, 3, 300, 300), nruns=100)
# ---
# <a id="3"></a>
# ## 3. Creating TorchScript modules
# To compile with TRTorch, the model must first be in **TorchScript**. TorchScript is a programming language included in PyTorch which removes the Python dependency normal PyTorch models have. This conversion is done via a JIT compiler which given a PyTorch Module will generate an equivalent TorchScript Module. There are two paths that can be used to generate TorchScript: **Tracing** and **Scripting**. <br>
# - Tracing follows execution of PyTorch generating ops in TorchScript corresponding to what it sees. <br>
# - Scripting does an analysis of the Python code and generates TorchScript, this allows the resulting graph to include control flow which tracing cannot do.
#
# Tracing however due to its simplicity is more likely to compile successfully with TRTorch (though both systems are supported).
model = ssd300.eval().to("cuda")
traced_model = torch.jit.trace(model, [torch.randn((1,3,300,300)).to("cuda")])
# If required, we can also save this model and use it independently of Python.
# This is just an example, and not required for the purposes of this demo
torch.jit.save(traced_model, "ssd_300_traced.jit.pt")
# Obtain the average time taken by a batch of input with Torchscript compiled modules
benchmark(traced_model, input_shape=(128, 3, 300, 300), nruns=100)
# ---
# <a id="4"></a>
# ## 4. Compiling with TRTorch
# TorchScript modules behave just like normal PyTorch modules and are intercompatible. From TorchScript we can now compile a TensorRT based module. This module will still be implemented in TorchScript but all the computation will be done in TensorRT.
# +
import trtorch
# The compiled module will have precision as specified by "op_precision".
# Here, it will have FP16 precision.
trt_model = trtorch.compile(traced_model, {
"inputs": [trtorch.Input((3, 3, 300, 300))],
"enabled_precisions": {torch.float, torch.half}, # Run with FP16
"workspace_size": 1 << 20
})
# -
# ---
# <a id="5"></a>
# ## 5. Running Inference
# Next, we run object detection
# +
# using a TRTorch module is exactly the same as how we usually do inference in PyTorch i.e. model(inputs)
detections_batch = trt_model(tensor.to(torch.half)) # convert the input to half precision
# By default, raw output from SSD network per input image contains 8732 boxes with
# localization and class probability distribution.
# Let’s filter this output to only get reasonable detections (confidence>40%) in a more comprehensive format.
results_per_input = utils.decode_results(detections_batch)
best_results_per_input_trt = [utils.pick_best(results, 0.40) for results in results_per_input]
# -
# Now, let's visualize our predictions!
#
# Visualize results with TRTorch/TensorRT
plot_results(best_results_per_input_trt)
# We get similar results as before!
# ---
# ## 6. Measuring Speedup
# We can run the benchmark function again to see the speedup gained! Compare this result with the same batch-size of input in the case without TRTorch/TensorRT above.
# +
batch_size = 128
# Recompiling with batch_size we use for evaluating performance
trt_model = trtorch.compile(traced_model, {
"inputs": [trtorch.Input((batch_size, 3, 300, 300))],
"enabled_precisions": {torch.float, torch.half}, # Run with FP16
"workspace_size": 1 << 20
})
benchmark(trt_model, input_shape=(batch_size, 3, 300, 300), nruns=100, dtype="fp16")
# -
# ---
# ## 7. Conclusion
#
# In this notebook, we have walked through the complete process of compiling a TorchScript SSD300 model with TRTorch, and tested the performance impact of the optimization. We find that using the TRTorch compiled model, we gain significant speedup in inference without any noticeable drop in performance!
# ### Details
# For detailed information on model input and output,
# training recipies, inference and performance visit:
# [github](https://github.com/NVIDIA/DeepLearningExamples/tree/master/PyTorch/Detection/SSD)
# and/or [NGC](https://ngc.nvidia.com/catalog/model-scripts/nvidia:ssd_for_pytorch)
#
# ### References
#
# - [SSD: Single Shot MultiBox Detector](https://arxiv.org/abs/1512.02325) paper
# - [Speed/accuracy trade-offs for modern convolutional object detectors](https://arxiv.org/abs/1611.10012) paper
# - [SSD on NGC](https://ngc.nvidia.com/catalog/model-scripts/nvidia:ssd_for_pytorch)
# - [SSD on github](https://github.com/NVIDIA/DeepLearningExamples/tree/master/PyTorch/Detection/SSD)
| notebooks/ssd-object-detection-demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setting up
# +
# Dependencies
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import seaborn as sns
from scipy.stats import sem
plt.style.use('seaborn')
# Hide warning messages in notebook
# import warnings
# warnings.filterwarnings('ignore')
# -
# # Importing 4 csv files and merging them into one
# Import datasets
demo_2016 = pd.read_csv("assets/data/2016_demo_data.csv")
demo_2017 = pd.read_csv("assets/data/2017_demo_data.csv")
demo_2018 = pd.read_csv("assets/data/2018_demo_data.csv")
demo_2019 = pd.read_csv("assets/data/2019_demo_data.csv")
# Append datasets
final_df = demo_2016.append(demo_2017, ignore_index=True)
final_df = final_df.append(demo_2018, ignore_index=True)
final_df = final_df.append(demo_2019, ignore_index=True)
final_df
# +
# Export the dataframe (do this Only Once!)
# final_df.to_csv("assets/data/final_demo_data.csv", index=False)
# -
# # Importing the final csv file
final_demo = pd.read_csv("assets/data/final_demo_data.csv")
final_demo.head()
# # Checking the dataset
# Type of variables
final_demo.dtypes
# Any NaN in the dataset
final_demo.isnull().sum()
# Any uplicates (or similarities, mis-spellings) in ethnicity and city
ethnicity = final_demo["ethnicity"].unique()
city = final_demo["city"].unique()
# # Cleaning the dataset
# Change the type of "student_id" to string
final_demo["student_id"] = final_demo["student_id"].astype(str)
# Drop NaN in the dataset
final_demo.dropna(inplace=True)
# Replace ethnicity categories
final_demo.replace({"Asian Indian": "General Asian",
"Cambodian": "General Asian",
"Chinese": "General Asian",
"Filipino": "General Asian",
"Hmong": "General Asian",
"Japanese": "General Asian",
"Korean": "General Asian",
"Laotian": "General Asian",
"Other Asian": "General Asian",
"Vietnamese": "General Asian",
"Samoan": "Pacific Islander",
"Other Pacific Islander": "Pacific Islander",
"Guamanian": "Pacific Islander",
"Tahitian": "Pacific Islander",
"Laotian": "Pacific Islander",
"Hawaiian": "Pacific Islander"}, inplace=True)
# Replace city categories
final_demo.replace({"So San Francisco": "South SF",
"South San Francisco": "South SF",
"So. San Francisco": "South SF",
"So San Francisco ": "South SF",
"So San Francisco": "South SF",
"So Sn Francisco": "South SF",
"So SanFrancisco": "South SF",
"So San Francisco": "South SF",
"So San Francico": "South SF",
"S San Francisco": "South SF",
"So San Fran": "South SF",
"south San Francisco": "South SF",
"South San Francisco ": "South SF",
"South San Francico": "South SF",
"So San Francsico": "South SF",
"So San Franicsco": "South SF",
"Concord ": "Concord",
"Burlingame ": "Burlingame",
"Pacifica ": "Pacifica",
"Daly cITY": "Daly City",
"Daly City ": "Daly City",
"Daly City ": "Daly City",
"Daly Citiy": "Daly City",
"Daly Ciy": "Daly City",
"Daly CIty": "Daly City",
"San Mateo ": "San Mateo"
}, inplace=True)
# # Creating yearly enrollment group
# Year subgroups
enroll2016 = final_demo.loc[final_demo["year"]==2016]
enroll2017 = final_demo.loc[final_demo["year"]==2017]
enroll2018 = final_demo.loc[final_demo["year"]==2018]
enroll2019 = final_demo.loc[final_demo["year"]==2019]
# ## + Creating subgroups - Ethnicity
# +
### YEAR 2016 ###
# Calcaulte number of enrollment based on ethnicity
enrollRace2016 = pd.DataFrame(enroll2016.groupby(["ethnicity"])["student_id"].count())
# Add year column
enrollRace2016["year"] = 2016
# Rename column name
enrollRace2016.rename({"student_id": "enrollment"}, axis=1, inplace=True)
# +
### YEAR 2017 ###
# Calcaulte number of enrollment based on ethnicity
enrollRace2017 = pd.DataFrame(enroll2017.groupby(["ethnicity"])["student_id"].count())
# Add year column
enrollRace2017["year"] = 2017
# Rename column name
enrollRace2017.rename({"student_id": "enrollment"}, axis=1, inplace=True)
# +
### YEAR 2018 ###
# Calcaulte number of enrollment based on ethnicity
enrollRace2018 = pd.DataFrame(enroll2018.groupby(["ethnicity"])["student_id"].count())
# Add year column
enrollRace2018["year"] = 2018
# Rename column name
enrollRace2018.rename({"student_id": "enrollment"}, axis=1, inplace=True)
# +
### YEAR 2019 ###
# Calcaulte number of enrollment based on ethnicity
enrollRace2019 = pd.DataFrame(enroll2019.groupby(["ethnicity"])["student_id"].count())
# Add year column
enrollRace2019["year"] = 2019
# Rename column name
enrollRace2019.rename({"student_id": "enrollment"}, axis=1, inplace=True)
# -
# Append 4 dataframes into one
enrollRace = enrollRace2016.append(enrollRace2017)
enrollRace = enrollRace.append(enrollRace2018)
enrollRace = enrollRace.append(enrollRace2019)
# Export to csv file
enrollRace.to_csv("assets/data/race_data.csv", index=True)
# ## + Creating subgroups - City
# +
### YEAR 2016 ###
# Calcaulte number of enrollment based on city
enrollCity2016 = pd.DataFrame(enroll2016.groupby(["city"])["student_id"].count())
# Add year column
enrollCity2016["year"] = 2016
# Rename column name
enrollCity2016.rename({"student_id": "enrollment"}, axis=1, inplace=True)
# -
enrollCity2016
| jupyter/ethnicity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Disambiguation
# +
import pprint
import subprocess
import sys
sys.path.append('../')
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
import matplotlib
import matplotlib.gridspec as gridspec
from mpl_toolkits.axes_grid1 import make_axes_locatable
import seaborn as sns
# %matplotlib inline
plt.rcParams['figure.figsize'] = (12.9, 12)
np.set_printoptions(suppress=True, precision=5)
sns.set(font_scale=3.5)
from network import Protocol, NetworkManager, BCPNNPerfect, TimedInput
from connectivity_functions import create_orthogonal_canonical_representation, build_network_representation
from connectivity_functions import get_weights_from_probabilities, get_probabilities_from_network_representation
from analysis_functions import calculate_recall_time_quantities, get_weights
from analysis_functions import get_weights_collections
from plotting_functions import plot_network_activity_angle, plot_weight_matrix
from analysis_functions import calculate_angle_from_history, calculate_winning_pattern_from_distances
from analysis_functions import calculate_patterns_timings
# -
epsilon = 10e-20
# +
def produce_overlaped_sequences(minicolumns, hypercolumns, n_patterns, s, r, mixed_start=False, contiguous=True):
n_r = int(r * n_patterns/2)
n_s = int(s * hypercolumns)
n_size = int(n_patterns / 2)
matrix = create_orthogonal_canonical_representation(minicolumns, hypercolumns)[:n_patterns]
sequence1 = matrix[:n_size]
sequence2 = matrix[n_size:]
if mixed_start:
start_index = 0
end_index = n_r
else:
start_index = max(int(0.5 * (n_size - n_r)), 0)
end_index = min(start_index + n_r, n_size)
for index in range(start_index, end_index):
if contiguous:
sequence2[index, :n_s] = sequence1[index, :n_s]
else:
sequence2[index, ...] = sequence1[index, ...]
sequence2[index, n_s:] = n_patterns + index
if False:
print(n_r)
print(n_size)
print(start_index)
print(end_index)
return sequence1, sequence2
def create_weights_from_two_sequences(nn, dt, n_patterns, s, r, mixed_start, contiguous,
training_time, inter_pulse_interval, inter_sequence_interval,
epochs, resting_time):
filtered = True
minicolumns = nn.minicolumns
hypercolumns = nn.hypercolumns
tau_z_pre_ampa = nn.tau_z_pre_ampa
tau_z_post_ampa = nn.tau_z_post_ampa
seq1, seq2 = produce_overlaped_sequences(minicolumns, hypercolumns, n_patterns, s, r,
mixed_start=mixed_start, contiguous=contiguous)
nr1 = build_network_representation(seq1, minicolumns, hypercolumns)
nr2 = build_network_representation(seq2, minicolumns, hypercolumns)
# Get the first
timed_input = TimedInput(nr1, dt, training_time, inter_pulse_interval=inter_pulse_interval,
inter_sequence_interval=inter_pulse_interval, epochs=epochs,
resting_time=resting_time)
S = timed_input.build_timed_input()
z_pre = timed_input.build_filtered_input_pre(tau_z_pre_ampa)
z_post = timed_input.build_filtered_input_post(tau_z_post_ampa)
pi1, pj1, P1 = timed_input.calculate_probabilities_from_time_signal(filtered=filtered)
w_timed1 = get_weights_from_probabilities(pi1, pj1, P1, minicolumns, hypercolumns)
t1 = timed_input.T_total
# Get the second
timed_input = TimedInput(nr2, dt, training_time, inter_pulse_interval=inter_pulse_interval,
inter_sequence_interval=inter_pulse_interval, epochs=epochs,
resting_time=resting_time)
S = timed_input.build_timed_input()
z_pre = timed_input.build_filtered_input_pre(tau_z_pre_ampa)
z_post = timed_input.build_filtered_input_post(tau_z_post_ampa)
t2 = timed_input.T_total
pi2, pj2, P2 = timed_input.calculate_probabilities_from_time_signal(filtered=filtered)
w_timed2 = get_weights_from_probabilities(pi2, pj2, P2, minicolumns, hypercolumns)
t_total = t1 + t2
# Mix
pi_total = (t1 / t_total) * pi1 + ((t_total - t1)/ t_total) * pi2
pj_total = (t1 / t_total) * pj1 + ((t_total - t1)/ t_total) * pj2
P_total = (t1 / t_total) * P1 + ((t_total - t1)/ t_total) * P2
w_total, beta = get_weights_from_probabilities(pi_total, pj_total, P_total, minicolumns, hypercolumns)
return seq1, seq2, nr1, nr2, w_total, beta
def calculate_recall_success_nr(manager, nr, T_recall, T_cue, debug=False, remove=0.020):
n_seq = nr.shape[0]
I_cue = nr[0]
# Do the recall
manager.run_network_recall(T_recall=T_recall, I_cue=I_cue, T_cue=T_cue,
reset=True, empty_history=True)
distances = calculate_angle_from_history(manager)
winning = calculate_winning_pattern_from_distances(distances)
timings = calculate_patterns_timings(winning, manager.dt, remove=remove)
pattern_sequence = [x[0] for x in timings]
# Calculate whether it was succesfull
success = 1.0
for index, pattern_index in enumerate(pattern_sequence[:n_seq]):
pattern = manager.patterns_dic[pattern_index]
goal_pattern = nr[index]
if not np.array_equal(pattern, goal_pattern):
success = 0.0
break
if debug:
return success, timings, pattern_sequence
else:
return success
# -
# ## An example
# +
always_learning = False
strict_maximum = True
perfect = False
z_transfer = False
k_perfect = True
diagonal_zero = False
normalized_currents = True
g_w_ampa = 2.0
g_w = 0.0
g_a = 10.0
tau_a = 0.250
G = 1.0
sigma = 0.0
tau_m = 0.020
tau_z_pre_ampa = 0.025
tau_z_post_ampa = 0.025
tau_p = 10.0
hypercolumns = 1
minicolumns = 20
n_patterns = 20
# Manager properties
dt = 0.001
values_to_save = ['o', 'i_ampa', 'a']
# Protocol
training_time = 0.100
inter_sequence_interval = 0.0
inter_pulse_interval = 0.0
resting_time = 2.0
epochs = 1
# Recall
T_recall = 1.0
T_cue = 0.020
# Patterns parameters
nn = BCPNNPerfect(hypercolumns, minicolumns, g_w_ampa=g_w_ampa, g_w=g_w, g_a=g_a, tau_a=tau_a, tau_m=tau_m,
sigma=sigma, G=G, tau_z_pre_ampa=tau_z_pre_ampa, tau_z_post_ampa=tau_z_post_ampa, tau_p=tau_p,
z_transfer=z_transfer, diagonal_zero=diagonal_zero, strict_maximum=strict_maximum,
perfect=perfect, k_perfect=k_perfect, always_learning=always_learning,
normalized_currents=normalized_currents)
# Build the manager
manager = NetworkManager(nn=nn, dt=dt, values_to_save=values_to_save)
# Build the protocol for training
mixed_start = False
contiguous = True
s = 1.0
r = 0.3
matrix = create_orthogonal_canonical_representation(minicolumns, hypercolumns)
aux = create_weights_from_two_sequences(nn, dt, n_patterns, s, r, mixed_start, contiguous,
training_time, inter_pulse_interval=inter_pulse_interval,
inter_sequence_interval=inter_sequence_interval, epochs=epochs,
resting_time=resting_time)
seq1, seq2, nr1, nr2, w_total, beta = aux
nr = np.concatenate((nr1, nr2))
aux, indexes = np.unique(nr, axis=0, return_index=True)
patterns_dic = {index:pattern for (index, pattern) in zip(indexes, aux)}
nn.w_ampa = w_total
manager.patterns_dic = patterns_dic
s = calculate_recall_success_nr(manager, nr1, T_recall, T_cue)
print('s1=', s)
plot_network_activity_angle(manager)
s = calculate_recall_success_nr(manager, nr2, T_recall, T_cue)
print('s2=', s)
plot_network_activity_angle(manager)
# -
plot_weight_matrix(nn, ampa=True)
# ## More systematic
# +
# %%time
always_learning = False
strict_maximum = True
perfect = False
z_transfer = False
k_perfect = True
diagonal_zero = False
normalized_currents = True
g_w_ampa = 2.0
g_w = 0.0
g_a = 10.0
tau_a = 0.250
g_beta = 1.0
G = 1.0
sigma = 0.0
tau_m = 0.010
tau_z_pre_ampa = 0.050
tau_z_post_ampa = 0.005
tau_p = 10.0
hypercolumns = 1
minicolumns = 20
n_patterns = 20
# Manager properties
dt = 0.001
values_to_save = ['o', 'i_ampa', 'a']
# Protocol
training_time = 0.100
inter_sequence_interval = 0.0
inter_pulse_interval = 0.0
epochs = 1
mixed_start = False
contiguous = True
s = 1.0
r = 0.25
# Recall
T_recall = 1.0
T_cue = 0.020
num = 10
r_space = np.linspace(0, 0.9, num=num)
success_vector = np.zeros(num)
factor = 0.2
g_w_ampa * (w_total[0, 0] - w_total[2, 0])
for r_index, r in enumerate(r_space):
print('r_index', r_index)
# The network
nn = BCPNNPerfect(hypercolumns, minicolumns, g_w_ampa=g_w_ampa, g_w=g_w, g_a=g_a, tau_a=tau_a, tau_m=tau_m,
sigma=sigma, G=G, tau_z_pre_ampa=tau_z_pre_ampa, tau_z_post_ampa=tau_z_post_ampa, tau_p=tau_p,
z_transfer=z_transfer, diagonal_zero=diagonal_zero, strict_maximum=strict_maximum,
perfect=perfect, k_perfect=k_perfect, always_learning=always_learning,
normalized_currents=normalized_currents, g_beta=g_beta)
# Build the manager
manager = NetworkManager(nn=nn, dt=dt, values_to_save=values_to_save)
