Model card for vit_base_patch16_224.owkin_pancancer
A Vision Transformer (ViT) image classification model.
Trained by Owkin on 40 million pan-cancer histology tiles from TCGA-COAD.
A version using the transformers library is also available here: https://huggingface.co/owkin/phikon
Model Details
- Model Type: Feature backbone
- Developed by: Owkin
- Funded by: Owkin and IDRIS
- Model Stats:
- Params: 85.8M (base)
- Image size: 224 x 224 x 3
- Patch size: 16 x 16 x 3
- Pre-training:
- Papers:
- Original: https://github.com/owkin/HistoSSLscaling
- License: Owkin non-commercial license
Model Usage
Image Embeddings
from urllib.request import urlopen
from PIL import Image
import timm
# get example histology image
img = Image.open(
urlopen(
"https://github.com/owkin/HistoSSLscaling/raw/main/assets/example.tif"
)
)
# load model from the hub
model = timm.create_model(
model_name="hf-hub:1aurent/vit_base_patch16_224.owkin_pancancer",
pretrained=True,
).eval()
# get model specific transforms (normalization, resize)
data_config = timm.data.resolve_model_data_config(model)
transforms = timm.data.create_transform(**data_config, is_training=False)
data = transforms(img).unsqueeze(0) # input is a (batch_size, num_channels, img_size, img_size) shaped tensor
output = model(data) # output is a (batch_size, num_features) shaped tensor
Citation
@article{Filiot2023.07.21.23292757,
author = {Alexandre Filiot and Ridouane Ghermi and Antoine Olivier and Paul Jacob and Lucas Fidon and Alice Mac Kain and Charlie Saillard and Jean-Baptiste Schiratti},
title = {Scaling Self-Supervised Learning for Histopathology with Masked Image Modeling},
elocation-id = {2023.07.21.23292757},
year = {2023},
doi = {10.1101/2023.07.21.23292757},
publisher = {Cold Spring Harbor Laboratory Press},
url = {https://www.medrxiv.org/content/early/2023/09/14/2023.07.21.23292757},
eprint = {https://www.medrxiv.org/content/early/2023/09/14/2023.07.21.23292757.full.pdf},
journal = {medRxiv}
}
- Downloads last month
- 39
Inference API (serverless) has been turned off for this model.
Model tree for 1aurent/vit_base_patch16_224.owkin_pancancer
Datasets used to train 1aurent/vit_base_patch16_224.owkin_pancancer
Evaluation results
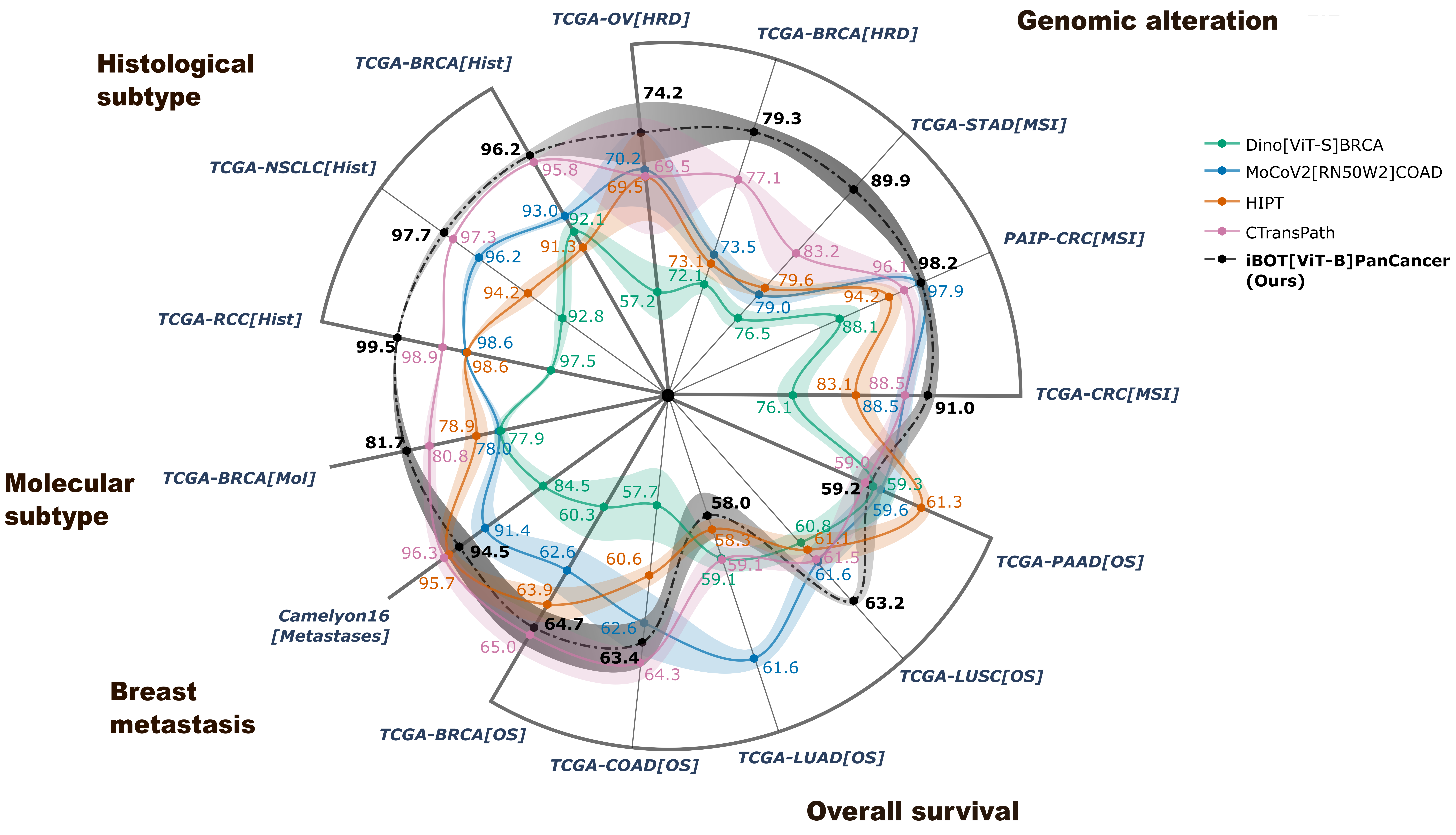
- ROC AUC on Camelyon16[Meta]self-reported94.5 ± 4.4
- ROC AUC on TCGA-BRCA[Hist]self-reported96.2 ± 3.3
- ROC AUC on TCGA-BRCA[HRD]self-reported79.3 ± 2.4
- ROC AUC on TCGA-BRCA[Mol]self-reported81.7 ± 1.6
- ROC AUC on TCGA-BRCA[OS]self-reported64.7 ± 5.7
- ROC AUC on TCGA-CRC[MSI]self-reported91.0 ± 2.2
- ROC AUC on TCGA-COAD[OS]self-reported63.4 ± 7.4
- ROC AUC on TCGA-NSCLC[CType]self-reported97.7 ± 1.3
- ROC AUC on TCGA-LUAD[OS]self-reported53.8 ± 4.5
- ROC AUC on TCGA-LUSC[OS]self-reported62.2 ± 2.9