File size: 7,302 Bytes
650e244 3c8069c 650e244 6ecf3f0 650e244 6ecf3f0 650e244 5c16b7b 3c8069c 650e244 5c16b7b fd8c5bc 3c8069c 650e244 3c8069c 650e244 6ecf3f0 650e244 3c8069c 650e244 3c8069c 650e244 f0039cf 650e244 3c8069c 650e244 f0039cf 650e244 8b5b5dc 650e244 8008a76 650e244 fd8c5bc 8b5b5dc 650e244 8008a76 650e244 3c8069c 650e244 8b5b5dc 650e244 8008a76 650e244 8b5b5dc 650e244 8008a76 650e244 8b5b5dc 650e244 8008a76 650e244 |
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 |
---
tags:
- monai
- medical
library_name: monai
license: apache-2.0
---
# Model Overview
A pre-trained model for segmenting nuclei cells with user clicks/interactions.



This model is trained using [BasicUNet](https://docs.monai.io/en/latest/networks.html#basicunet) over [ConSeP](https://warwick.ac.uk/fac/cross_fac/tia/data/hovernet) dataset.
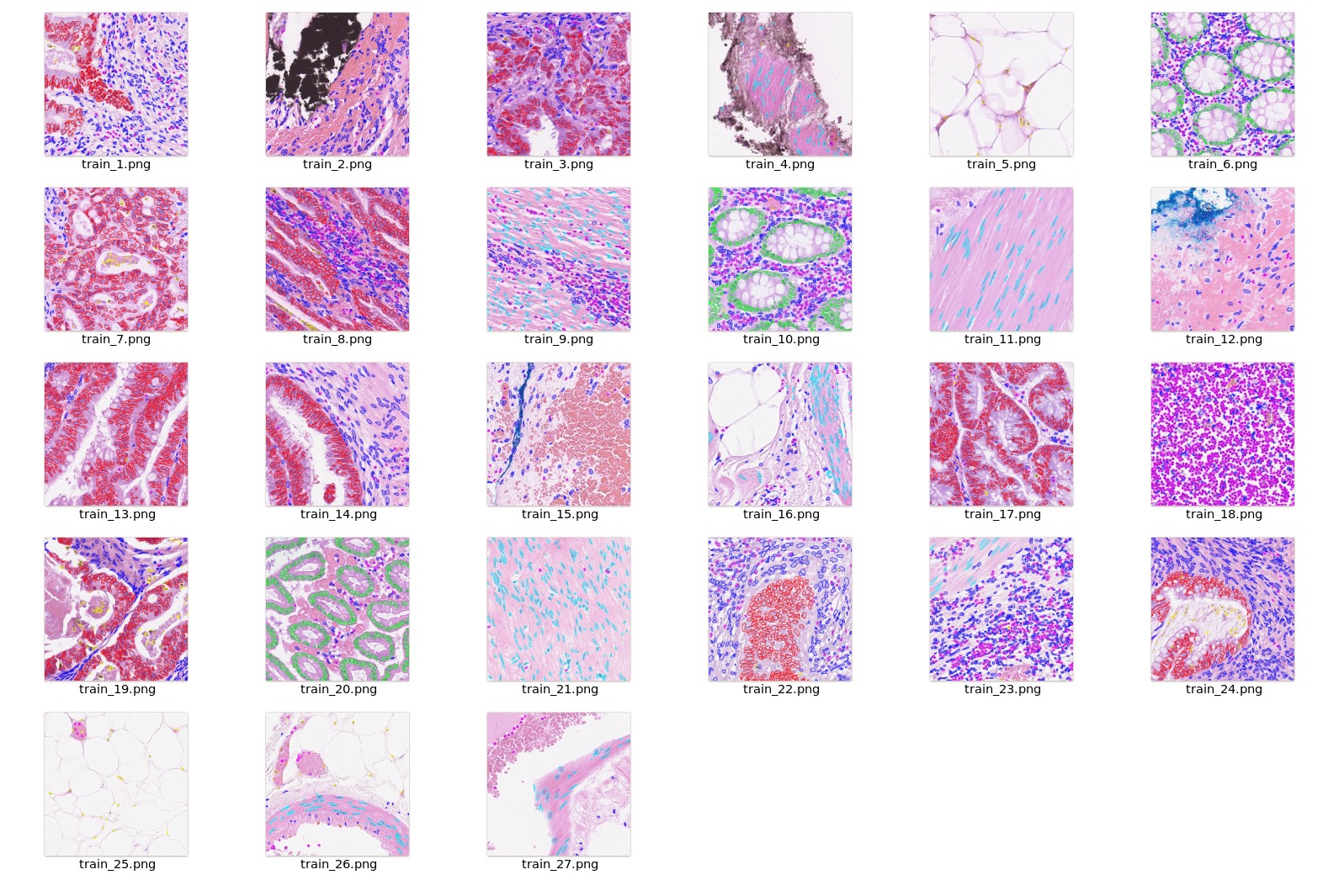
## Data
The training dataset is from https://warwick.ac.uk/fac/cross_fac/tia/data/hovernet
```commandline
wget https://warwick.ac.uk/fac/cross_fac/tia/data/hovernet/consep_dataset.zip
unzip -q consep_dataset.zip
```
<br/>
### Preprocessing
After [downloading this dataset](https://warwick.ac.uk/fac/cross_fac/tia/data/hovernet/consep_dataset.zip),
python script `data_process.py` from `scripts` folder can be used to preprocess and generate the final dataset for training.