# The sequences
matrix = create_orthogonal_canonical_representation(minicolumns, hypercolumns)
aux = create_weights_from_two_sequences(nn, dt, n_patterns, s, r, mixed_start, contiguous,
training_time, inter_pulse_interval=inter_pulse_interval,
inter_sequence_interval=inter_sequence_interval, epochs=epochs,
resting_time=resting_time)
seq1, seq2, nr1, nr2, w_total, beta = aux
nr = np.concatenate((nr1, nr2))
aux, indexes = np.unique(nr, axis=0, return_index=True)
patterns_dic = {index:pattern for (index, pattern) in zip(indexes, aux)}
nn.w_ampa = w_total
nn.beta = beta
manager.patterns_dic = patterns_dic
current = g_w_ampa * (w_total[0, 0] - w_total[2, 0])
noise = factor * current
nn.sigma = noise
# Recall
aux = calculate_recall_success_nr(manager, nr1, T_recall, T_cue, debug=True, remove=0.020)
s1, timings, pattern_sequence = aux
print('1', s1, pattern_sequence, seq1)
aux = calculate_recall_success_nr(manager, nr2, T_recall, T_cue, debug=True, remove=0.020)
s2, timings, pattern_sequence = aux
print('2', s2, pattern_sequence, seq2)
success_vector[r_index] = 0.5 * (s1 + s2)
# +
markersize = 15
linewdith = 8
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
ax.plot(r_space, success_vector, 'o-', lw=linewdith, ms=markersize)
ax.axhline(0, ls='--', color='gray')
ax.axvline(0, ls='--', color='gray')
ax.set_xlabel('Overlap')
ax.set_ylabel('Recall')
# -
# #### tau_z
# +
# %%time
always_learning = False
strict_maximum = True
perfect = False
z_transfer = False
k_perfect = True
diagonal_zero = False
normalized_currents = True
g_w_ampa = 2.0
g_w = 0.0
g_a = 10.0
tau_a = 0.250
G = 1.0
sigma = 0.0
tau_m = 0.010
tau_z_pre_ampa = 0.025
tau_z_post_ampa = 0.025
tau_p = 10.0
hypercolumns = 1
minicolumns = 20
n_patterns = 20
# Manager properties
dt = 0.001
values_to_save = ['o']
# Protocol
training_time = 0.100
inter_sequence_interval = 0.0
inter_pulse_interval = 0.0
epochs = 1
mixed_start = False
contiguous = True
s = 1.0
r = 0.25
# Recall
T_recall = 1.0
T_cue = 0.020
num = 10
r_space = np.linspace(0, 0.9, num=num)
success_vector = np.zeros(num)
tau_z_list = [0.025, 0.035, 0.050, 0.075]
#tau_z_list = [0.025, 0.100, 0.250]
#tau_z_list = [0.025, 0.050]
success_list = []
for tau_z_pre_ampa in tau_z_list:
success_vector = np.zeros(num)
print(tau_z_pre_ampa)
for r_index, r in enumerate(r_space):
# The network
nn = BCPNNPerfect(hypercolumns, minicolumns, g_w_ampa=g_w_ampa, g_w=g_w, g_a=g_a, tau_a=tau_a, tau_m=tau_m,
sigma=sigma, G=G, tau_z_pre_ampa=tau_z_pre_ampa, tau_z_post_ampa=tau_z_post_ampa, tau_p=tau_p,
z_transfer=z_transfer, diagonal_zero=diagonal_zero, strict_maximum=strict_maximum,
perfect=perfect, k_perfect=k_perfect, always_learning=always_learning,
normalized_currents=normalized_currents)
# Build the manager
manager = NetworkManager(nn=nn, dt=dt, values_to_save=values_to_save)
# The sequences
matrix = create_orthogonal_canonical_representation(minicolumns, hypercolumns)
aux = create_weights_from_two_sequences(nn, dt, n_patterns, s, r, mixed_start, contiguous,
training_time, inter_pulse_interval=inter_pulse_interval,
inter_sequence_interval=inter_sequence_interval, epochs=epochs,
resting_time=resting_time)
seq1, seq2, nr1, nr2, w_total, beta = aux
nr = np.concatenate((nr1, nr2))
aux, indexes = np.unique(nr, axis=0, return_index=True)
patterns_dic = {index:pattern for (index, pattern) in zip(indexes, aux)}
nn.w_ampa = w_total
manager.patterns_dic = patterns_dic
# Recall
s1 = calculate_recall_success_nr(manager, nr1, T_recall, T_cue)
s2 = calculate_recall_success_nr(manager, nr2, T_recall, T_cue)
success_vector[r_index] = 0.5 * (s1 + s2)
success_list.append(np.copy(success_vector))
# +
markersize = 15
linewdith = 8
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
for tau_z, success_vector in zip(tau_z_list, success_list):
ax.plot(r_space, success_vector, 'o-', lw=linewdith, ms=markersize, label=str(tau_z))
ax.axhline(0, ls='--', color='gray')
ax.axvline(0, ls='--', color='gray')
ax.set_xlabel('Overlap')
ax.set_ylabel('Recall')
ax.legend();
# -
# #### Scale
# +
# %%time
always_learning = False
strict_maximum = True
perfect = False
z_transfer = False
k_perfect = True
diagonal_zero = False
normalized_currents = True
g_w_ampa = 2.0
g_w = 0.0
g_a = 10.0
tau_a = 0.250
G = 1.0
sigma = 0.0
tau_m = 0.010
tau_z_pre_ampa = 0.025
tau_z_post_ampa = 0.025
tau_p = 10.0
hypercolumns = 1
minicolumns = 20
n_patterns = 20
# Manager properties
dt = 0.001
values_to_save = ['o']
# Protocol
training_time = 0.100
inter_sequence_interval = 0.0
inter_pulse_interval = 0.0
epochs = 1
mixed_start = False
contiguous = True
s = 1.0
r = 0.25
# Recall
T_recall = 1.0
T_cue = 0.020
num = 10
r_space = np.linspace(0, 0.9, num=num)
success_vector = np.zeros(num)
hypercolumns_list = [1, 3, 7, 10]
#tau_z_list = [0.025, 0.100, 0.250]
#tau_z_list = [0.025, 0.050]
success_list = []
for hypercolumns in hypercolumns_list:
success_vector = np.zeros(num)
print(hypercolumns)
for r_index, r in enumerate(r_space):
# The network
nn = BCPNNPerfect(hypercolumns, minicolumns, g_w_ampa=g_w_ampa, g_w=g_w, g_a=g_a, tau_a=tau_a, tau_m=tau_m,
sigma=sigma, G=G, tau_z_pre_ampa=tau_z_pre_ampa, tau_z_post_ampa=tau_z_post_ampa, tau_p=tau_p,
z_transfer=z_transfer, diagonal_zero=diagonal_zero, strict_maximum=strict_maximum,
perfect=perfect, k_perfect=k_perfect, always_learning=always_learning,
normalized_currents=normalized_currents)
# Build the manager
manager = NetworkManager(nn=nn, dt=dt, values_to_save=values_to_save)
# The sequences
matrix = create_orthogonal_canonical_representation(minicolumns, hypercolumns)
aux = create_weights_from_two_sequences(nn, dt, n_patterns, s, r, mixed_start, contiguous,
training_time, inter_pulse_interval=inter_pulse_interval,
inter_sequence_interval=inter_sequence_interval, epochs=epochs,
resting_time=resting_time)
seq1, seq2, nr1, nr2, w_total, beta = aux
nr = np.concatenate((nr1, nr2))
aux, indexes = np.unique(nr, axis=0, return_index=True)
patterns_dic = {index:pattern for (index, pattern) in zip(indexes, aux)}
nn.w_ampa = w_total
manager.patterns_dic = patterns_dic
# Recall
s1 = calculate_recall_success_nr(manager, nr1, T_recall, T_cue)
s2 = calculate_recall_success_nr(manager, nr2, T_recall, T_cue)
success_vector[r_index] = 0.5 * (s1 + s2)
success_list.append(np.copy(success_vector))
# +
markersize = 15
linewdith = 8
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
for hypercolumns, success_vector in zip(hypercolumns_list, success_list):
ax.plot(r_space, success_vector, 'o-', lw=linewdith, ms=markersize, label=str(hypercolumns))
ax.axhline(0, ls='--', color='gray')
ax.axvline(0, ls='--', color='gray')
ax.set_xlabel('Overlap')
ax.set_ylabel('Recall')
ax.legend();
# -
# #### tau_m
# +
# %%time
always_learning = False
strict_maximum = True
perfect = False
z_transfer = False
k_perfect = True
diagonal_zero = False
normalized_currents = True
g_w_ampa = 2.0
g_w = 0.0
g_a = 10.0
tau_a = 0.250
G = 1.0
sigma = 0.0
tau_m = 0.010
tau_z_pre_ampa = 0.025
tau_z_post_ampa = 0.025
tau_p = 10.0
hypercolumns = 1
minicolumns = 20
n_patterns = 20
# Manager properties
dt = 0.001
values_to_save = ['o']
# Protocol
training_time = 0.100
inter_sequence_interval = 0.0
inter_pulse_interval = 0.0
epochs = 1
mixed_start = False
contiguous = True
s = 1.0
r = 0.25
# Recall
T_recall = 1.0
T_cue = 0.020
num = 10
r_space = np.linspace(0, 0.9, num=num)
success_vector = np.zeros(num)
tau_m_list = [0.001, 0.008, 0.020]
success_list = []
for tau_m in tau_m_list:
success_vector = np.zeros(num)
print(tau_m)
for r_index, r in enumerate(r_space):
# The network
nn = BCPNNPerfect(hypercolumns, minicolumns, g_w_ampa=g_w_ampa, g_w=g_w, g_a=g_a, tau_a=tau_a, tau_m=tau_m,
sigma=sigma, G=G, tau_z_pre_ampa=tau_z_pre_ampa, tau_z_post_ampa=tau_z_post_ampa, tau_p=tau_p,
z_transfer=z_transfer, diagonal_zero=diagonal_zero, strict_maximum=strict_maximum,
perfect=perfect, k_perfect=k_perfect, always_learning=always_learning,
normalized_currents=normalized_currents)
# Build the manager
manager = NetworkManager(nn=nn, dt=dt, values_to_save=values_to_save)
# The sequences
matrix = create_orthogonal_canonical_representation(minicolumns, hypercolumns)
aux = create_weights_from_two_sequences(nn, dt, n_patterns, s, r, mixed_start, contiguous,
training_time, inter_pulse_interval=inter_pulse_interval,
inter_sequence_interval=inter_sequence_interval, epochs=epochs,
resting_time=resting_time)
seq1, seq2, nr1, nr2, w_total, beta = aux
nr = np.concatenate((nr1, nr2))
aux, indexes = np.unique(nr, axis=0, return_index=True)
patterns_dic = {index:pattern for (index, pattern) in zip(indexes, aux)}
nn.w_ampa = w_total
manager.patterns_dic = patterns_dic
# Recall
s1 = calculate_recall_success_nr(manager, nr1, T_recall, T_cue)
s2 = calculate_recall_success_nr(manager, nr2, T_recall, T_cue)
success_vector[r_index] = 0.5 * (s1 + s2)
success_list.append(np.copy(success_vector))
# +
markersize = 15
linewdith = 8
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
for tau_m, success_vector in zip(tau_m_list, success_list):
ax.plot(r_space, success_vector, 'o-', lw=linewdith, ms=markersize, label=str(tau_m))
ax.axhline(0, ls='--', color='gray')
ax.axvline(0, ls='--', color='gray')
ax.set_xlabel('Overlap')
ax.set_ylabel('Recall')
ax.legend();
# -
# #### training time
# +
# %%time
always_learning = False
strict_maximum = True
perfect = False
z_transfer = False
k_perfect = True
diagonal_zero = False
normalized_currents = True
g_w_ampa = 2.0
g_w = 0.0
g_a = 10.0
tau_a = 0.250
G = 1.0
sigma = 0.0
tau_m = 0.010
tau_z_pre_ampa = 0.025
tau_z_post_ampa = 0.025
tau_p = 10.0
hypercolumns = 1
minicolumns = 20
n_patterns = 20
# Manager properties
dt = 0.001
values_to_save = ['o']
# Protocol
training_time = 0.100
inter_sequence_interval = 0.0
inter_pulse_interval = 0.0
epochs = 1
mixed_start = False
contiguous = True
s = 1.0
r = 0.25
# Recall
T_recall = 1.0
T_cue = 0.020
num = 10
r_space = np.linspace(0, 0.9, num=num)
success_vector = np.zeros(num)
training_time_list = [0.050, 0.100, 0.250, 0.500]
success_list = []
for training_time in training_time_list:
success_vector = np.zeros(num)
print(training_time)
for r_index, r in enumerate(r_space):
# The network
nn = BCPNNPerfect(hypercolumns, minicolumns, g_w_ampa=g_w_ampa, g_w=g_w, g_a=g_a, tau_a=tau_a, tau_m=tau_m,
sigma=sigma, G=G, tau_z_pre_ampa=tau_z_pre_ampa, tau_z_post_ampa=tau_z_post_ampa, tau_p=tau_p,
z_transfer=z_transfer, diagonal_zero=diagonal_zero, strict_maximum=strict_maximum,
perfect=perfect, k_perfect=k_perfect, always_learning=always_learning,
normalized_currents=normalized_currents)
# Build the manager
manager = NetworkManager(nn=nn, dt=dt, values_to_save=values_to_save)
# The sequences
matrix = create_orthogonal_canonical_representation(minicolumns, hypercolumns)
aux = create_weights_from_two_sequences(nn, dt, n_patterns, s, r, mixed_start, contiguous,
training_time, inter_pulse_interval=inter_pulse_interval,
inter_sequence_interval=inter_sequence_interval, epochs=epochs,
resting_time=resting_time)
seq1, seq2, nr1, nr2, w_total, beta = aux
nr = np.concatenate((nr1, nr2))
aux, indexes = np.unique(nr, axis=0, return_index=True)
patterns_dic = {index:pattern for (index, pattern) in zip(indexes, aux)}
nn.w_ampa = w_total
manager.patterns_dic = patterns_dic
# Recall
s1 = calculate_recall_success_nr(manager, nr1, T_recall, T_cue)
s2 = calculate_recall_success_nr(manager, nr2, T_recall, T_cue)
success_vector[r_index] = 0.5 * (s1 + s2)
success_list.append(np.copy(success_vector))
# +
markersize = 15
linewdith = 8
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
for training_time, success_vector in zip(training_time_list, success_list):
ax.plot(r_space, success_vector, 'o-', lw=linewdith, ms=markersize, label=str(training_time))
ax.axhline(0, ls='--', color='gray')
ax.axvline(0, ls='--', color='gray')
ax.set_xlabel('Overlap')
ax.set_ylabel('Recall')
ax.legend();
# -
# ## Systematic with noise
# +
# %%time
always_learning = False
strict_maximum = True
perfect = False
z_transfer = False
k_perfect = True
diagonal_zero = False
normalized_currents = True
g_w_ampa = 2.0
g_w = 0.0
g_a = 10.0
tau_a = 0.250
g_beta = 0.0
G = 1.0
sigma = 0.0
tau_m = 0.010
tau_z_pre_ampa = 0.050
tau_z_post_ampa = 0.005
tau_p = 10.0
hypercolumns = 1
minicolumns = 20
n_patterns = 20
# Manager properties
dt = 0.001
values_to_save = ['o', 'i_ampa', 'a']
# Protocol
training_time = 0.100
inter_sequence_interval = 0.0
inter_pulse_interval = 0.0
epochs = 1
mixed_start = False
contiguous = True
s = 1.0
r = 0.25
# Recall
T_recall = 1.0
T_cue = 0.020
num = 15
trials = 25
r_space = np.linspace(0, 0.6, num=num)
success_vector = np.zeros((num, trials))
factor = 0.1
for r_index, r in enumerate(r_space):
print(r_index)
# The network
nn = BCPNNPerfect(hypercolumns, minicolumns, g_w_ampa=g_w_ampa, g_w=g_w, g_a=g_a, tau_a=tau_a, tau_m=tau_m,
sigma=sigma, G=G, tau_z_pre_ampa=tau_z_pre_ampa, tau_z_post_ampa=tau_z_post_ampa, tau_p=tau_p,
z_transfer=z_transfer, diagonal_zero=diagonal_zero, strict_maximum=strict_maximum,
perfect=perfect, k_perfect=k_perfect, always_learning=always_learning,
normalized_currents=normalized_currents, g_beta=g_beta)
# Build the manager
manager = NetworkManager(nn=nn, dt=dt, values_to_save=values_to_save)
# The sequences
matrix = create_orthogonal_canonical_representation(minicolumns, hypercolumns)
aux = create_weights_from_two_sequences(nn, dt, n_patterns, s, r, mixed_start, contiguous,
training_time, inter_pulse_interval=inter_pulse_interval,
inter_sequence_interval=inter_sequence_interval, epochs=epochs,
resting_time=resting_time)
seq1, seq2, nr1, nr2, w_total, beta = aux
nr = np.concatenate((nr1, nr2))
aux, indexes = np.unique(nr, axis=0, return_index=True)
patterns_dic = {index:pattern for (index, pattern) in zip(indexes, aux)}
manager.patterns_dic = patterns_dic
nn.w_ampa = w_total
nn.beta = beta
current = g_w_ampa * (w_total[0, 0] - w_total[2, 0])
noise = factor * current
nn.sigma = noise
print(nn.sigma)
# Recall
for trial in range(trials):
s1 = calculate_recall_success_nr(manager, nr1, T_recall, T_cue)
s2 = calculate_recall_success_nr(manager, nr2, T_recall, T_cue)
success_vector[r_index, trial] = 0.5 * (s1 + s2)
# +
markersize = 15
linewdith = 8
current_palette = sns.color_palette()
index = 0
alpha = 0.5
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
mean_success = success_vector.mean(axis=1)
std = success_vector.std(axis=1)
ax.plot(r_space, mean_success, 'o-', lw=linewdith, ms=markersize)
ax.fill_between(r_space, mean_success - std, mean_success + std,
color=current_palette[index], alpha=alpha)
ax.axhline(0, ls='--', color='gray')
ax.axvline(0, ls='--', color='gray')
ax.set_xlabel('Overlap')
ax.set_ylabel('Recall')
# +
# %%time
always_learning = False
strict_maximum = True
perfect = False
z_transfer = False
k_perfect = True
diagonal_zero = False
normalized_currents = True
g_w_ampa = 2.0
g_w = 0.0
g_a = 10.0
tau_a = 0.250
g_beta = 0.0
G = 1.0
sigma = 0.0
tau_m = 0.010
tau_z_pre_ampa = 0.050
tau_z_post_ampa = 0.005
tau_p = 10.0
hypercolumns = 1
minicolumns = 20
n_patterns = 20
# Manager properties
dt = 0.001
values_to_save = ['o', 'i_ampa', 'a']
# Protocol
training_time = 0.100
inter_sequence_interval = 0.0
inter_pulse_interval = 0.0
epochs = 1
mixed_start = False
contiguous = True
s = 1.0
r = 0.25
# Recall
T_recall = 1.0
T_cue = 0.020
num = 15
trials = 25
r_space = np.linspace(0, 0.6, num=num)
success_vector = np.zeros((num, trials))
successes = []
factors = [0.0, 0.1, 0.2, 0.3]
for factor in factors:
print(factor)
for r_index, r in enumerate(r_space):
print(r_index)
# The network
nn = BCPNNPerfect(hypercolumns, minicolumns, g_w_ampa=g_w_ampa, g_w=g_w, g_a=g_a, tau_a=tau_a, tau_m=tau_m,
sigma=sigma, G=G, tau_z_pre_ampa=tau_z_pre_ampa, tau_z_post_ampa=tau_z_post_ampa, tau_p=tau_p,
z_transfer=z_transfer, diagonal_zero=diagonal_zero, strict_maximum=strict_maximum,
perfect=perfect, k_perfect=k_perfect, always_learning=always_learning,
normalized_currents=normalized_currents, g_beta=g_beta)
# Build the manager
manager = NetworkManager(nn=nn, dt=dt, values_to_save=values_to_save)
# The sequences
matrix = create_orthogonal_canonical_representation(minicolumns, hypercolumns)
aux = create_weights_from_two_sequences(nn, dt, n_patterns, s, r, mixed_start, contiguous,
training_time, inter_pulse_interval=inter_pulse_interval,
inter_sequence_interval=inter_sequence_interval, epochs=epochs,
resting_time=resting_time)
seq1, seq2, nr1, nr2, w_total, beta = aux
nr = np.concatenate((nr1, nr2))
aux, indexes = np.unique(nr, axis=0, return_index=True)
patterns_dic = {index:pattern for (index, pattern) in zip(indexes, aux)}
manager.patterns_dic = patterns_dic
nn.w_ampa = w_total
nn.beta = beta
current = g_w_ampa * (w_total[0, 0] - w_total[2, 0])
noise = factor * current
nn.sigma = noise
# Recall
for trial in range(trials):
s1 = calculate_recall_success_nr(manager, nr1, T_recall, T_cue)
s2 = calculate_recall_success_nr(manager, nr2, T_recall, T_cue)
success_vector[r_index, trial] = 0.5 * (s1 + s2)
successes.append(np.copy(success_vector))
# +
markersize = 15
linewdith = 8
current_palette = sns.color_palette()
index = 0
alpha = 0.5
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
for index, success_vector in enumerate(successes):
mean_success = success_vector.mean(axis=1)
std = success_vector.std(axis=1)
ax.plot(r_space, mean_success, 'o-', lw=linewdith, ms=markersize, label=str(factors[index]))
ax.fill_between(r_space, mean_success - std, mean_success + std,
color=current_palette[index], alpha=alpha)
ax.axhline(0, ls='--', color='gray')
ax.axvline(0, ls='--', color='gray')
ax.set_xlabel('Overlap')
ax.set_ylabel('Recall')
ax.legend();
# -
| jupyter/2018-05-23(Disambiguation).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Run the following two cells before you begin.**
# %autosave 10
# ______________________________________________________________________
# **First, import your data set and define the sigmoid function.**
# <details>
# <summary>Hint:</summary>
# The definition of the sigmoid is $f(x) = \frac{1}{1 + e^{-X}}$.
# </details>
# +
# Import the data set
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
import seaborn as sns
df = pd.read_csv('cleaned_data.csv')
# -
# Define the sigmoid function
def sigmoid(X):
Y = 1 / (1 + np.exp(-X))
return Y
# **Now, create a train/test split (80/20) with `PAY_1` and `LIMIT_BAL` as features and `default payment next month` as values. Use a random state of 24.**
# Create a train/test split
X_train, X_test, y_train, y_test = train_test_split(df[['PAY_1', 'LIMIT_BAL']].values, df['default payment next month'].values,test_size=0.2, random_state=24)
# ______________________________________________________________________
# **Next, import LogisticRegression, with the default options, but set the solver to `'liblinear'`.**
lr_model = LogisticRegression(solver='liblinear')
lr_model
# ______________________________________________________________________
# **Now, train on the training data and obtain predicted classes, as well as class probabilities, using the testing data.**
# Fit the logistic regression model on training data
lr_model.fit(X_train,y_train)
# Make predictions using `.predict()`
y_pred = lr_model.predict(X_test)
# Find class probabilities using `.predict_proba()`
y_pred_proba = lr_model.predict_proba(X_test)
# ______________________________________________________________________
# **Then, pull out the coefficients and intercept from the trained model and manually calculate predicted probabilities. You'll need to add a column of 1s to your features, to multiply by the intercept.**
# Add column of 1s to features
ones_and_features = np.hstack([np.ones((X_test.shape[0],1)), X_test])
print(ones_and_features)
np.ones((X_test.shape[0],1)).shape
# Get coefficients and intercepts from trained model
intercept_and_coefs = np.concatenate([lr_model.intercept_.reshape(1,1), lr_model.coef_], axis=1)
intercept_and_coefs
# Manually calculate predicted probabilities
X_lin_comb = np.dot(intercept_and_coefs, np.transpose(ones_and_features))
y_pred_proba_manual = sigmoid(X_lin_comb)
# ______________________________________________________________________
# **Next, using a threshold of `0.5`, manually calculate predicted classes. Compare this to the class predictions output by scikit-learn.**
# Manually calculate predicted classes
y_pred_manual = y_pred_proba_manual >= 0.5
y_pred_manual.shape
y_pred.shape
# Compare to scikit-learn's predicted classes
np.array_equal(y_pred.reshape(1,-1), y_pred_manual)
y_test.shape
y_pred_proba_manual.shape
# ______________________________________________________________________
# **Finally, calculate ROC AUC using both scikit-learn's predicted probabilities, and your manually predicted probabilities, and compare.**
# + eid="e7697"
# Use scikit-learn's predicted probabilities to calculate ROC AUC
from sklearn.metrics import roc_auc_score
roc_auc_score(y_test, y_pred_proba_manual.reshape(y_pred_proba_manual.shape[1],))
# -
# Use manually calculated predicted probabilities to calculate ROC AUC
roc_auc_score(y_test, y_pred_proba[:,1])
| Mini-Project-2/Project 4/Fitting_a_Logistic_Regression_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # ProVis: Attention Visualizer for Proteins
# + pycharm={"is_executing": false, "name": "#%%\n"}
import io
import urllib
import torch
from Bio.Data import SCOPData
from Bio.PDB import PDBParser, PPBuilder
from tape import TAPETokenizer, ProteinBertModel
import nglview
attn_color = [0.937, .522, 0.212]
# + pycharm={"name": "#%%\n"}
def get_structure(pdb_id):
resource = urllib.request.urlopen(f'https://files.rcsb.org/download/{pdb_id}.pdb')
content = resource.read().decode('utf8')
handle = io.StringIO(content)
parser = PDBParser(QUIET=True)
return parser.get_structure(pdb_id, handle)
# + pycharm={"name": "#%%\n"}
def get_attn_data(chain, layer, head, min_attn, start_index=0, end_index=None, max_seq_len=1024):
tokens = []
coords = []
for res in chain:
t = SCOPData.protein_letters_3to1.get(res.get_resname(), "X")
tokens += t
if t == 'X':
coord = None
else:
coord = res['CA'].coord.tolist()
coords.append(coord)
last_non_x = None
for i in reversed(range(len(tokens))):
if tokens[i] != 'X':
last_non_x = i
break
assert last_non_x is not None
tokens = tokens[:last_non_x + 1]
coords = coords[:last_non_x + 1]
tokenizer = TAPETokenizer()
model = ProteinBertModel.from_pretrained('bert-base', output_attentions=True)
if max_seq_len:
tokens = tokens[:max_seq_len - 2] # Account for SEP, CLS tokens (added in next step)
token_idxs = tokenizer.encode(tokens).tolist()
if max_seq_len:
assert len(token_idxs) == min(len(tokens) + 2, max_seq_len)
else:
assert len(token_idxs) == len(tokens) + 2
inputs = torch.tensor(token_idxs).unsqueeze(0)
with torch.no_grad():
attns = model(inputs)[-1]
# Remove attention from <CLS> (first) and <SEP> (last) token
attns = [attn[:, :, 1:-1, 1:-1] for attn in attns]
attns = torch.stack([attn.squeeze(0) for attn in attns])
attn = attns[layer, head]
if end_index is None:
end_index = len(tokens)
attn_data = []
for i in range(start_index, end_index):
for j in range(i, end_index):
# Currently non-directional: shows max of two attns
a = max(attn[i, j].item(), attn[j, i].item())
if a is not None and a >= min_attn:
attn_data.append((a, coords[i], coords[j]))
return attn_data
# -
# ### Visualize head 7-1 (targets binding sites)
# + pycharm={"is_executing": false, "name": "#%%\n"}
# Example for head 7-1 (targets binding sites)
pdb_id = '7HVP'
chain_ids = None # All chains
layer = 7
head = 1
min_attn = 0.1
attn_scale = .9
layer_zero_indexed = layer - 1
head_zero_indexed = head - 1
structure = get_structure(pdb_id)
view = nglview.show_biopython(structure)
view.stage.set_parameters(**{
"backgroundColor": "black",
"fogNear": 50, "fogFar": 100,
})
models = list(structure.get_models())
if len(models) > 1:
print('Warning:', len(models), 'models. Using first one')
prot_model = models[0]
if chain_ids is None:
chain_ids = [chain.id for chain in prot_model]
for chain_id in chain_ids:
print('Loading chain', chain_id)
chain = prot_model[chain_id]
attn_data = get_attn_data(chain, layer_zero_indexed, head_zero_indexed, min_attn)
for att, coords_from, coords_to in attn_data:
view.shape.add_cylinder(coords_from, coords_to, attn_color, att * attn_scale)
view
# -
# ### Visualize head 12-4 (targets contact maps)
# +
# Example for head 12-4 (targets contact maps)
pdb_id = '2KC7'
chain_ids = None # All chains
layer = 12
head = 4
min_attn = 0.2
attn_scale = .5
layer_zero_indexed = layer - 1
head_zero_indexed = head - 1
structure = get_structure(pdb_id)
view2 = nglview.show_biopython(structure)
view2.stage.set_parameters(**{
"backgroundColor": "black",
"fogNear": 50, "fogFar": 100,
})
models = list(structure.get_models())
if len(models) > 1:
print('Warning:', len(models), 'models. Using first one')
prot_model = models[0]
if chain_ids is None:
chain_ids = [chain.id for chain in prot_model]
for chain_id in chain_ids:
print('Loading chain', chain_id)
chain = prot_model[chain_id]
attn_data = get_attn_data(chain, layer_zero_indexed, head_zero_indexed, min_attn)
for att, coords_from, coords_to in attn_data:
view2.shape.add_cylinder(coords_from, coords_to, attn_color, att * attn_scale)
view2
# To save: view2.download_image(filename="testing.png")
# +
| notebooks/provis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SBTi-Finance Tool - Portfolio Aggregation
# In this notebook we'll give some examples on how the portfolio aggregation methods can be used.
#
# Please see the [methodology](https://sciencebasedtargets.org/wp-content/uploads/2020/09/Temperature-Rating-Methodology-V1.pdf), [guidance](https://sciencebasedtargets.org/wp-content/uploads/2020/10/Financial-Sector-Science-Based-Targets-Guidance-Pilot-Version.pdf) and the [technical documentation](http://getting-started.sbti-tool.org/) for more details on the different aggregation methods.
#
# See 1_analysis_example (on [Colab](https://colab.research.google.com/github/OFBDABV/SBTi/blob/master/examples/1_analysis_example.ipynb) or [Github](https://github.com/OFBDABV/SBTi/blob/master/examples/1_analysis_example.ipynb)) for more in depth example of how to work with Jupyter Notebooks in general and SBTi notebooks in particular.
#
# ## Setting up
# First we will set up the imports, data providers, and load the portfolio.
#
# For more examples of this process, please refer to notebook 1 & 2 (analysis and quick calculation example).
#
# !pip install SBTi
# %load_ext autoreload
# %autoreload 2
import SBTi
from SBTi.data.excel import ExcelProvider
from SBTi.portfolio_aggregation import PortfolioAggregationMethod
from SBTi.portfolio_coverage_tvp import PortfolioCoverageTVP
from SBTi.temperature_score import TemperatureScore, Scenario, ScenarioType, EngagementType
from SBTi.target_validation import TargetProtocol
from SBTi.interfaces import ETimeFrames, EScope
# %aimport -pandas
import pandas as pd
# +
# Download the dummy data
import urllib.request
import os
if not os.path.isdir("data"):
os.mkdir("data")
if not os.path.isfile("data/data_provider_example.xlsx"):
urllib.request.urlretrieve("https://github.com/OFBDABV/SBTi/raw/master/examples/data/data_provider_example.xlsx", "data/data_provider_example.xlsx")
if not os.path.isfile("data/example_portfolio.csv"):
urllib.request.urlretrieve("https://github.com/OFBDABV/SBTi/raw/master/examples/data/example_portfolio.csv", "data/example_portfolio.csv")
# -
provider = ExcelProvider(path="data/data_provider_example.xlsx")
df_portfolio = pd.read_csv("data/example_portfolio.csv", encoding="iso-8859-1")
companies = SBTi.utils.dataframe_to_portfolio(df_portfolio)
scores_collection = {}
temperature_score = TemperatureScore(time_frames=list(SBTi.interfaces.ETimeFrames), scopes=[EScope.S1S2, EScope.S3, EScope.S1S2S3])
amended_portfolio = temperature_score.calculate(data_providers=[provider], portfolio=companies)
# ## Calculate the aggregated temperature score
# Calculate an aggregated temperature score. This can be done using different aggregation methods. The termperature scores are calculated per time-frame/scope combination.
# + [markdown] pycharm={"name": "#%% md\n"}
# ### WATS
# Weighted Average Temperature Score (WATS): Temperature scores are allocated based on portfolio weights.
# This method uses the "investment_value" field to be defined in your portfolio data.
# + pycharm={"name": "#%%\n"}
temperature_score.aggregation_method = PortfolioAggregationMethod.WATS
aggregated_scores = temperature_score.aggregate_scores(amended_portfolio)
df_wats = pd.DataFrame(aggregated_scores.dict()).applymap(lambda x: round(x['all']['score'], 2))
scores_collection.update({'WATS': df_wats})
df_wats
# + [markdown] pycharm={"name": "#%% md\n"}
# ### TETS
# Total emissions weighted temperature score (TETS): Temperature scores are allocated based on historical emission weights using total company emissions.
# In addition to the portfolios "investment value" the TETS method requires company emissions, please refer to [Data Legends - Fundamental Data](https://ofbdabv.github.io/SBTi/Legends.html#fundamental-data) for more details
# + pycharm={"name": "#%%\n"}
temperature_score.aggregation_method = PortfolioAggregationMethod.TETS
aggregated_scores = temperature_score.aggregate_scores(amended_portfolio)
df_tets = pd.DataFrame(aggregated_scores.dict()).applymap(lambda x: round(x['all']['score'], 2))
scores_collection.update({'TETS': df_tets})
df_tets
# + [markdown] pycharm={"name": "#%% md\n"}
# ### MOTS
# Market Owned emissions weighted temperature score (MOTS): Temperature scores are allocated based on an equity ownership approach.
# In addition to the portfolios "investment value" the MOTS method requires company emissions and market cap, please refer to [Data Legends - Fundamental Data](https://ofbdabv.github.io/SBTi/Legends.html#fundamental-data) for more details
# + pycharm={"name": "#%%\n"}
temperature_score.aggregation_method = PortfolioAggregationMethod.MOTS
aggregated_scores = temperature_score.aggregate_scores(amended_portfolio)
df_mots = pd.DataFrame(aggregated_scores.dict()).applymap(lambda x: round(x['all']['score'], 2))
scores_collection.update({'MOTS': df_mots})
df_mots
# + [markdown] pycharm={"name": "#%% md\n"}
# ### EOTS
# Enterprise Owned emissions weighted temperature score (EOTS): Temperature scores are allocated based
# on an enterprise ownership approach.
# In addition to the portfolios "investment value" the EOTS method requires company emissions and enterprise value, please refer to [Data Legends - Fundamental Data](https://ofbdabv.github.io/SBTi/Legends.html#fundamental-data) for more details
# + pycharm={"name": "#%%\n"}
temperature_score.aggregation_method = PortfolioAggregationMethod.EOTS
aggregated_scores = temperature_score.aggregate_scores(amended_portfolio)
df_eots = pd.DataFrame(aggregated_scores.dict()).applymap(lambda x: round(x['all']['score'], 2))
scores_collection.update({'EOTS': df_eots})
df_eots
# + [markdown] pycharm={"name": "#%% md\n"}
# ### ECOTS
# Enterprise Value + Cash emissions weighted temperature score (ECOTS): Temperature scores are allocated based on an enterprise value (EV) plus cash & equivalents ownership approach.
# In addition to the portfolios "investment value" the ECOTS method requires company emissions, company cash equivalents and enterprise value; please refer to [Data Legends - Fundamental Data](https://ofbdabv.github.io/SBTi/Legends.html#fundamental-data) for more details
# + pycharm={"name": "#%%\n"}
temperature_score.aggregation_method = PortfolioAggregationMethod.ECOTS
aggregated_scores = temperature_score.aggregate_scores(amended_portfolio)
df_ecots = pd.DataFrame(aggregated_scores.dict()).applymap(lambda x: round(x['all']['score'], 2))
scores_collection.update({'ECOTS': df_ecots})
df_ecots
# + [markdown] pycharm={"name": "#%% md\n"}
# ### AOTS
# Total Assets emissions weighted temperature score (AOTS): Temperature scores are allocated based on a total assets ownership approach.