```
python scripts/data_process.py --input /path/to/data/CoNSeP --output /path/to/data/CoNSePNuclei
```
After generating the output files, please modify the `dataset_dir` parameter specified in `configs/train.json` and `configs/inference.json` to reflect the output folder which contains new dataset.json.
Class values in dataset are
- 1 = other
- 2 = inflammatory
- 3 = healthy epithelial
- 4 = dysplastic/malignant epithelial
- 5 = fibroblast
- 6 = muscle
- 7 = endothelial
As part of pre-processing, the following steps are executed.
- Crop and Extract each nuclei Image + Label (128x128) based on the centroid given in the dataset.
- Combine classes 3 & 4 into the epithelial class and 5,6 & 7 into the spindle-shaped class.
- Update the label index for the target nuclei based on the class value
- Other cells which are part of the patch are modified to have label idx = 255
Example dataset.json
```json
{
"training": [
{
"image": "/workspace/data/CoNSePNuclei/Train/Images/train_1_3_0001.png",
"label": "/workspace/data/CoNSePNuclei/Train/Labels/train_1_3_0001.png",
"nuclei_id": 1,
"mask_value": 3,
"centroid": [
64,
64
]
}
],
"validation": [
{
"image": "/workspace/data/CoNSePNuclei/Test/Images/test_1_3_0001.png",
"label": "/workspace/data/CoNSePNuclei/Test/Labels/test_1_3_0001.png",
"nuclei_id": 1,
"mask_value": 3,
"centroid": [
64,
64
]
}
]
}
```
## Training Configuration
The training was performed with the following:
- GPU: at least 12GB of GPU memory
- Actual Model Input: 5 x 128 x 128
- AMP: True
- Optimizer: Adam
- Learning Rate: 1e-4
- Loss: DiceLoss
### Memory Consumption
- Dataset Manager: CacheDataset
- Data Size: 13,136 PNG images
- Cache Rate: 1.0
- Single GPU - System RAM Usage: 4.7G
### Memory Consumption Warning
If you face memory issues with CacheDataset, you can either switch to a regular Dataset class or lower the caching rate `cache_rate` in the configurations within range [0, 1] to minimize the System RAM requirements.