# In addition to the portfolios "investment value" the AOTS method requires company emissions and company total assets; please refer to [Data Legends - Fundamental Data](https://ofbdabv.github.io/SBTi/Legends.html#fundamental-data) for more details
# + pycharm={"name": "#%%\n"}
temperature_score.aggregation_method = PortfolioAggregationMethod.AOTS
aggregated_scores = temperature_score.aggregate_scores(amended_portfolio)
df_aots = pd.DataFrame(aggregated_scores.dict()).applymap(lambda x: round(x['all']['score'], 2))
scores_collection.update({'AOTS': df_aots})
df_aots
# + [markdown] pycharm={"name": "#%% md\n"}
# ### ROTS
# Revenue owned emissions weighted temperature score (ROTS): Temperature scores are allocated based on the share of revenue.
# In addition to the portfolios "investment value" the ROTS method requires company emissions and company revenue; please refer to [Data Legends - Fundamental Data](https://ofbdabv.github.io/SBTi/Legends.html#fundamental-data) for more details
# + pycharm={"name": "#%%\n"}
temperature_score.aggregation_method = PortfolioAggregationMethod.ROTS
aggregated_scores = temperature_score.aggregate_scores(amended_portfolio)
df_rots = pd.DataFrame(aggregated_scores.dict()).applymap(lambda x: round(x['all']['score'], 2))
scores_collection.update({'ROTS': df_rots})
df_rots
# + [markdown] pycharm={"name": "#%% md\n"}
# See below how each aggregation method impact the scores on for each time frame and scope combination
# + pycharm={"name": "#%%\n"}
pd.concat(scores_collection, axis=0)
| examples/4_portfolio_aggregations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] colab_type="text" id="1Pi_B2cvdBiW"
# ##### Copyright 2019 The TF-Agents Authors.
# + [markdown] colab_type="text" id="f5926O3VkG_p"
# ### Get Started
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/agents/blob/master/tf_agents/colabs/8_networks_tutorial.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/agents/blob/master/tf_agents/colabs/8_networks_tutorial.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + colab_type="code" id="xsLTHlVdiZP3" colab={}
# Note: If you haven't installed tf-agents yet, run:
# !pip install tf-nightly
# !pip install tfp-nightly
# !pip install tf-agents-nightly
# + [markdown] colab_type="text" id="lEgSa5qGdItD"
# ### Imports
# + colab_type="code" id="sdvop99JlYSM" colab={}
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import abc
import tensorflow as tf
import numpy as np
from tf_agents.environments import random_py_environment
from tf_agents.environments import tf_py_environment
from tf_agents.networks import encoding_network
from tf_agents.networks import network
from tf_agents.networks import utils
from tf_agents.specs import array_spec
from tf_agents.utils import common as common_utils
from tf_agents.utils import nest_utils
tf.compat.v1.enable_v2_behavior()
# + [markdown] colab_type="text" id="31uij8nIo5bG"
# # Introduction
#
# In this colab we will cover how to define custom networks for your agents. The networks help us define the model that is trained by agents. In TF-Agents you will find several different types of networks which are useful across agents:
#
# **Main Networks**
#
# * **QNetwork**: Used in Qlearning for environments with discrete actions, this network maps an observation to value estimates for each possible action.
# * **CriticNetworks**: Also referred to as `ValueNetworks` in literature, learns to estimate some version of a Value function mapping some state into an estimate for the expected return of a policy. These networks estimate how good the state the agent is currently in is.
# * **ActorNetworks**: Learn a mapping from observations to actions. These networks are usually used by our policies to generate actions.
# * **ActorDistributionNetworks**: Similar to `ActorNetworks` but these generate a distribution which a policy can then sample to generate actions.
#
# **Helper Networks**
# * **EncodingNetwork**: Allows users to easily define a mapping of pre-processing layers to apply to a network's input.
# * **DynamicUnrollLayer**: Automatically resets the network's state on episode boundaries as it is applied over a time sequence.
# * **ProjectionNetwork**: Networks like `CategoricalProjectionNetwork` or `NormalProjectionNetwork` take inputs and generate the required parameters to generate Categorical, or Normal distributions.
#
# All examples in TF-Agents come with pre-configured networks. However these networks are not setup to handle complex observations.
#
# If you have an environment which exposes more than one observation/action and you need to customize your networks then this tutorial is for you!
# + [markdown] id="ums84-YP_21F" colab_type="text"
# #Defining Networks
#
# ##Network API
#
# In TF-Agents we subclass from Keras [Networks](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/keras/engine/network.py). With it we can:
#
# * Simplify copy operations required when creating target networks.
# * Perform automatic variable creation when calling `network.variables()`.
# * Validate inputs based on network input_specs.
#
# ##EncodingNetwork
# As mentioned above the `EncodingNetwork` allows us to easily define a mapping of pre-processing layers to apply to a network's input to generate some encoding.
#
# The EncodingNetwork is composed of the following mostly optional layers:
#
# * Preprocessing layers
# * Preprocessing combiner
# * Conv2D
# * Flatten
# * Dense
#
# The special thing about encoding networks is that input preprocessing is applied. Input preprocessing is possible via `preprocessing_layers` and `preprocessing_combiner` layers. Each of these can be specified as a nested structure. If the `preprocessing_layers` nest is shallower than `input_tensor_spec`, then the layers will get the subnests. For example, if:
#
# ```
# input_tensor_spec = ([TensorSpec(3)] * 2, [TensorSpec(3)] * 5)
# preprocessing_layers = (Layer1(), Layer2())
# ```
#
# then preprocessing will call:
#
# ```
# preprocessed = [preprocessing_layers[0](observations[0]),
# preprocessing_layers[1](obsrevations[1])]
# ```
#
# However if
#
# ```
# preprocessing_layers = ([Layer1() for _ in range(2)],
# [Layer2() for _ in range(5)])
# ```
#
# then preprocessing will call:
#
# ```python
# preprocessed = [
# layer(obs) for layer, obs in zip(flatten(preprocessing_layers),
# flatten(observations))
# ]
# ```
#
# + [markdown] id="RP3H1bw0ykro" colab_type="text"
# ## Custom Networks
#
# To create your own networks you will only have to override the `__init__` and `__call__` methods. Let's create a custom network using what we learned about `EncodingNetworks` to create an ActorNetwork that takes observations which contain an image and a vector.
#
# + id="Zp0TjAJhYo4s" colab_type="code" colab={}
class ActorNetwork(network.Network):
def __init__(self,
observation_spec,
action_spec,
preprocessing_layers=None,
preprocessing_combiner=None,
conv_layer_params=None,
fc_layer_params=(75, 40),
dropout_layer_params=None,
activation_fn=tf.keras.activations.relu,
enable_last_layer_zero_initializer=False,
name='ActorNetwork'):
super(ActorNetwork, self).__init__(
input_tensor_spec=observation_spec, state_spec=(), name=name)
# For simplicity we will only support a single action float output.
self._action_spec = action_spec
flat_action_spec = tf.nest.flatten(action_spec)
if len(flat_action_spec) > 1:
raise ValueError('Only a single action is supported by this network')
self._single_action_spec = flat_action_spec[0]
if self._single_action_spec.dtype not in [tf.float32, tf.float64]:
raise ValueError('Only float actions are supported by this network.')
kernel_initializer = tf.keras.initializers.VarianceScaling(
scale=1. / 3., mode='fan_in', distribution='uniform')
self._encoder = encoding_network.EncodingNetwork(
observation_spec,
preprocessing_layers=preprocessing_layers,
preprocessing_combiner=preprocessing_combiner,
conv_layer_params=conv_layer_params,
fc_layer_params=fc_layer_params,
dropout_layer_params=dropout_layer_params,
activation_fn=activation_fn,
kernel_initializer=kernel_initializer,
batch_squash=False)
initializer = tf.keras.initializers.RandomUniform(
minval=-0.003, maxval=0.003)
self._action_projection_layer = tf.keras.layers.Dense(
flat_action_spec[0].shape.num_elements(),
activation=tf.keras.activations.tanh,
kernel_initializer=initializer,
name='action')
def call(self, observations, step_type=(), network_state=()):
outer_rank = nest_utils.get_outer_rank(observations, self.input_tensor_spec)
# We use batch_squash here in case the observations have a time sequence
# compoment.
batch_squash = utils.BatchSquash(outer_rank)
observations = tf.nest.map_structure(batch_squash.flatten, observations)
state, network_state = self._encoder(
observations, step_type=step_type, network_state=network_state)
actions = self._action_projection_layer(state)
actions = common_utils.scale_to_spec(actions, self._single_action_spec)
actions = batch_squash.unflatten(actions)
return tf.nest.pack_sequence_as(self._action_spec, [actions]), network_state
# + [markdown] id="Fm-MbMMLYiZj" colab_type="text"
# Let's create a `RandomPyEnvironment` to generate structured observations and validate our implementation.
# + id="E2XoNuuD66s5" colab_type="code" colab={}
action_spec = array_spec.BoundedArraySpec((3,), np.float32, minimum=0, maximum=10)
observation_spec = {
'image': array_spec.BoundedArraySpec((16, 16, 3), np.float32, minimum=0,
maximum=255),
'vector': array_spec.BoundedArraySpec((5,), np.float32, minimum=-100,
maximum=100)}
random_env = random_py_environment.RandomPyEnvironment(observation_spec, action_spec=action_spec)
# Convert the environment to a TFEnv to generate tensors.
tf_env = tf_py_environment.TFPyEnvironment(random_env)
# + [markdown] id="LM3uDTD7TNVx" colab_type="text"
# Since we've defined the observations to be a dict we need to create preprocessing layers to handle these.
# + id="r9U6JVevTAJw" colab_type="code" colab={}
preprocessing_layers = {
'image': tf.keras.models.Sequential([tf.keras.layers.Conv2D(8, 4),
tf.keras.layers.Flatten()]),
'vector': tf.keras.layers.Dense(5)
}
preprocessing_combiner = tf.keras.layers.Concatenate(axis=-1)
actor = ActorNetwork(tf_env.observation_spec(),
tf_env.action_spec(),
preprocessing_layers=preprocessing_layers,
preprocessing_combiner=preprocessing_combiner)
# + [markdown] id="mM9qedlwc41U" colab_type="text"
# Now that we have the actor network we can process observations from the environment.
# + id="JOkkeu7vXoei" colab_type="code" colab={}
time_step = tf_env.reset()
actor(time_step.observation, time_step.step_type)
# + [markdown] id="ALGxaQLWc9GI" colab_type="text"
# This same strategy can be used to customize any of the main networks used by the agents. You can define whatever preprocessing and connect it to the rest of the network. As you define your own custom make sure the output layer definitions of the network match.
| tf_agents/colabs/8_networks_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Python for STEM Teachers<br/>[Oregon Curriculum Network](http://4dsolutions.net/ocn/)
#
# # Atoms R Us
#
# 
#
# +
import json
series_types = ["Don't Know", "Other nonmetal", "Alkali metal",
"Alkaline earth metal", "Nobel gas", "Metalloid",
"Halogen", "Transition metal", "Post-transition metal",
"Lanthanoid", "Actinoid"]
class Element:
fields = "protons symbol long_name mass series"
repstr = ("Atom(protons={protons}, symbol='{symbol}', "
"long_name='{long_name}', "
"mass={mass}, series='{series}')")
def __init__(self, protons: int, symbol: str,
long_name: str, mass: float, series: str):
# build self.__dict__
self.protons = protons
self.symbol = symbol
self.long_name = long_name
self.__dict__['mass'] = mass # same idea
self.series = series
def __getitem__(self, idx): # simulates collection.namedtuple behavior
return self.__dict__[self.fields[idx]]
def __repr__(self):
return self.repstr.format(**self.__dict__)
Atom = Element # synonyms
lithium = Atom(3, "Li", "Lithium", 6.941, "Alkali metal")
print(lithium) # __str__, then __repr__
print(lithium.__dict__)
print(lithium.protons) # print(lithium.__getattr__('protons'))
# +
import unittest
class Test_Element(unittest.TestCase):
def test_instance(self):
lithium = Atom(3, "Li", "Lithium", 6.941, "Alkali metal")
self.assertEqual(lithium.protons, 3, "Houston, we have a problem")
a = Test_Element() # the test suite
suite = unittest.TestLoader().loadTestsFromModule(a) # fancy boilerplate
unittest.TextTestRunner().run(suite) # run the test suite
# +
class ElementEncoder(json.JSONEncoder):
"""
See: https://docs.python.org/3.5/library/json.html
"""
def default(self, obj):
if isinstance(obj, Element): # how to encode an Element
return [obj.protons, obj.symbol, obj.long_name, obj.mass, obj.series]
return json.JSONEncoder.default(self, obj) # just do your usual
# Element = namedtuple("Atom", "protons abbrev long_name mass")
def load_elements():
global all_elements # <--- will be visible to entire module
try:
the_file = "periodic_table.json"
f = open(the_file, "r") # <--- open the_file instead
except IOError:
print("Sorry, no such file!")
else:
the_dict = json.load(f)
f.close()
all_elements = {}
for symbol, data in the_dict.items():
all_elements[symbol] = Atom(*data) # "explode" data into 5 inputs
print("File:", the_file, 'loaded.')
load_elements() # actually do it
# -
# 
# <div align="center">graphic by <NAME></div>
# +
def print_periodic_table(sortby=1):
"""
sort all_elements by number of protons, ordered_elements local only
What about series?
Sort Order:
1. protons
2. symbol
3. series
"""
print("Selected:", sortby)
if sortby == 1:
ordered_elements = sorted(all_elements.values(), key = lambda k: k.protons)
elif sortby == 2:
ordered_elements = sorted(all_elements.values(), key = lambda k: k.symbol)
elif sortby == 3:
ordered_elements = sorted(all_elements.values(), key = lambda k: k.series)
print("PERIODIC TABLE OF THE ELEMENTS")
print("-" * 70)
print("Symbol |Long Name |Protons |Mass |Series " )
print("-" * 70)
for the_atom in ordered_elements:
print("{:6} | {:20} | {:6} | {:5.2f} | {:15}".format(the_atom.symbol,
the_atom.long_name,
the_atom.protons,
the_atom.mass,
the_atom.series))
print_periodic_table() # do it for real
# -
# 
# <div align="center">by <NAME></div>
| Atoms in Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflowjs as tfjs
import json
# After saving keras model with ZeroPadding3D as model.json Modify with this code:
# Open json file to modify
with open(output_folder + 'model.json') as f:
model_dict = json.load(f)
# Convert
for layer in model_dict['modelTopology']['model_config']['config']['layers']:
if layer['class_name'] == "ZeroPadding3D":
model_dict['modelTopology']['model_config']['config']['layers'].remove(layer)
prev_layer_name = ""
for layer in model_dict['modelTopology']['model_config']['config']['layers']:
if layer['class_name'] == "InputLayer":
layer["config"]["batch_input_shape"] = [None, 38, 38, 38, 1]
if layer['class_name'] == "Conv3D":
layer["config"]["padding"] = "same"
layer["config"]["data_format"] = "channels_last"
layer['inbound_nodes'][0][0][0] = prev_layer_name
prev_layer_name = layer["config"]["name"]
# Verification
for layer in model_dict['modelTopology']['model_config']['config']['layers']:
print(layer)
print("-------------------------------------------------------")
# Save model.json file
with open(output_folder + 'model.json', 'w') as fp:
json.dump(model_dict, fp)
| python/Conversion/Convert model.json with ZeroPadding3D to tfjs compatible json.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="eerkOkIpLQD2"
# Author: <NAME>, <EMAIL>; <NAME>, <EMAIL>; <NAME>, <EMAIL> (2021)
# + id="EAcBQ5Y9E-aw" executionInfo={"status": "ok", "timestamp": 1636962210309, "user_tz": 480, "elapsed": 354, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNA4F55Dr3Wpfy_3xDEtdDeKDmfL_WiSi81FRmoQ=s64", "userId": "11440633380440656636"}}
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import warnings
import seaborn as sns
import matplotlib
matplotlib.rcParams.update({'font.size': 13})
warnings.filterwarnings("ignore")
# + [markdown] id="Gh2ObfFDZOWa"
# ## Two Datasets
#
# Dataset1: n = 200
#
# Dataset2: n = 50
# + id="ppTpI6E7ExzO" executionInfo={"status": "ok", "timestamp": 1636962210309, "user_tz": 480, "elapsed": 6, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNA4F55Dr3Wpfy_3xDEtdDeKDmfL_WiSi81FRmoQ=s64", "userId": "11440633380440656636"}}
np.random.seed(10)
dataset1 = np.random.multivariate_normal(np.array([2,1]), np.array([[1, 1.5], [1.5, 3]]), 200)
dataset2 = dataset1[:50,:] + np.random.multivariate_normal(np.array([0,0]), np.array([[0.1, 0], [0, 0.1]]), 50)
dataset1 = pd.DataFrame(dataset1,columns = ['X','Y'])
dataset2 = pd.DataFrame(dataset2,columns = ['X','Y'])
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="Jcd97pw8E-7D" executionInfo={"status": "ok", "timestamp": 1636962210727, "user_tz": 480, "elapsed": 423, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNA4F55Dr3Wpfy_3xDEtdDeKDmfL_WiSi81FRmoQ=s64", "userId": "11440633380440656636"}} outputId="09bec41e-e969-4e2e-d9a1-7b5ec30ddb7b"
plt.scatter(dataset1['X'],dataset1['Y'])
plt.xlabel('X')
plt.ylabel('Y')
plt.title('Dataset 1: # of sample: 200')
plt.xlim(-1,5)
plt.ylim(-5,6)
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="8JBzBJlhFdFb" executionInfo={"status": "ok", "timestamp": 1614120461791, "user_tz": 480, "elapsed": 1791, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikqN0fra_DdutXH7CWyxA7GsB2Na6ivKIwnTh_OTw=s64", "userId": "14594676763170300704"}} outputId="29a567c3-2a29-475b-9004-685cd3d65de2"
plt.scatter(dataset2['X'],dataset2['Y'],c = 'C1')
plt.xlabel('X')
plt.ylabel('Y')
plt.title('Dataset2: # of sample: 50')
plt.xlim(-1,5)
plt.ylim(-5,6)
# + [markdown] id="XyabMB_OZf0b"
# ## Fit linear regression
# + id="jn9RfVenFg6p"
import statsmodels.formula.api as smf
linear_reg_dataset1 = smf.ols(formula='Y ~ X', data=dataset1)
linear_reg_dataset1 = linear_reg_dataset1.fit()
linear_reg_dataset2 = smf.ols(formula='Y ~ X', data=dataset2)
linear_reg_dataset2 = linear_reg_dataset2.fit()
# + [markdown] id="E6K71IBxZsZI"
# Linear regression: fit dataset 1
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="wBTgUMtybDO6" executionInfo={"status": "ok", "timestamp": 1614120462695, "user_tz": 480, "elapsed": 2687, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikqN0fra_DdutXH7CWyxA7GsB2Na6ivKIwnTh_OTw=s64", "userId": "14594676763170300704"}} outputId="04ef5372-a20e-4ee6-fd27-7d5a9e1c4d6e"
X_pred = np.linspace(-2,6,num = 100)
reg = linear_reg_dataset1
data = dataset1
plt.plot(data['X'], data['Y'], 'o', label="Data")
plt.plot(X_pred,reg.predict(exog=dict(X=X_pred)), 'r-', label="Predicted")
plt.title('Dataset 1, '+'R2: '+str(np.round(reg.rsquared,2)))
plt.legend(loc="best")
plt.xlim(-1,5)
plt.ylim(-5,6)
plt.xlabel('X')
plt.ylabel('Y')
# + colab={"base_uri": "https://localhost:8080/"} id="LTdxc8XaYKrH" executionInfo={"status": "ok", "timestamp": 1614120462696, "user_tz": 480, "elapsed": 2683, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikqN0fra_DdutXH7CWyxA7GsB2Na6ivKIwnTh_OTw=s64", "userId": "14594676763170300704"}} outputId="393c194f-4639-429f-b5c0-905072a14d6b"
print(linear_reg_dataset1.summary())
# + [markdown] id="mEHBuLgaZx9o"
# Linear regression: fit dataset 2
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="kQLt4QkbbELu" executionInfo={"status": "ok", "timestamp": 1614120462893, "user_tz": 480, "elapsed": 2876, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikqN0fra_DdutXH7CWyxA7GsB2Na6ivKIwnTh_OTw=s64", "userId": "14594676763170300704"}} outputId="5142baeb-383f-4bbb-82bd-6af2235cbec9"
X_pred = np.linspace(-2,6,num = 100)
reg = linear_reg_dataset2
data = dataset2
plt.plot(data['X'], data['Y'], 'o', label="Data",c = 'C1')
plt.plot(X_pred,reg.predict(exog=dict(X=X_pred)), 'r-', label="Predicted")
plt.title('Dataset 2, '+'R2: '+str(np.round(reg.rsquared,2)))
plt.legend(loc="best")
plt.xlim(-1,5)
plt.ylim(-5,6)
plt.xlabel('X')
plt.ylabel('Y')
# + colab={"base_uri": "https://localhost:8080/"} id="T6epM9JTZrvW" executionInfo={"status": "ok", "timestamp": 1614120462894, "user_tz": 480, "elapsed": 2873, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikqN0fra_DdutXH7CWyxA7GsB2Na6ivKIwnTh_OTw=s64", "userId": "14594676763170300704"}} outputId="6d6c0c8b-a69e-45ec-9d2b-5d528dd8d76d"
print(linear_reg_dataset2.summary())
# + [markdown] id="nS0lLJM8Z6CG"
# ## Residual
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="3Dqi4YlVYTFX" executionInfo={"status": "ok", "timestamp": 1614120463329, "user_tz": 480, "elapsed": 3304, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikqN0fra_DdutXH7CWyxA7GsB2Na6ivKIwnTh_OTw=s64", "userId": "14594676763170300704"}} outputId="800b8f7f-8367-4ff3-940f-ac2099e8f81b"
plt.plot(linear_reg_dataset1.resid,'.', label = 'Residual')
plt.hlines(y = 0, xmin = 0, xmax = 200, color = 'r')
plt.xlabel('# of samples')
plt.ylabel('Residual')
plt.title('Dataset 1, residual')
plt.legend(loc="best")
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="AOB-9W9kF7Yi" executionInfo={"status": "ok", "timestamp": 1614120463674, "user_tz": 480, "elapsed": 3643, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikqN0fra_DdutXH7CWyxA7GsB2Na6ivKIwnTh_OTw=s64", "userId": "14594676763170300704"}} outputId="03b8ec3a-e2cc-4abb-f0e5-15a5e96338e7"
plt.plot(linear_reg_dataset2.resid,'.', label = 'Residual',c = 'C1')
plt.hlines(y = 0, xmin = 0, xmax = 50, color = 'r')
plt.xlabel('# of samples')
plt.ylabel('Residual')
plt.title('Dataset 2, residual')
plt.legend(loc="best")
# + [markdown] id="Rr-8e4T4bc47"
# ## 95% confidence interval
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="UoXUXKeCDq4X" executionInfo={"status": "ok", "timestamp": 1614120464154, "user_tz": 480, "elapsed": 4118, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikqN0fra_DdutXH7CWyxA7GsB2Na6ivKIwnTh_OTw=s64", "userId": "14594676763170300704"}} outputId="fcfe240e-484f-445d-c333-e25ea2630609"
import seaborn as sns
data = dataset1
sns.regplot(data['X'],data['Y'],color = 'C0')
plt.title('Dataset 1: confidence interval')
plt.xlim(-1,5)
plt.ylim(-5,6)
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="mnJwvy9_ErQS" executionInfo={"status": "ok", "timestamp": 1614120464315, "user_tz": 480, "elapsed": 4274, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikqN0fra_DdutXH7CWyxA7GsB2Na6ivKIwnTh_OTw=s64", "userId": "14594676763170300704"}} outputId="0a0278e3-cb21-48c4-cd16-458e408ac0a7"
data = dataset1
sns.regplot(data['X'],data['Y'],color = 'C0',scatter = False)
plt.title('Dataset 1: confidence interval')
plt.xlim(-1,5)
plt.ylim(-5,6)
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="lsJbFTHEDaoC" executionInfo={"status": "ok", "timestamp": 1614120464769, "user_tz": 480, "elapsed": 4723, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikqN0fra_DdutXH7CWyxA7GsB2Na6ivKIwnTh_OTw=s64", "userId": "14594676763170300704"}} outputId="2c6aa292-b914-4473-9efa-2dec94842045"
data = dataset2
sns.regplot(data['X'],data['Y'],color = 'C1')
plt.title('Dataset 2: confidence interval')
plt.xlim(-1,5)
plt.ylim(-5,6)
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="t_EykV-hEusv" executionInfo={"status": "ok", "timestamp": 1614120465088, "user_tz": 480, "elapsed": 5037, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikqN0fra_DdutXH7CWyxA7GsB2Na6ivKIwnTh_OTw=s64", "userId": "14594676763170300704"}} outputId="81e1483f-af13-494e-9a91-b19abd69a66f"
data = dataset2
sns.regplot(data['X'],data['Y'],color = 'C1',scatter = False)
plt.title('Dataset 2: confidence interval')
plt.xlim(-1,5)
plt.ylim(-5,6)
| Ch3_SpatialAggregation/Colabs/Spatial Aggregation_ linear regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Attribute data operations {#attr}
#
# ## Prerequisites
#| echo: false
import pandas as pd
import matplotlib.pyplot as plt
pd.options.display.max_rows = 6
pd.options.display.max_columns = 6
pd.options.display.max_colwidth = 35
plt.rcParams["figure.figsize"] = (5, 5)
# Packages...
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import geopandas as gpd
import rasterio
# Sample data...
#| echo: false
from pathlib import Path
data_path = Path("data")
file_path = Path("data/landsat.tif")
if not file_path.exists():
if not data_path.is_dir():
os
os.mkdir(data_path)
import os
print("Attempting to get the data")
import requests
r = requests.get("https://github.com/geocompr/py/releases/download/0.1/landsat.tif")
with open(file_path, "wb") as f:
f.write(r.content)
world = gpd.read_file("data/world.gpkg")
src_elev = rasterio.open("data/elev.tif")
src_multi_rast = rasterio.open("data/landsat.tif")
# ## Introduction
#
# ...
#
# ## Vector attribute manipulation
#
# As mentioned previously (...), vector layers (`GeoDataFrame`, from package `geopandas`) are basically extended tables (`DataFrame` from package `pandas`), the difference being that a vector layer has a geometry column. Since `GeoDataFrame` extends `DataFrame`, all ordinary table-related operations from package `pandas` are supported for vector laters as well, as shown below.
#
# ### Vector attribute subsetting
#
# `pandas` supports several subsetting interfaces, though the most [recommended](https://stackoverflow.com/questions/38886080/python-pandas-series-why-use-loc) ones are:
#
# * `.loc`, which uses pandas indices, and
# * `.iloc`, which uses (implicit) numpy-style numeric indices.
#
# In both cases the method is followed by square brackets, and two indices, separated by a comma. Each index can comprise:
#
# * A specific value, as in `1`
# * A slice, as in `0:3`
# * A `list`, as in `[0,2,4]`
# * `:`—indicating "all" indices
#
# The once exception which we are going to with subsetting by indices is when selecting columns, directly using a list, as in `df[["a","b"]]`, instead of `df.loc[:, ["a","b"]]`, to select columns `"a"` and `"b"` from `df`.
#
# Here are few examples of subsetting the `GeoDataFrame` of world countries.
#
# Subsetting rows by position:
world.iloc[0:3, :]
# Subsetting columns by position:
world.iloc[:, 0:3]
# Subsetting rows and columns by position:
world.iloc[0:3, 0:3]
# Subsetting columns by name:
world[["name_long", "geometry"]]
# "Slice" of columns between given ones:
world.loc[:, "name_long":"pop"]
# Subsetting by a boolean series:
x = np.array([1, 1, 0, 0, 0, 0, 0, 1, 1, 0, 0], dtype=bool)
world.iloc[:, x]
# We can remove specific columns using the `.drop` method and `axis=1` (i.e., columns):
world.drop(["name_long", "continent"], axis=1)
# We can rename (some of) the selected columns using the `.rename` method:
world[["name_long", "pop"]].rename(columns={"pop": "population"})
# The standard `numpy` comparison operators can be used in boolean subsetting, as illustrated in Table ...
#
# TABLE ...: Comparison operators that return Booleans (TRUE/FALSE).
#
# |`Symbol` | `Name` |
# |---|---|
# | `==` | Equal to |
# | `!=` | Not equal to |
# | `>`, `<` | Greater/Less than |
# | `>=`, `<=` | Greater/Less than or equal |
# | `&`, `|`, `~` | Logical operators: And, Or, Not |
#
# A demonstration of the utility of using logical vectors for subsetting is shown in the code chunk below. This creates a new object, small_countries, containing nations whose surface area is smaller than 10,000 km^2^:
i_small = world["area_km2"] < 10000 ## a logical 'Series'
small_countries = world[i_small]
small_countries
# The intermediary `i_small` (short for index representing small countries) is a boolean `Series` that can be used to subset the seven smallest countries in the world by surface area. A more concise command, which omits the intermediary object, generates the same result:
small_countries = world[world["area_km2"] < 10000]
# The various methods shown above can be chained for any combination with several subsetting steps. For example:
world[world["continent"] == "Asia"] \
.loc[:, ["name_long", "continent"]] \
.iloc[0:5, :]
# ### Vector attribute aggregation
#
# Aggregation involves summarizing data with one or more *grouping variables*, typically from columns in the table to be aggregated (geographic aggregation is covered in the next chapter). An example of attribute aggregation is calculating the number of people per continent based on country-level data (one row per country). The `world` dataset contains the necessary ingredients: the columns `pop` and `continent`, the population and the grouping variable, respectively. The aim is to find the `sum()` of country populations for each continent, resulting in a smaller data frame (aggregation is a form of data reduction and can be a useful early step when working with large datasets). This can be done with a combination of `.groupby` and `.sum`:
world_agg1 = world[['continent', 'pop']].groupby('continent').sum()
world_agg1
# The result is a (non-spatial) table with eight rows, one per continent, and two columns reporting the name and population of each continent.
#
# Alternatively, to include the geometry in the aggregation result, we can use the `.dissolve` method. That way, in addition to the summed population we also get the associated geometry per continent, i.e., the union of all countries. Note that we use the `by` parameter to choose which column(s) are used for grouping, and the `aggfunc` parameter to choose the summary function for non-geometry columns:
world_agg2 = world[['continent', 'pop', 'geometry']] \
.dissolve(by='continent', aggfunc='sum')
world_agg2
# Here is a plot of the result:
world_agg2.plot(column='pop');
# The resulting `world_agg2` object is a vector layer containing 8 features representing the continents of the world (and the open ocean).
#
# Other options for the `aggfunc` parameter in `.dissolve` [include](https://geopandas.org/en/stable/docs/user_guide/aggregation_with_dissolve.html):
#
# * `'first'`
# * `'last'`
# * `'min'`
# * `'max'`
# * `'sum'`
# * `'mean'`
# * `'median'`
#
# Additionally, we can pass a custom functiom.
#
# For example, here is how we can calculate the summed population, summed area, and count of countries, per continent. We do this in two steps, then join the results:
world_agg3a = world[['continent', 'area_km2', 'geometry']] \
.dissolve(by='continent', aggfunc='sum')
world_agg3b = world[['continent', 'name_long', 'geometry']] \
.dissolve(by='continent', aggfunc=lambda x: x.nunique()) \
.rename(columns={"name_long": "n"})
world_agg = pd.merge(world_agg3a, world_agg3b, on='continent')
# ...
#
# ### Vector attribute joining
#
# Join by attribute...
coffee_data = pd.read_csv("data/coffee_data.csv")
coffee_data
# Join by `"name_long"` column...
world_coffee = pd.merge(world, coffee_data, on="name_long", how="left")
world_coffee
# Plot...
base = world.plot(color="white", edgecolor="lightgrey")
world_coffee.plot(ax=base, column="coffee_production_2017");
# ### Creating attributes and removing spatial information
#
# Calculate new column...
world2 = world.copy()
world2["pop_dens"] = world2["pop"] / world2["area_km2"]
# Unite columns...
world2["con_reg"] = world["continent"] + ":" + world2["region_un"]
world2 = world2.drop(["continent", "region_un"], axis=1)
# Split column...
world2[["continent", "region_un"]] = world2["con_reg"] \
.str.split(":", expand=True)
# Rename...
world2.rename(columns={"name_long": "name"})
# Renaming all columns...
new_names =["i", "n", "c", "r", "s", "t", "a", "p", "l", "gP", "geom"]
world.columns = new_names
# Dropping geometry...
pd.DataFrame(world.drop(columns="geom"))
# ## Manipulating raster objects
#
# ### Raster subsetting
#
# When using `rasterio`, raster values are accessible through a `numpy` array, which can be imported with the `.read` method:
elev = src_elev.read(1)
elev
# Then, we can access any subset of cell values using `numpy` methods. For example:
elev[0, 0] ## Value at row 1, column 1
# Cell values can be modified by overwriting existing values in conjunction with a subsetting operation. The following expression, for example, sets the upper left cell of elev to 0:
elev[0, 0] = 0
elev
# Multiple cells can also be modified in this way:
elev[0, 0:2] = 0
elev
# ### Summarizing raster objects
#
# Global summaries of raster values can be calculated by applying `numpy` summary functions---such as `np.mean`---on the array with raster values. For example:
np.mean(elev)
# Note that "No Data"-safe functions--such as `np.nanmean`---should be used in case the raster contains "No Data" values which need to be ignored:
elev[0, 2] = np.nan
elev
np.mean(elev)
np.nanmean(elev)
# Raster value statistics can be visualized in a variety of ways. One approach is to "flatten" the raster values into a one-dimensional array, then use a graphical function such as `plt.hist` or `plt.boxplot` (from `matplotlib.pyplot`). For example:
x = elev.flatten()
plt.hist(x);
# ## Exercises
#
| ipynb/03-attribute-operations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# #lec11
with open('jmu_news.txt','r') as jmu_news:
news_content = jmu_news.read()
print(news_content)
from collections import Counter
# +
count_result = Counter(['a','b','b'])
print (count_result.most_common(1))
# -
with open ('jmu_news.txt','r') as jmu_news:
new_content = jmu_news.read()
word_list = news_content.split()
count_result = Counter(word_list)
for word, count in count_result.most_common(10):
print(word,count)
num_list = [1,2,3,4]
new_list = [i+1 for i in num_list]
print(new_list)
# #Ex1
with open ('jmu_news.txt','r') as jmu_news:
new_content = jmu_news.read()
word_list = news_content.split()
count_result = Counter(word_list)
for word, count in count_result.most_common(10):
print(word,count)
with open ('jmu_news.txt','r') as jmu_news:
new_content = jmu_news.read()
word_list = news_content.split()
low_case_list = [word.lower() for word in word_list]
print(low_case_list)
# #EX2
with open ('jmu_news.txt','r') as jmu_news:
new_content = jmu_news.read()
word_list = news_content.split()
low_case_list = [word.lower() for word in word_list]
count_result = Counter(word_list)
for word, count in count_result.most_common(10):
print(word,count)
print("{} salary is${}".format('Tom',60000))
# +
import json
from pprint import pprint
# -
with open('demo.json','r') as json_file:
json_dict = json.load(json_file)
pprint(json_dict)
import urllib.request
# +
url = 'https://www.jmu.edu'
res = urllib.request.urlopen(url)
web_html = res.read()
print(web_html.decode('utf-8'))
# -
| Lec11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Code Challenges!
#
#
# +
# Codewars
# https://www.codewars.com/kata/5467e4d82edf8bbf40000155/train/python
# Descending Orders
# -
def descending_order(num):
# Bust a move right here
num = sorted([char for char in str(num)], reverse = True)
num = num = int("".join(num))
return num
descending_order(5020016)
| Python Code Challenges/.ipynb_checkpoints/Descending Order-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Custom Types
#
# Often, the behavior for a field needs to be customized to support a particular shape or validation method that ParamTools does not support out of the box. In this case, you may use the `register_custom_type` function to add your new `type` to the ParamTools type registry. Each `type` has a corresponding `field` that is used for serialization and deserialization. ParamTools will then use this `field` any time it is handling a `value`, `label`, or `member` that is of this `type`.
#
# ParamTools is built on top of [`marshmallow`](https://github.com/marshmallow-code/marshmallow), a general purpose validation library. This means that you must implement a custom `marshmallow` field to go along with your new type. Please refer to the `marshmallow` [docs](https://marshmallow.readthedocs.io/en/stable/) if you have questions about the use of `marshmallow` in the examples below.
#
#
# ## 32 Bit Integer Example
#
# ParamTools's default integer field uses NumPy's `int64` type. This example shows you how to define an `int32` type and reference it in your `defaults`.