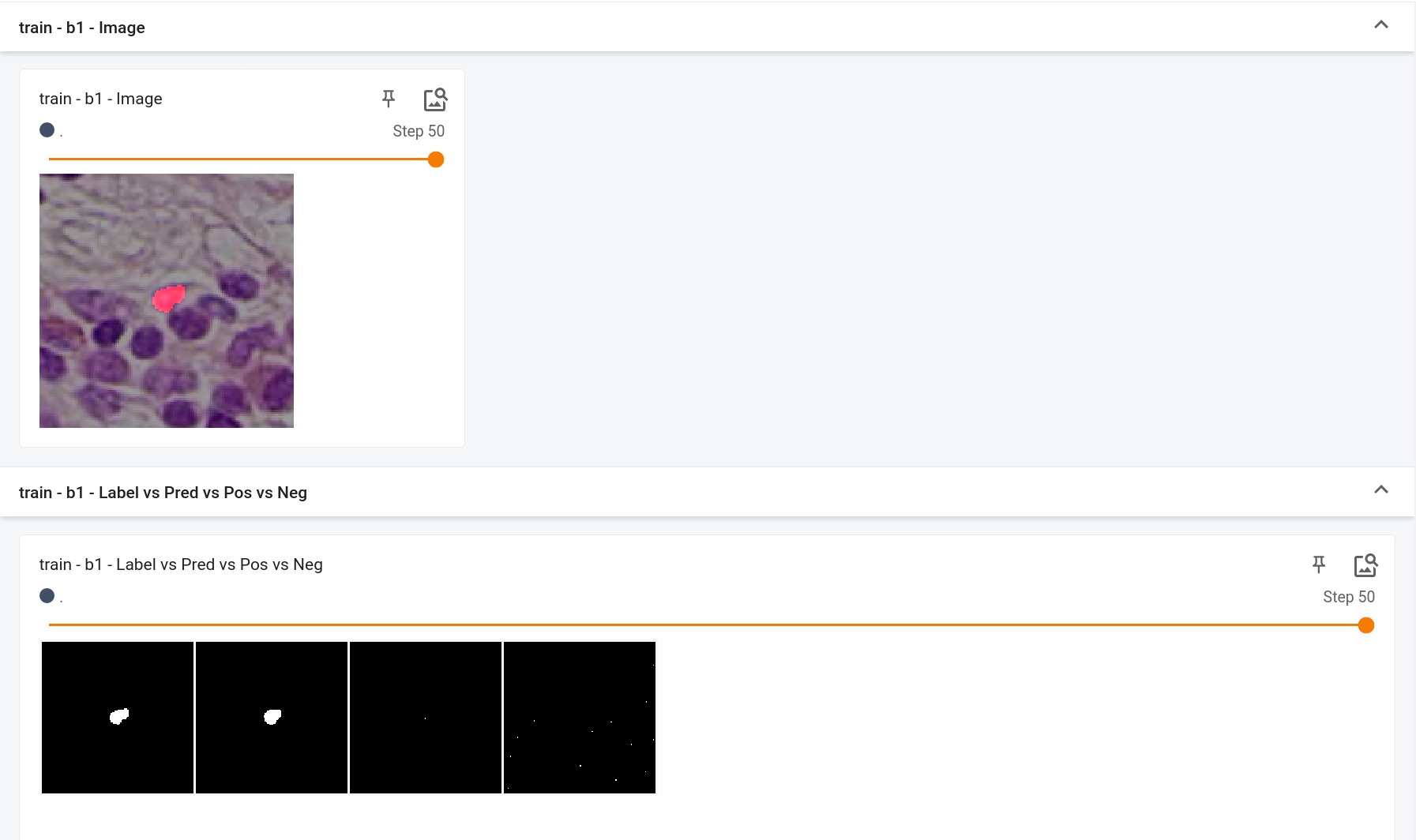
## Input
5 channels
- 3 RGB channels
- +ve signal channel (this nuclei)
- -ve signal channel (other nuclei)
## Output
2 channels
- 0 = Background
- 1 = Nuclei
## Performance
This model achieves the following Dice score on the validation data provided as part of the dataset:
- Train Dice score = 0.89
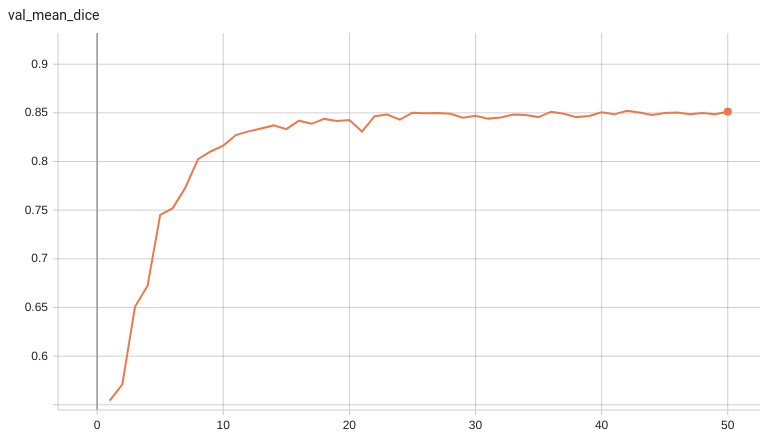
- Validation Dice score = 0.85
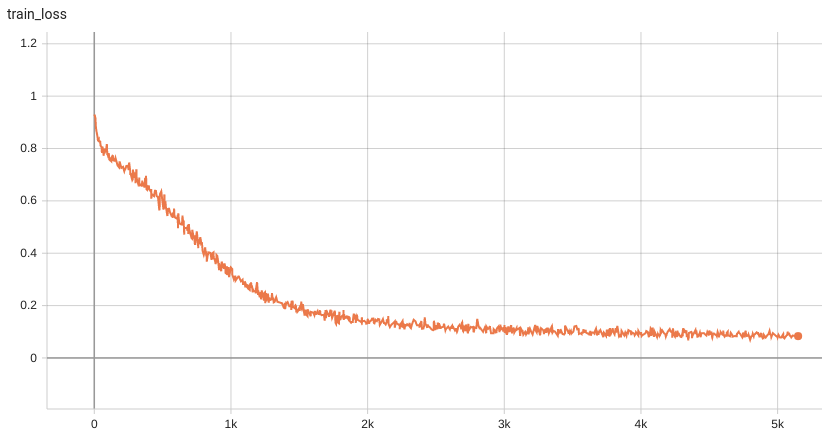
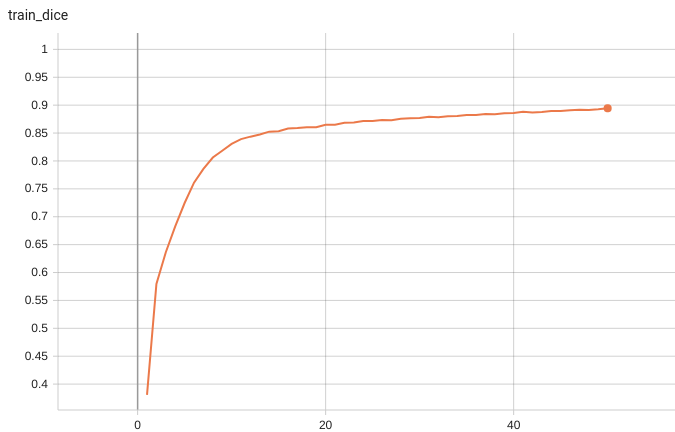
#### Training Loss and Dice
A graph showing the training Loss and Dice over 50 epochs.
 <br>
 <br>
#### Validation Dice
A graph showing the validation mean Dice over 50 epochs.
 <br>
## MONAI Bundle Commands
In addition to the Pythonic APIs, a few command line interfaces (CLI) are provided to interact with the bundle. The CLI supports flexible use cases, such as overriding configs at runtime and predefining arguments in a file.
For more details usage instructions, visit the [MONAI Bundle Configuration Page](https://docs.monai.io/en/latest/config_syntax.html).
#### Execute training:
```
python -m monai.bundle run --config_file configs/train.json
```
Please note that if the default dataset path is not modified with the actual path in the bundle config files, you can also override it by using `--dataset_dir`:
```
python -m monai.bundle run --config_file configs/train.json --dataset_dir <actual dataset path>
```
#### Override the `train` config to execute multi-GPU training:
```
torchrun --standalone --nnodes=1 --nproc_per_node=2 -m monai.bundle run --config_file "['configs/train.json','configs/multi_gpu_train.json']"