#
# First, let's define the Marshmallow class:
#
# +
import marshmallow as ma
import numpy as np
class Int32(ma.fields.Field):
"""
A custom type for np.int32.
https://numpy.org/devdocs/reference/arrays.dtypes.html
"""
# minor detail that makes this play nice with array_first
np_type = np.int32
def _serialize(self, value, *args, **kwargs):
"""Convert np.int32 to basic, serializable Python int."""
return value.tolist()
def _deserialize(self, value, *args, **kwargs):
"""Cast value from JSON to NumPy Int32."""
converted = np.int32(value)
return converted
# -
# Now, reference it in our defaults JSON/dict object:
#
# +
import paramtools as pt
# add int32 type to the paramtools type registry
pt.register_custom_type("int32", Int32())
class Params(pt.Parameters):
defaults = {
"small_int": {
"title": "Small integer",
"description": "Demonstrate how to define a custom type",
"type": "int32",
"value": 2
}
}
params = Params(array_first=True)
print(f"value: {params.small_int}, type: {type(params.small_int)}")
# -
# One problem with this is that we could run into some deserialization issues. Due to integer overflow, our deserialized result is not the number that we passed in--it's negative!
#
params.adjust(dict(
# this number wasn't chosen randomly.
small_int=2147483647 + 1
))
# ### Marshmallow Validator
#
# Fortunately, you can specify a custom validator with `marshmallow` or ParamTools. Making this works requires modifying the `_deserialize` method to check for overflow like this:
#
class Int32(ma.fields.Field):
"""
A custom type for np.int32.
https://numpy.org/devdocs/reference/arrays.dtypes.html
"""
# minor detail that makes this play nice with array_first
np_type = np.int32
def _serialize(self, value, *args, **kwargs):
"""Convert np.int32 to basic Python int."""
return value.tolist()
def _deserialize(self, value, *args, **kwargs):
"""Cast value from JSON to NumPy Int32."""
converted = np.int32(value)
# check for overflow and let range validator
# display the error message.
if converted != int(value):
return int(value)
return converted
# Now, let's see how to use `marshmallow` to fix this problem:
#
# +
import marshmallow as ma
import paramtools as pt
# get the minimum and maxium values for 32 bit integers.
min_int32 = -2147483648 # = np.iinfo(np.int32).min
max_int32 = 2147483647 # = np.iinfo(np.int32).max
# add int32 type to the paramtools type registry
pt.register_custom_type(
"int32",
Int32(validate=[
ma.validate.Range(min=min_int32, max=max_int32)
])
)
class Params(pt.Parameters):
defaults = {
"small_int": {
"title": "Small integer",
"description": "Demonstrate how to define a custom type",
"type": "int32",
"value": 2
}
}
params = Params(array_first=True)
params.adjust(dict(
small_int=np.int64(max_int32) + 1
))
# -
# ### ParamTools Validator
#
# Finally, we will use ParamTools to solve this problem. We need to modify how we create our custom `marshmallow` field so that it's wrapped by ParamTools's `PartialField`. This makes it clear that your field still needs to be initialized, and that your custom field is able to receive validation information from the `defaults` configuration:
#
# +
import paramtools as pt
# add int32 type to the paramtools type registry
pt.register_custom_type(
"int32",
pt.PartialField(Int32)
)
class Params(pt.Parameters):
defaults = {
"small_int": {
"title": "Small integer",
"description": "Demonstrate how to define a custom type",
"type": "int32",
"value": 2,
"validators": {
"range": {"min": -2147483648, "max": 2147483647}
}
}
}
params = Params(array_first=True)
params.adjust(dict(
small_int=2147483647 + 1
))
# -
| docs/api/custom-types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: gpu
# language: python
# name: gpu
# ---
# +
import pandas as pd
import re
import folium
import os
from folium.plugins import HeatMap
# -
excel = pd.read_excel('data.xlsx')
# +
df = excel
clean_frame = pd.DataFrame(columns=['Tag', 'Latitude', 'Longitude'])
i = 0
for index, row in df.iterrows():
tags = re.findall(r"#(\w+)", row['Tweet content'])
latidute = row['Latitude']
longitude = row['Longitude']
for tag in tags:
clean_frame.loc[i] = [tag, latidute, longitude]
i += 1
# -
clean_frame = clean_frame[['Latitude', 'Longitude', 'Tag']]
top_tags = clean_frame['Tag'].value_counts()
clean_frame['Tag'] = pd.factorize(clean_frame.Tag)[0]
top_tags_factorized = clean_frame['Tag'].value_counts()
print(top_tags_factorized.head())
# +
m = folium.Map([48., 5.], zoom_start=6)
colors = ['red', 'blue', 'lime']
top_hash_indexes = top_tags_factorized.head(3).index.tolist()
for i, k in enumerate(top_hash_indexes):
data = clean_frame.loc[clean_frame['Tag'] == k].values.tolist()
HeatMap(data, radius=15, gradient={0: 'white', 1: colors[i]}).add_to(m)
m.save('Heatmap.html')
m
# +
import numpy as np
data = (np.random.normal(size=(20, 3)) *
np.array([[1, 1, 1]]) +
np.array([[48, 5, 1]])).tolist()
data2 = (np.random.normal(size=(20, 3)) *
np.array([[1, 1, 1]]) +
np.array([[48, 20, 1]])).tolist()
from folium.plugins import HeatMap
m = folium.Map([48., 5.], zoom_start=6)
HeatMap(data, 'first', radius=15, gradient={0: 'white', 1: 'red'}).add_to(m)
HeatMap(data2, 'second', radius=15, gradient={0: 'white', 1: 'blue'}).add_to(m)
m.save(os.path.join('Heatmap.html'))
m
# -
| lab1/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: simclr
# language: python
# name: simclr
# ---
# +
import os
import cv2
import numpy as np
import pandas as pd
from tqdm import tqdm
import torch
import torch.nn as nn
from torch.nn import functional as F
import transformers
import gc
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
# -
# ## TfidfVectorizer model
def read_dataset(is_train=True):
if is_train:
df = pd.read_csv('train.csv')
image_paths = 'train_images/' + df['image']
else:
df = pd.read_csv('test.csv')
image_paths = 'test_images/' + df['image']
return df, image_paths
def combine_predictions(row):
x = np.concatenate([ row['text_predictions'], row['phash']])
return ' '.join( np.unique(x) )
def get_text_predictions_torch(df, max_features=25_000, th=0.75):
model = TfidfVectorizer(stop_words='english', binary=True, max_features=max_features)
text_embeddings = model.fit_transform(df['title']).toarray().astype(np.float16)
text_embeddings=torch.from_numpy(text_embeddings).to('cuda:0')
preds = []
CHUNK = 1024*4
print('Finding similar titles...')
CTS = len(df) // CHUNK
if (len(df)%CHUNK) != 0: CTS += 1
for j in tqdm(range( CTS )):
a = j * CHUNK
b = (j+1) * CHUNK
b = min(b, len(df))
#print('chunk',a,'to',b)
# COSINE SIMILARITY DISTANCE
cts = torch.matmul(text_embeddings, text_embeddings[a:b].T).T
for k in range(b-a):
IDX = torch.where(cts[k,] > th)[0].cpu().numpy()
o = df.iloc[IDX].posting_id.values
preds.append(o)
del model, text_embeddings
gc.collect()
torch.cuda.empty_cache()
return preds
df,image_paths = read_dataset()
df.head()
text_predictions = get_text_predictions_torch(df, max_features=25_000)
# ### phash
phash = df.groupby('image_phash').posting_id.agg('unique').to_dict()
df['phash'] = df.image_phash.map(phash)
df.head()
# ### TfidfVectorizer + phash
df['text_predictions'] = text_predictions
df['matches'] = df.apply(combine_predictions, axis=1)
df[['posting_id', 'matches']].to_csv('submission.csv', index=False)
# LB: 0.652
# +
def getMetric(col):
def f1score(row):
n = len(np.intersect1d(row.target, row[col]))
return 2*n / (len(row.target) + len(row[col]))
return f1score
def combine_for_cv(row):
x = np.concatenate([row['phash'], row['text_predictions']])
return np.unique(x)
df['text_predictions'] = text_predictions
phash = df.groupby('image_phash').posting_id.agg('unique').to_dict()
df['phash'] = df.image_phash.map(phash)
df['matches_CV'] = df.apply(combine_for_cv, axis=1)
tmp = df.groupby('label_group').posting_id.agg('unique').to_dict()
df['target'] = df.label_group.map(tmp)
MyCVScore = df.apply(getMetric('matches_CV'), axis=1)
print('CV score =', MyCVScore.mean())
# -
# ## Transformer
class CFG:
batch_size = 16
seed = 42
device = 'cuda'
classes = 11014
scale = 30
margin = 0.5
CV = False
num_workers=4
transformer_model = 'sentence-transformer-models/paraphrase-xlm-r-multilingual-v1/0_Transformer'
text_model_path = 'best-multilingual-model/sentence_transfomer_xlm_best_loss_num_epochs_25_arcface.bin'
model_params = {
'n_classes':11014,
'model_name':transformer_model,
'use_fc':False,
'fc_dim':512,
'dropout':0.3,
}
tokenizer = transformers.AutoTokenizer.from_pretrained(CFG.transformer_model)
# +
class ShopeeTextDataset(Dataset):
def __init__(self, csv):
self.csv = csv.reset_index()
def __len__(self):
return self.csv.shape[0]
def __getitem__(self, index):
row = self.csv.iloc[index]
text = row.title
text = tokenizer(text, padding='max_length', truncation=True, max_length=128, return_tensors="pt")
input_ids = text['input_ids'][0]
attention_mask = text['attention_mask'][0]
return input_ids, attention_mask
class ShopeeTextNet(nn.Module):
def __init__(self,
n_classes,
model_name='bert-base-uncased',
use_fc=False,
fc_dim=512,
dropout=0.0):
"""
:param n_classes:
:param model_name: name of model from pretrainedmodels
e.g. resnet50, resnext101_32x4d, pnasnet5large
:param pooling: One of ('SPoC', 'MAC', 'RMAC', 'GeM', 'Rpool', 'Flatten', 'CompactBilinearPooling')
:param loss_module: One of ('arcface', 'cosface', 'softmax')
"""
super(ShopeeTextNet, self).__init__()
self.transformer = transformers.AutoModel.from_pretrained(model_name)
final_in_features = self.transformer.config.hidden_size
self.use_fc = use_fc
if use_fc:
self.dropout = nn.Dropout(p=dropout)
self.fc = nn.Linear(final_in_features, fc_dim)
self.bn = nn.BatchNorm1d(fc_dim)
self._init_params()
final_in_features = fc_dim
def _init_params(self):
nn.init.xavier_normal_(self.fc.weight)
nn.init.constant_(self.fc.bias, 0)
nn.init.constant_(self.bn.weight, 1)
nn.init.constant_(self.bn.bias, 0)
def forward(self, input_ids,attention_mask):
feature = self.extract_feat(input_ids,attention_mask)
return F.normalize(feature)
def extract_feat(self, input_ids,attention_mask):
x = self.transformer(input_ids=input_ids,attention_mask=attention_mask)
features = x[0]
features = features[:,0,:]
if self.use_fc:
features = self.dropout(features)
features = self.fc(features)
features = self.bn(features)
return features
# -
def get_text_embeddings(df):
embeds = []
model = ShopeeTextNet(**CFG.model_params)
model.eval()
model.load_state_dict(dict(list(torch.load(CFG.TEXT_MODEL_PATH).items())[:-1]))
model = model.to(CFG.device)
text_dataset = ShopeeTextDataset(df)
text_loader = torch.utils.data.DataLoader(
text_dataset,
batch_size=CFG.batch_size,
pin_memory=True,
drop_last=False,
num_workers=CFG.num_workers
)
with torch.no_grad():
for input_ids, attention_mask in tqdm(text_loader):
input_ids = input_ids.cuda()
attention_mask = attention_mask.cuda()
feat = model(input_ids, attention_mask)
text_embeddings = feat.detach().cpu().numpy()
embeds.append(text_embeddings)
del model
text_embeddings = np.concatenate(embeds)
print(f'Our text embeddings shape is {text_embeddings.shape}')
del embeds
gc.collect()
return text_embeddings
def f1_score(y_true, y_pred):
y_true = y_true.apply(lambda x: set(x.split()))
y_pred = y_pred.apply(lambda x: set(x.split()))
intersection = np.array([len(x[0] & x[1]) for x in zip(y_true, y_pred)])
len_y_pred = y_pred.apply(lambda x: len(x)).values
len_y_true = y_true.apply(lambda x: len(x)).values
f1 = 2 * intersection / (len_y_pred + len_y_true)
return f1
def get_neighbours_cos_sim(df,embeddings, threshold=0.6):
'''
When using cos_sim use normalized features else use normal features
'''
embeddings = cupy.array(embeddings)
if CFG.GET_CV:
thresholds = list(np.arange(0.5,0.7,0.05))
scores = []
for threshold in thresholds:
preds = []
CHUNK = 1024*4
print('Finding similar titles...for threshold :',threshold)
CTS = len(embeddings)//CHUNK
if len(embeddings)%CHUNK!=0: CTS += 1
for j in range( CTS ):
a = j*CHUNK
b = (j+1)*CHUNK
b = min(b,len(embeddings))
cts = cupy.matmul(embeddings,embeddings[a:b].T).T
for k in range(b-a):
IDX = cupy.where(cts[k,]>threshold)[0]
o = df.iloc[cupy.asnumpy(IDX)].posting_id.values
o = ' '.join(o)
preds.append(o)
df['pred_matches'] = preds
df['f1'] = f1_score(df['matches'], df['pred_matches'])
score = df['f1'].mean()
print(f'Our f1 score for threshold {threshold} is {score}')
scores.append(score)
thresholds_scores = pd.DataFrame({'thresholds': thresholds, 'scores': scores})
max_score = thresholds_scores[thresholds_scores['scores'] == thresholds_scores['scores'].max()]
best_threshold = max_score['thresholds'].values[0]
best_score = max_score['scores'].values[0]
print(f'Our best score is {best_score} and has a threshold {best_threshold}')
else:
preds = []
CHUNK = 1024*4
print('Finding similar texts...for threshold :',threshold)
CTS = len(embeddings)//CHUNK
if len(embeddings)%CHUNK!=0: CTS += 1
for j in range( CTS ):
a = j*CHUNK
b = (j+1)*CHUNK
b = min(b,len(embeddings))
print('chunk',a,'to',b)
cts = cupy.matmul(embeddings,embeddings[a:b].T).T
for k in range(b-a):
IDX = cupy.where(cts[k,]>threshold)[0]
o = df.iloc[cupy.asnumpy(IDX)].posting_id.values
preds.append(o)
return df, preds
df,df_cu,image_paths = read_dataset()
df.head()
text_embeddings = get_text_embeddings(df)
df, text_predictions = get_neighbours_cos_sim(df, text_embeddings)
# ### CV Score for transformer
df['text_predictions'] = text_predictions
phash = df.groupby('image_phash').posting_id.agg('unique').to_dict()
df['phash'] = df.image_phash.map(phash)
df['matches_CV'] = df.apply(combine_for_cv, axis=1)
tmp = df.groupby('label_group').posting_id.agg('unique').to_dict()
df['target'] = df.label_group.map(tmp)
MyCVScore = df.apply(getMetric('matches_CV'), axis=1)
print('CV score =', MyCVScore.mean())
| Text.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# # Part 1: Data Ingestion
#
# This demo showcases financial fraud prevention and using the MLRun feature store to define complex features that help identify fraud. Fraud prevention specifically is a challenge as it requires processing raw transaction and events in real-time and being able to quickly respond and block transactions before they occur.
#
# To address this, we create a development pipeline and a production pipeline. Both pipelines share the same feature engineering and model code, but serve data very differently. Furthermore, we automate the data and model monitoring process, identify drift and trigger retraining in a CI/CD pipeline. This process is described in the diagram below:
#
# 
# The raw data is described as follows:
#
# | TRANSACTIONS || ║ |USER EVENTS ||
# |-----------------|----------------------------------------------------------------|----------|-----------------|----------------------------------------------------------------|
# | **age** | age group value 0-6. Some values are marked as U for unknown | ║ | **source** | The party/entity related to the event |
# | **gender** | A character to define the age | ║ | **event** | event, such as login or password change |
# | **zipcodeOri** | ZIP code of the person originating the transaction | ║ | **timestamp** | The date and time of the event |
# | **zipMerchant** | ZIP code of the merchant receiving the transaction | ║ | | |
# | **category** | category of the transaction (e.g., transportation, food, etc.) | ║ | | |
# | **amount** | the total amount of the transaction | ║ | | |
# | **fraud** | whether the transaction is fraudulent | ║ | | |
# | **timestamp** | the date and time in which the transaction took place | ║ | | |
# | **source** | the ID of the party/entity performing the transaction | ║ | | |
# | **target** | the ID of the party/entity receiving the transaction | ║ | | |
# | **device** | the device ID used to perform the transaction | ║ | | |
# This notebook introduces how to **Ingest** different data sources to the **Feature Store**.
#
# The following FeatureSets will be created:
# - **Transactions**: Monetary transactions between a source and a target.
# - **Events**: Account events such as account login or a password change.
# - **Label**: Fraud label for the data.
#
# By the end of this tutorial you’ll learn how to:
#
# - Create an ingestion pipeline for each data source.
# - Define preprocessing, aggregation and validation of the pipeline.
# - Run the pipeline locally within the notebook.
# - Launch a real-time function to ingest live data.
# - Schedule a cron to run the task when needed.
project_name = 'fraud-demo'
# +
import mlrun
# Initialize the MLRun project object
project = mlrun.get_or_create_project(project_name, context="./", user_project=True)
# -
# ## Step 1 - Fetch, Process and Ingest our datasets
# ## 1.1 - Transactions
# ### Transactions
# + tags=["hide-cell"]
# Helper functions to adjust the timestamps of our data
# while keeping the order of the selected events and
# the relative distance from one event to the other
def date_adjustment(sample, data_max, new_max, old_data_period, new_data_period):
'''
Adjust a specific sample's date according to the original and new time periods
'''
sample_dates_scale = ((data_max - sample) / old_data_period)
sample_delta = new_data_period * sample_dates_scale
new_sample_ts = new_max - sample_delta
return new_sample_ts
def adjust_data_timespan(dataframe, timestamp_col='timestamp', new_period='2d', new_max_date_str='now'):
'''
Adjust the dataframe timestamps to the new time period
'''
# Calculate old time period
data_min = dataframe.timestamp.min()
data_max = dataframe.timestamp.max()
old_data_period = data_max-data_min
# Set new time period
new_time_period = pd.Timedelta(new_period)
new_max = pd.Timestamp(new_max_date_str)
new_min = new_max-new_time_period
new_data_period = new_max-new_min
# Apply the timestamp change
df = dataframe.copy()
df[timestamp_col] = df[timestamp_col].apply(lambda x: date_adjustment(x, data_max, new_max, old_data_period, new_data_period))
return df
# +
import pandas as pd
# Fetch the transactions dataset from the server
transactions_data = pd.read_csv('https://s3.wasabisys.com/iguazio/data/fraud-demo-mlrun-fs-docs/data.csv', parse_dates=['timestamp'], nrows=500)
# Adjust the samples timestamp for the past 2 days
transactions_data = adjust_data_timespan(transactions_data, new_period='2d')
# Preview
transactions_data.head(3)
# -
# ### Transactions - Create a FeatureSet and Preprocessing Pipeline
# Create the FeatureSet (data pipeline) definition for the **credit transaction processing** which describes the offline/online data transformations and aggregations.<br>
# The feature store will automatically add an offline `parquet` target and an online `NoSQL` target by using `set_targets()`.
#
# The data pipeline consists of:
#
# * **Extracting** the data components (hour, day of week)
# * **Mapping** the age values
# * **One hot encoding** for the transaction category and the gender
# * **Aggregating** the amount (avg, sum, count, max over 2/12/24 hour time windows)
# * **Aggregating** the transactions per category (over 14 days time windows)
# * **Writing** the results to **offline** (Parquet) and **online** (NoSQL) targets
# Import MLRun's Feature Store
import mlrun.feature_store as fstore
from mlrun.feature_store.steps import OneHotEncoder, MapValues, DateExtractor
# Define the transactions FeatureSet
transaction_set = fstore.FeatureSet("transactions",
entities=[fstore.Entity("source")],
timestamp_key='timestamp',
description="transactions feature set")
# +
# Define and add value mapping
main_categories = ["es_transportation", "es_health", "es_otherservices",
"es_food", "es_hotelservices", "es_barsandrestaurants",
"es_tech", "es_sportsandtoys", "es_wellnessandbeauty",
"es_hyper", "es_fashion", "es_home", "es_contents",
"es_travel", "es_leisure"]
# One Hot Encode the newly defined mappings
one_hot_encoder_mapping = {'category': main_categories,
'gender': list(transactions_data.gender.unique())}
# Define the graph steps
transaction_set.graph\
.to(DateExtractor(parts = ['hour', 'day_of_week'], timestamp_col = 'timestamp'))\
.to(MapValues(mapping={'age': {'U': '0'}}, with_original_features=True))\
.to(OneHotEncoder(mapping=one_hot_encoder_mapping))
# Add aggregations for 2, 12, and 24 hour time windows
transaction_set.add_aggregation(name='amount',
column='amount',
operations=['avg','sum', 'count','max'],
windows=['2h', '12h', '24h'],
period='1h')
# Add the category aggregations over a 14 day window
for category in main_categories:
transaction_set.add_aggregation(name=category,column=f'category_{category}',
operations=['count'], windows=['14d'], period='1d')
# Add default (offline-parquet & online-nosql) targets
transaction_set.set_targets()
# Plot the pipeline so we can see the different steps
transaction_set.plot(rankdir="LR", with_targets=True)
# -
# ### Transactions - Ingestion
# +
# Ingest our transactions dataset through our defined pipeline
transactions_df = fstore.ingest(transaction_set, transactions_data,
infer_options=fstore.InferOptions.default())
transactions_df.head(3)
# -
# ## 1.2 - User Events
# ### User Events - Fetching
# +
# Fetch our user_events dataset from the server
user_events_data = pd.read_csv('https://s3.wasabisys.com/iguazio/data/fraud-demo-mlrun-fs-docs/events.csv',
index_col=0, quotechar="\'", parse_dates=['timestamp'], nrows=500)
# Adjust to the last 2 days to see the latest aggregations in our online feature vectors
user_events_data = adjust_data_timespan(user_events_data, new_period='2d')
# Preview
user_events_data.head(3)
# -
# ### User Events - Create a FeatureSet and Preprocessing Pipeline
#
# Now we will define the events feature set.
# This is a pretty straight forward pipeline in which we only one hot encode the event categories and save the data to the default targets.
user_events_set = fstore.FeatureSet("events",
entities=[fstore.Entity("source")],
timestamp_key='timestamp',
description="user events feature set")
# +
# Define and add value mapping
events_mapping = {'event': list(user_events_data.event.unique())}
# One Hot Encode
user_events_set.graph.to(OneHotEncoder(mapping=events_mapping))
# Add default (offline-parquet & online-nosql) targets
user_events_set.set_targets()
# Plot the pipeline so we can see the different steps
user_events_set.plot(rankdir="LR", with_targets=True)
# -
# ### User Events - Ingestion
# Ingestion of our newly created events feature set
events_df = fstore.ingest(user_events_set, user_events_data)
events_df.head(3)
# ## Step 2 - Create a labels dataset for model training
# ### Label Set - Create a FeatureSet
# This feature set contains the label for the fraud demo, it will be ingested directly to the default targets without any changes
def create_labels(df):
labels = df[['fraud','source','timestamp']].copy()
labels = labels.rename(columns={"fraud": "label"})
labels['timestamp'] = labels['timestamp'].astype("datetime64[ms]")
labels['label'] = labels['label'].astype(int)
labels.set_index('source', inplace=True)
return labels
# +
# Define the "labels" feature set
labels_set = fstore.FeatureSet("labels",
entities=[fstore.Entity("source")],
timestamp_key='timestamp',
description="training labels",
engine="pandas")
labels_set.graph.to(name="create_labels", handler=create_labels)
# specify only Parquet (offline) target since its not used for real-time
labels_set.set_targets(['parquet'], with_defaults=False)
labels_set.plot(with_targets=True)
# -
# ### Label Set - Ingestion
# Ingest the labels feature set
labels_df = fstore.ingest(labels_set, transactions_data)
labels_df.head(3)
# ## Step 3 - Deploy a real-time pipeline
#
# When dealing with real-time aggregation, it's important to be able to update these aggregations in real-time.
# For this purpose, we will create live serving functions that will update the online feature store of the `transactions` FeatureSet and `Events` FeatureSet.
#
# Using MLRun's `serving` runtime, craetes a nuclio function loaded with our feature set's computational graph definition
# and an `HttpSource` to define the HTTP trigger.
#
# Notice that the implementation below does not require any rewrite of the pipeline logic.
# ## 3.1 - Transactions
# ### Transactions - Deploy our FeatureSet live endpoint
# Create iguazio v3io stream and transactions push API endpoint
transaction_stream = f'v3io:///projects/{project.name}/streams/transaction'
transaction_pusher = mlrun.datastore.get_stream_pusher(transaction_stream)
# +
# Define the source stream trigger (use v3io streams)
# we will define the `key` and `time` fields (extracted from the Json message).
source = mlrun.datastore.sources.StreamSource(path=transaction_stream , key_field='source', time_field='timestamp')
# Deploy the transactions feature set's ingestion service over a real-time (Nuclio) serverless function
# you can use the run_config parameter to pass function/service specific configuration
transaction_set_endpoint = fstore.deploy_ingestion_service(featureset=transaction_set, source=source)
# -
# ### Transactions - Test the feature set HTTP endpoint
# By defining our `transactions` feature set we can now use MLRun and Storey to deploy it as a live endpoint, ready to ingest new data!
#
# Using MLRun's `serving` runtime, we will create a nuclio function loaded with our feature set's computational graph definition and an `HttpSource` to define the HTTP trigger.
# +
import requests
import json
# Select a sample from the dataset and serialize it to JSON
transaction_sample = json.loads(transactions_data.sample(1).to_json(orient='records'))[0]
transaction_sample['timestamp'] = str(pd.Timestamp.now())
transaction_sample
# -
# Post the sample to the ingestion endpoint
requests.post(transaction_set_endpoint, json=transaction_sample).text
# ## 3.2 - User Events
# ### User Events - Deploy our FeatureSet live endpoint
# Deploy the events feature set's ingestion service using the feature set and all the previously defined resources.
# Create iguazio v3io stream and transactions push API endpoint
events_stream = f'v3io:///projects/{project.name}/streams/events'
events_pusher = mlrun.datastore.get_stream_pusher(events_stream)
# +
# Define the source stream trigger (use v3io streams)
# we will define the `key` and `time` fields (extracted from the Json message).
source = mlrun.datastore.sources.StreamSource(path=events_stream , key_field='source', time_field='timestamp')
# Deploy the transactions feature set's ingestion service over a real-time (Nuclio) serverless function
# you can use the run_config parameter to pass function/service specific configuration
events_set_endpoint = fstore.deploy_ingestion_service(featureset=user_events_set, source=source)
# -
# ### User Events - Test the feature set HTTP endpoint
# Select a sample from the events dataset and serialize it to JSON
user_events_sample = json.loads(user_events_data.sample(1).to_json(orient='records'))[0]
user_events_sample['timestamp'] = str(pd.Timestamp.now())
user_events_sample
# Post the sample to the ingestion endpoint
requests.post(events_set_endpoint, json=user_events_sample).text
# ## Done!
#
# You've completed Part 1 of the data-ingestion with the feature store.
# Proceed to [Part 2](02-create-training-model.ipynb) to learn how to train an ML model using the feature store data.
| docs/feature-store/end-to-end-demo/01-ingest-datasources.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import re
from operator import itemgetter
Corpus = {
'l o w _':5,
'l o w e r _':2,
'n e w e s t _':6,
'w i d e s t _':3,
'h a p p i e r _':2
}
Corpus
def getPairCounts(Corpus):
pairs = {}
for word,fr in Corpus.items():
symbols = word.split(' ')
for i in range(len(symbols)-1):
pair = (symbols[i],symbols[i+1])
cfr = pairs.get(pair,0)
pairs[pair] = cfr+fr
return pairs
pairsCounts = getPairCounts(Corpus)
pairsCounts
def getBestPair(pairsCounts):
return max(pairsCounts,key=pairsCounts.get)
print(getBestPair(pairsCounts))
def mergeInCorpus(bestPair,Corpus):
newCorpus = {}
for word in Corpus:
newWord = re.sub(' '.join(bestPair),''.join(bestPair),word)
newCorpus[newWord] = Corpus[word]
return newCorpus
bestPair = getBestPair(pairsCounts)
newCorpus = mergeInCorpus(bestPair,Corpus)
newCorpus
def runBPE(Corpus,k):
bpeStats = {}
for i in range(k):
pairsCounts = getPairCounts(Corpus)
if not pairsCounts:
break
bestPair = getBestPair(pairsCounts)
bpeStats[bestPair] = i
Corpus = mergeInCorpus(bestPair,Corpus)
return Corpus,bpeStats
Corpus = {
'l o w _':5,
'l o w e r _':2,
'n e w e s t _':6,
'w i d e s t _':3,
'h a p p i e r _':2
}
newCorpus,bpeStats = runBPE(Corpus,10)
newCorpus
bpeStats
newWord = 'lowest'
newWord2 = ' '.join(list(newWord))+' _'
def getAllPairs(word):
pairs = []
word = word.split(' ')
prevChar = word[0]
for char in word[1:]:
pairs.append((prevChar,char))
prevChar = char
return pairs
pairs = getAllPairs(newWord2)
pairs
def getPairToBeMerged(bpeStats,pairs):
#bpeCodes = [(pair,bpeStats[pair]) for pair in pairs if pair in bpeStats]
bpeCodes = []
for pair in pairs:
if pair in bpeStats:
bpeCodes.append((pair,bpeStats[pair]))
if len(bpeCodes) == 0:
return (-1,-1)
pairToBeMerged = min(bpeCodes,key=itemgetter(1))[0]
return pairToBeMerged
pairToBeMerged = getPairToBeMerged(bpeStats,pairs)
def mergeLetters(word,pairToBeMerged):
newWord = re.sub(' '.join(pairToBeMerged),''.join(pairToBeMerged),word)
return newWord
print(mergeLetters(newWord2,pairToBeMerged))
def bpeTokenize(word,bpeStats):
if len(word) == 1:
return word
word = ' '.join(list(word))+' _'
while True:
pairs = getAllPairs(word)
pairToBeMerged = getPairToBeMerged(bpeStats,pairs)
if pairToBeMerged[0] == -1:
break
word = mergeLetters(word,pairToBeMerged)
return word
newWord = bpeTokenize('lowest',bpeStats)
newWord
| 01-Code/2b_TextProcessing_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import tensorflow as tf
from sklearn.decomposition import PCA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
from sklearn.cluster import KMeans
# +
fashion_mnist = tf.keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# -
X = np.concatenate([train_images,test_images])
Y = np.concatenate([train_labels,test_labels])
X = X.reshape(X.shape[0], 28*28)
pca = PCA(n_components=20).fit(X)
pca2 = PCA(n_components=2).fit(X)
pca3 = PCA(n_components=3).fit(X)
X2 = pca2.transform(X)
X3 = pca3.transform(X)
kmeans_per_k = [KMeans(n_clusters=k, random_state=42, init='random').fit(X2) for k in [4, 7, 10]]
for model in kmeans_per_k:
print(model.cluster_centers_)
x = X2[:300]
labels = [model.predict(x) for model in kmeans_per_k]
fig, ax = plt.subplots()
x = X2[:300,0]
y = X2[:300,1]
ax.scatter(x, y,marker="")
for i, txt in enumerate(labels[0]):
ax.annotate(txt, (x[i], y[i]))
fig, ax = plt.subplots()
x = X2[:300,0]
y = X2[:300,1]
ax.scatter(x, y,marker="")
for i, txt in enumerate(labels[1]):
ax.annotate(txt, (x[i], y[i]))
fig, ax = plt.subplots()
x = X2[:300,0]
y = X2[:300,1]
ax.scatter(x, y,marker="")
for i, txt in enumerate(labels[2]):
ax.annotate(txt, (x[i], y[i]))
pca_s.
| 7-c.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# <a href="https://hub.callysto.ca/jupyter/hub/user-redirect/git-pull?repo=https%3A%2F%2Fgithub.com%2Fcallysto%2Fcurriculum-notebooks&branch=master&subPath=Mathematics/Percentage/Percents.ipynb&depth=1" target="_parent"><img src="https://raw.githubusercontent.com/callysto/curriculum-notebooks/master/open-in-callysto-button.svg?sanitize=true" width="123" height="24" alt="Open in Callysto"/></a>
# **Run the cell below, this will add two buttons. Click on the "initialize" button before proceeding through the notebook**
# + tags=["hide-input"]
import uiButtons
# %uiButtons
# + tags=["hide-input"] language="html"
# <script src="https://d3js.org/d3.v3.min.js"></script>
# -
# # Percentages
# ## Introduction
# In this notebook we will discuss what percentages are and why this way of representing data is helpful in many different contexts. Common examples of percentages are sales tax or a mark for an assignment.
#
# The word percent comes from the Latin adverbial phrase *per centum* meaning “*by the hundred*”.
#
# For example, if the sales tax is $5\%$, this means that for every dollar you spend the tax adds $5$ cents to the total price of the purchase.
#
# A percentage simply represents a fraction (per hundred). For example, $90\%$ is the same as saying $\dfrac{90}{100}$. It is used to represent a ratio.
#
# What makes percentages so powerful is that they can represent any ratio.
#
# For example, getting $\dfrac{22}{25}$ on a math exam can be represented as $88\%$: $22$ is $88\%$ of $25$.
# ## How to Get a Percentage
# As mentioned in the introduction, a percentage is simply a fraction represented as a portion of 100.
#
# For this notebook we will only talk about percentages between 0% and 100%.
#
# This means the corresponding fraction will always be a value between $0$ and $1$.
#
# Let's look at our math exam mark example from above. The student correctly answered $22$ questions out of $25$, so the student received a grade of $\dfrac{22}{25}$.
#
# To represent this ratio as a percentage we first convert $\dfrac{22}{25}$ to its decimal representation (simply do the division in your calculator).
#
# $$
# \dfrac{22}{25} = 22 \div 25 = 0.88
# $$
#
# We are almost done: we now have the ratio represented as a value between 0 and 1. To finish getting the answer to our problem all we need to do is multiply this value by $100$ to get our percentage. $$0.88 \times 100 = 88\%$$
#
# Putting it all together we can say $22$ is $88\%$ of $25$.
#
# Think of a grade you recently received (as a fraction) and convert it to a percentage. Once you think you have an answer you can use the widget below to check your answer.
#
# Simply add the total marks of the test/assignment then move the slider until you get to your grade received.
# + tags=["hide-input"] language="html"
# <style>
# .main {
# font-family: 'Segoe UI', Tahoma, Geneva, Verdana, sans-serif;
# }
#
# .slider {
# width: 100px;
# }
#
# #maxVal {
# border:1px solid #cccccc;
# border-radius: 5px;
# width: 50px;
# }
# </style>
# <div class="main" style="border:2px solid black; width: 400px; padding: 20px;border-radius: 10px; margin: 0 auto; box-shadow: 3px 3px 12px #acacac">
# <div>
# <label for="maxValue">Enter the assignment/exam total marks</label>
# <input type="number" id="maxVal" value="100">
# </div>
# <div>
# <input type="range" min="0" max="100" value="0" class="slider" id="mySlider" style="width: 300px; margin-top: 20px;">
# </div>
# <h4 id="sliderVal">0</h3>
# </div>
#
# <script>
# var slider = document.getElementById('mySlider');
# var sliderVal = document.getElementById('sliderVal');
#
# slider.oninput = function () {
# var sliderMax = document.getElementById('maxVal').value;
# if(sliderMax < 0 || isNaN(sliderMax)) {
# sliderMax = 100;
# document.getElementById('maxVal').value = 100;
# }
# d3.select('#mySlider').attr('max', sliderMax);
# sliderVal.textContent = "If you answered " + this.value + "/" + sliderMax + " correct questions your grade will be " + ((
# this.value / sliderMax) * 100).toPrecision(3) + "%";
# }
# </script>
# -
# ## Solving Problems Using Percentages
#
# Now that we understand what percentages mean and how to get them from fractions, let's look at solving problems using percentages. Start by watching the video below to get a basic understanding.