```
Please note that the distributed training-related options depend on the actual running environment; thus, users may need to remove `--standalone`, modify `--nnodes`, or do some other necessary changes according to the machine used. For more details, please refer to [pytorch's official tutorial](https://pytorch.org/tutorials/intermediate/ddp_tutorial.html).
#### Override the `train` config to execute evaluation with the trained model:
```
python -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json']"
```
#### Override the `train` config and `evaluate` config to execute multi-GPU evaluation:
```
torchrun --standalone --nnodes=1 --nproc_per_node=2 -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json','configs/multi_gpu_evaluate.json']"
```
#### Execute inference:
```
python -m monai.bundle run --config_file configs/inference.json
```
# References
[1] Koohbanani, Navid Alemi, et al. "NuClick: a deep learning framework for interactive segmentation of microscopic images." Medical Image Analysis 65 (2020): 101771. https://arxiv.org/abs/2005.14511.
[2] S. Graham, Q. D. Vu, S. E. A. Raza, A. Azam, Y-W. Tsang, J. T. Kwak and N. Rajpoot. "HoVer-Net: Simultaneous Segmentation and Classification of Nuclei in Multi-Tissue Histology Images." Medical Image Analysis, Sept. 2019. [[doi](https://doi.org/10.1016/j.media.2019.101563)]
[3] NuClick [PyTorch](https://github.com/mostafajahanifar/nuclick_torch) Implementation
# License
Copyright (c) MONAI Consortium
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
|