# + tags=["hide-input"] language="html"
# <div align="middle">
# <iframe id="percentVid" width="640" height="360" src="https://www.youtube.com/embed/rR95Cbcjzus?end=368" frameborder="0" allow="autoplay; encrypted-media" allowfullscreen style="box-shadow: 3px 3px 12px #ACACAC">
# </iframe>
# <p><a href="https://www.youtube.com/channel/UC4a-Gbdw7vOaccHmFo40b9g" target="_blank">Click here</a> for more videos by Math Antics</p>
# </div>
# <script>
# $(function() {
# var reachable = false;
# var myFrame = $('#percentVid');
# var videoSrc = myFrame.attr("src");
# myFrame.attr("src", videoSrc)
# .on('load', function(){reachable = true;});
# setTimeout(function() {
# if(!reachable) {
# var ifrm = myFrame[0];
# ifrm = (ifrm.contentWindow) ? ifrm.contentWindow : (ifrm.contentDocument.document) ? ifrm.contentDocument.document : ifrm.contentDocument;
# ifrm.document.open();
# ifrm.document.write('If the video does not start click <a href="' + videoSrc + '" target="_blank">here</a>');
# ifrm.document.close();
# }
# }, 2000)
# });
# </script>
# -
# As shown in the video, taking $25\%$ of 20 "things" is the same as saying $\dfrac{25}{100}\times\dfrac{20}{1}=\dfrac{500}{100}=\dfrac{5}{1}=5$.
#
# Let's do another example, assume a retail store is having a weekend sale. The sale is $30\%$ off everything in store.
#
# Sam thinks this is a great time to buy new shoes, and the shoes she is interested in are regular price $\$89.99$.<br>
# If Sam buys these shoes this weekend how much will they cost? If the sales tax is $5\%$, what will the total price be?
#
# <img src="https://orig00.deviantart.net/5c3e/f/2016/211/b/d/converse_shoes_free_vector_by_superawesomevectors-dabxj2k.jpg" width="300">
# <img src="https://www.publicdomainpictures.net/pictures/170000/nahled/30-korting.jpg" width="300">
#
# Let's start by figuring out the sale price of the shoes before calculating the tax. To figure out the new price we must first take $30\%$ off the original price.
#
# So the shoes are regular priced at $\$89.99$ and the sale is for $30\%$ off
#
# $$
# \$89.99\times 30\%=\$89.99\times\frac{30}{100}=\$26.997
# $$
#
# We can round $\$26.997$ to $\$27$.
#
# Ok we now know how much Sam will save on her new shoes, but let's not forget that the question is asking how much her new shoes will cost, not how much she will save. All we need to do now is take the total price minus the savings to get the new price:
#
# $$
# \$89.99- \$27=\$62.99
# $$
#
# Wow, what savings!
#
# Now for the second part of the question: what will the total price be if the tax is $5\%$?
#
# We must now figure out what $5\%$ of $\$62.99$ is
#
# $$
# \$62.99\times5\%=\$62.99\times\frac{5}{100}=\$3.15
# $$
#
# Now we know that Sam will need to pay $\$3.15$ of tax on her new shoes so the final price is
#
# $$
# \$62.99+\$3.15=\$66.14
# $$
#
# A shortcut for finding the total price including the sales tax is to add 1 to the tax ratio, let's see how this works:
#
# $$
# \$62.99\times\left(\frac{5}{100}+1\right)=\$62.99\times1.05=\$66.14
# $$
#
# You can use this trick to quickly figure out a price after tax.
# ## Multiplying Percentages together
# Multiplying two or more percentages together is probably not something you would encounter often but it is easy to do if you remember that percentages are really fractions.
#
# Since percentages is simply a different way to represent a fraction, the rules for multiplying them are the same. Recall that multiplying two fractions together is the same as saying a *a fraction of a fraction*. For example $\dfrac{1}{2}\times\dfrac{1}{2}$ is the same as saying $\dfrac{1}{2}$ of $\dfrac{1}{2}$.
#
# Therefore if we write $50\%\times 20\%$ we really mean $50\%$ of $20\%$.
#
# The simplest approach to doing this is to first convert each fraction into their decimal representation (divide them by 100), so
#
# $$
# 50\%\div 100=0.50$$ and $$20\%\div 100=0.20
# $$
#
# Now that we have each fraction shown as their decimal representation we simply multiply them together:
#
# $$
# 0.50\times0.20=0.10
# $$
#
# and again to get this decimal to a percent we multiply by 100
#
# $$
# 0.10\times100=10\%
# $$
#
# Putting this into words we get: *$50\%$ of $20\%$ is $10\%$ (One half of $20\%$ is $10\%$)*.
# ## Sports Example
#
# As we know, statistics play a huge part in sports. Keeping track of a team's wins/losses or how many points a player has are integral parts of today's professional sports. Some of these stats may require more interesting mathematical formulas to figure them out. One such example is a goalie’s save percentage in hockey.
#
# The save percentage is the ratio of how many shots the goalie saved over how many he/she has faced. If you are familiar with the NHL you will know this statistic for goalies as Sv\% and is represented as a number like 0.939. In this case the $0.939$ is the percentage we are interested in. You can multiply this number by $100$ to get it in the familiar form $93.9\%$. This means the Sv\% is $93.9\%$, so this particular goalie has saved $93.9\%$ of the shots he's/she's faced.
#
# You will see below a "sport" like game. The objective of the game is to score on your opponent and protect your own net. As you play the game you will see (in real time) below the game window your Sv% and your opponents Sv%. Play a round or two before we discuss how to get this value.
#
# _**How to play:** choose the winning score from the drop down box then click "Start". In game use your mouse to move your paddle up and down (inside the play area). Don't let the ball go in your net!_
# + tags=["hide-input"] language="html"
# <style>
# .mainBody {
# font-family: Arial, Helvetica, sans-serif;
# }
# #startBtn {
# background-color: cornflowerblue;
# border: none;
# border-radius: 3px;
# font-size: 14px;
# color: white;
# font-weight: bold;
# padding: 2px 8px;
# text-transform: uppercase;
# }
# </style>
# <div class="mainBody">
# <div style="padding-bottom: 10px;">
# <label for="winningScore">Winning Score: </label>
# <select name="Winning Score" id="winningScore">
# <option value="3">3</option>
# <option value="5">5</option>
# <option value="7">7</option>
# <option value="10">10</option>
# </select>
# <button type="button" id="startBtn">Start</button>
# </div>
# <canvas id="gameCanvas" width="600" height="350" style="border: solid 1px black"></canvas>
#
# <div>
# <ul>
# <li>Player's point save average: <output id="playerAvg"></output></li>
# <li>Computer's point save average: <output id="compAvg"></output></li>
# </ul>
# </div>
# </div>
# -
# If you look below the game screen you will see "Player's point save average" and "Computer's point save average". You might also have noticed these values changed every time a save was made (unless Sv% was 1) or a score happened, can you come up with a formula to get these values?
#
# The Sv% value is the ratio of how many saves was made over how many total shots the player faced so our formula is
#
# $$
# Sv\%=\frac{saved \ shots}{total \ shots}
# $$
#
# Let's assume the player faced $33$ shots and let in $2$, then the player's Sv% is
#
# $$
# Sv\%=\frac{(33-2)}{33}=0.939
# $$
#
# *Note: $(33-2)$ is how many shots where saved since the total was $33$ and the player let in $2$*
# ## Questions
# + tags=["hide-input"] language="html"
# <style>
# hr {
# width: 60%;
# margin-left: 20px;
# }
# </style>
# <main>
# <div class="questions">
# <ul style="list-style: none">
# <h4>Question #1</h4>
# <li>
# <label for="q1" class="question">A new goalie played his first game and got a shutout (did not let
# the other team score) and made 33 saves, what is his Sv%? </label>
# </li>
# <li>
# <input type="text" id="q1" class="questionInput">
# <button id="q1Btn" onclick="checkAnswer('q1')" class="ansBtn">Check Answer</button>
# </li>
# <li>
# <p class="q1Ans" id="q1True" style="display: none">✓ That's right! Until the goalie let's
# his/her
# first goal in he/she will have a Sv% of 1</p>
# </li>
# <li>
# <p class="q1Ans" id="q1False" style="display: none">Not quite, don't forget to take the total
# amount of shots minus how many went in the net</p>
# </li>
# </ul>
# </div>
# <hr>
# <div class="questions">
# <ul style="list-style: none">
# <h4>Question #2</h4>
# <li>
# <label for="q2" class="question">If a goalie has a Sv% of .990 can he/she ever get back to a Sv% of
# 1.00?</label>
# </li>
# <li>
# <select id="q2">
# <option value="Yes">Yes</option>
# <option value="No">No</option>
# </select>
# <button id="q2Btn" onclick="checkAnswer('q2')" class="ansBtn">Check Answer</button>
# </li>
# <li>
# <p class="q2Ans" id="q2True" style="display: none">✓ That's correct, the goalie could get back
# up to
# 0.999 but never 1.00</p>
# </li>
# <li>
# <p class="q2Ans" id="q2False" style="display: none">Not quite, the goalie could get back up to 0.999
# but never 1.00</p>
# </li>
# </ul>
# </div>
# <hr>
# <div class="questions">
# <ul style="list-style: none">
# <h4>Question #3</h4>
# <li>
# <label for="q3" class="question">A student received a mark of 47/50 on his unit exam, what
# percentage did he get?</label>
# </li>
# <li>
# <input type="text" id="q3" class="questionInput">
# <button id="q3tn" onclick="checkAnswer('q3')" class="ansBtn">Check Answer</button>
# </li>
# <li>
# <p class="q3Ans" id="q3True" style="display: none">✓ That's correct!</p>
# </li>
# <li>
# <p class="q3Ans" id="q3False" style="display: none">Not quite, try again</p>
# </li>
# </ul>
# </div>
# <hr>
# <div class="questions">
# <ul style="list-style: none">
# <h4>Question #4</h4>
# <li>
# <label for="q4" class="question">In a class of 24 students, 8 students own cats, 12 students own dogs
# and 6 students own both cats and dogs. What is the percentage of students who own both cats and
# dogs?</label>
# </li>
# <li>
# <input type="text" id="q4" class="questionInput">
# <button id="q4tn" onclick="checkAnswer('q4')" class="ansBtn">Check Answer</button>
# </li>
# <li>
# <p class="q4Ans" id="q4True" style="display: none">✓ That's correct!</p>
# </li>
# <li>
# <p class="q4Ans" id="q4False" style="display: none">Not quite, try again</p>
# </li>
# </ul>
# </div>
#
# </main>
# <script>
# checkAnswer = function(q) {
# var val = document.getElementById(q).value;
# var isCorrect = false;
# $("."+q+"Ans").css("display", "none");
# switch(q) {
# case 'q1' : Number(val) === 1 ? isCorrect = true : isCorrect = false; break;
# case 'q2' : val === 'No' ? isCorrect = true : isCorrect = false; break;
# case 'q3' : (val === '94%'|| val === '94.0%' || Number(val) === 94) ? isCorrect = true : isCorrect = false;break;
# case 'q4' : (Number(val) === 25 || val === '25%' || val === '25.0%') ? isCorrect = true : isCorrect = false; break;
# default : return false;
# }
#
# if(isCorrect) {
# $("#"+q+"True").css("display", "block");
# } else {
# $("#"+q+"False").css("display", "block");
# }
# }
# </script>
#
# -
# ## Conclusion
#
# As we saw in this notebook, percentages show up in many different ways and are very useful when describing a ratio. It allows for demonstrating any ratio on a familiar scale ($100$) to make data easier to understand. In this notebook we covered the following:
# - A percentage simply represents a fraction
# - To convert any fraction to a percent we turn it into it's decimal form and add $100$
# - A percentage of an amount is simply a fraction multiplication problem
# - To add or subtract a percentage of an amount we first find the percent value than add/subtract from the original value
# - When adding a percentage to an amount we an use the decimal form of percent and add $1$ to it (for example $\$12\times(0.05+1)=\$12.60$)
#
# Keep practising converting fractions to percentages and it will eventually become second nature!
# + tags=["hide-input"] language="html"
# <script>
# var canvas;
# var canvasContext;
# var isInitialized;
#
# var ballX = 50;
# var ballY = 50;
# var ballSpeedX = 5;
# var ballSpeedY = 3;
#
# var leftPaddleY = 250;
# var rightPaddleY = 250;
#
# var playerSaves = 0;
# var playerSOG = 0;
# var compSaves = 0;
# var compSOG = 0;
#
# var playerScore = 0;
# var compScore = 0;
# var winningScore = 3;
# var winScreen = false;
#
# var PADDLE_WIDTH = 10;
# var PADDLE_HEIGHT = 100;
# var BALL_RADIUS = 10;
# var COMP_SPEED = 4;
#
# document.getElementById('startBtn').onclick = function () {
# initGame();
# var selection = document.getElementById('winningScore');
# winningScore = Number(selection.options[selection.selectedIndex].value);
# canvas = document.getElementById('gameCanvas');
# canvasContext = canvas.getContext('2d');
# canvasContext.font = '50px Arial';
# ballReset();
#
# if (!isInitialized) {
# var framesPerSec = 60;
# setInterval(function () {
# moveAll();
# drawAll();
# }, 1000 / framesPerSec);
# isInitialized = true;
# }
#
# canvas.addEventListener('mousemove', function (event) {
# var mousePos = mouseYPos(event);
# leftPaddleY = mousePos.y - PADDLE_HEIGHT / 2;
# });
# }
#
# function updateSaveAvg() {
# var playerSaveAvgTxt = document.getElementById('playerAvg');
# var compSaveAvgTxt = document.getElementById('compAvg');
#
# var playerSaveAvg = playerSaves / playerSOG;
# var compSaveAvg = compSaves / compSOG;
#
# playerSaveAvgTxt.textContent = ((playerSaveAvg < 0 || isNaN(playerSaveAvg)) ? Number(0).toPrecision(3) + (' (0.0%)') :
# playerSaveAvg.toPrecision(3) + (' (' + (playerSaveAvg * 100).toPrecision(3) + '%)'));
# compSaveAvgTxt.textContent = ((compSaveAvg < 0 || isNaN(compSaveAvg)) ? Number(0).toPrecision(3) + (' (0.0%)') :
# compSaveAvg.toPrecision(
# 3) + (' (' + (compSaveAvg * 100).toPrecision(3) + '%)'));
#
# }
#
# function initGame() {
# playerScore = 0;
# compScore = 0;
# playerSaves = 0;
# playerSOG = 0;
# compSaves = 0;
# compSOG = 0;
# ballSpeedX = 5;
# ballSpeedY = 3;
# }
#
# function ballReset() {
# if (playerScore >= winningScore || compScore >= winningScore) {
# winScreen = true;
# }
# if (winScreen) {
# updateSaveAvg();
# if (confirm('Another game?')) {
# winScreen = false;
# initGame();
# } else {
# return;
# }
# }
# ballX = canvas.width / 2;
# ballY = canvas.height / 2;
# ballSpeedY = Math.floor(Math.random() * 4) + 1;
# var randomizer = Math.floor(Math.random() * 2) + 1;
# if (randomizer % 2 === 0) {
# ballSpeedY -= ballSpeedY;
# }
# flipSide();
# }
#
# function flipSide() {
# ballSpeedX = -ballSpeedX;
# }
#
# function moveAll() {
# if (winScreen) {
# return;
# }
# computerMove();
# ballX += ballSpeedX;
# if (ballX < (0 + BALL_RADIUS)) {
# if (ballY > leftPaddleY && ballY < leftPaddleY + PADDLE_HEIGHT) {
# playerSaves++;
# playerSOG++;
# flipSide();
# var deltaY = ballY - (leftPaddleY + PADDLE_HEIGHT / 2);
# ballSpeedY = deltaY * 0.35;
# } else {
# playerSOG++;
# compScore++;
# if (compScore === winningScore) {
# updateSaveAvg();
# drawAll();
# alert('Computer wins, final score: ' + playerScore + '-' + compScore);
# }
# ballReset();
# }
# }
# if (ballX >= canvas.width - BALL_RADIUS) {
# if (ballY > rightPaddleY && ballY < rightPaddleY + PADDLE_HEIGHT) {
# compSaves++;
# compSOG++;
# flipSide();
# var deltaY = ballY - (rightPaddleY + PADDLE_HEIGHT / 2);
# ballSpeedY = deltaY * 0.35;
# } else {
# compSOG++;
# playerScore++;
# if (playerScore === winningScore) {
# updateSaveAvg();
# drawAll();
# alert('You win, final score: ' + playerScore + '-' + compScore);
# }
# ballReset();
# }
# }
# ballY += ballSpeedY;
# if (ballY >= canvas.height - BALL_RADIUS || ballY < 0 + BALL_RADIUS) {
# ballSpeedY = -ballSpeedY;
# }
# updateSaveAvg();
# }
#
# function computerMove() {
# var rightPaddleYCenter = rightPaddleY + (PADDLE_HEIGHT / 2)
# if (rightPaddleYCenter < ballY - 20) {
# rightPaddleY += COMP_SPEED;
# } else if (rightPaddleYCenter > ballY + 20) {
# rightPaddleY -= COMP_SPEED;
# }
# }
#
# function mouseYPos(event) {
# var rect = canvas.getBoundingClientRect();
# var root = document.documentElement;
# var mouseX = event.clientX - rect.left - root.scrollLeft;
# var mouseY = event.clientY - rect.top - root.scrollTop;
# return {
# x: mouseX,
# y: mouseY
# };
# }
#
# function drawAll() {
#
# colorRect(0, 0, canvas.width, canvas.height, 'black');
# if (winScreen) {
# drawNet();
# drawScore();
# return;
# }
# //Left paddle
# colorRect(1, leftPaddleY, PADDLE_WIDTH, PADDLE_HEIGHT, 'white');
# //Right paddle
# colorRect(canvas.width - PADDLE_WIDTH - 1, rightPaddleY, PADDLE_WIDTH, PADDLE_HEIGHT, 'white');
# //Ball
# colorCircle(ballX, ballY, BALL_RADIUS, 'white');
#
# drawNet();
#
# drawScore();
#
# }
#
# function colorRect(x, y, width, height, drawColor) {
# canvasContext.fillStyle = drawColor;
# canvasContext.fillRect(x, y, width, height);
# }
#
# function colorCircle(centerX, centerY, radius, drawColor) {
# canvasContext.fillStyle = 'drawColor';
# canvasContext.beginPath();
# canvasContext.arc(centerX, centerY, radius, 0, Math.PI * 2, true);
# canvasContext.fill();
# }
#
# function drawScore() {
# canvasContext.fillText(playerScore, (canvas.width / 2) - (canvas.width / 4) - 25, 100);
# canvasContext.fillText(compScore, (canvas.width / 2) + (canvas.width / 4) - 25, 100);
# }
#
# function drawNet() {
# for (var i = 0; i < 60; i++) {
# if (i % 2 === 1) {
# colorRect(canvas.width / 2 - 3, i * 10, 6, 10, 'white')
# }
# }
# }
# </script>
# -
# [](https://github.com/callysto/curriculum-notebooks/blob/master/LICENSE.md)
| _build/html/_sources/curriculum-notebooks/Mathematics/Percentage/percentage.ipynb |
End of preview. Expand
in Data Studio
This is starcoderdata, but with leading boilerplate text/license text removed, and with short sequences filtered out. It also removes the extra tags at the beginning of some of the files, like <reponame>.
- Downloads last month
- 1,216