The full dataset viewer is not available (click to read why). Only showing a preview of the rows.
Error code: DatasetGenerationCastError
Exception: DatasetGenerationCastError
Message: An error occurred while generating the dataset
All the data files must have the same columns, but at some point there are 3 new columns ({'id', 'instances', 'name'}) and 2 missing columns ({'output', 'input'}).
This happened while the json dataset builder was generating data using
hf://datasets/BAAI/OPI/OPI_DATA/AP/Function/test/CASPSimilarSeq_function_test.jsonl (at revision 893d4028ed394a960d1730c82cb4694bba1efcb2)
Please either edit the data files to have matching columns, or separate them into different configurations (see docs at https://hf.co/docs/hub/datasets-manual-configuration#multiple-configurations)
Traceback: Traceback (most recent call last):
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/builder.py", line 1870, in _prepare_split_single
writer.write_table(table)
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/arrow_writer.py", line 622, in write_table
pa_table = table_cast(pa_table, self._schema)
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/table.py", line 2292, in table_cast
return cast_table_to_schema(table, schema)
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/table.py", line 2240, in cast_table_to_schema
raise CastError(
datasets.table.CastError: Couldn't cast
id: string
name: string
instruction: string
instances: list<item: struct<input: string, output: string>>
child 0, item: struct<input: string, output: string>
child 0, input: string
child 1, output: string
to
{'instruction': Value(dtype='string', id=None), 'input': Value(dtype='string', id=None), 'output': Value(dtype='string', id=None)}
because column names don't match
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "/src/services/worker/src/worker/job_runners/config/parquet_and_info.py", line 1438, in compute_config_parquet_and_info_response
parquet_operations = convert_to_parquet(builder)
File "/src/services/worker/src/worker/job_runners/config/parquet_and_info.py", line 1050, in convert_to_parquet
builder.download_and_prepare(
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/builder.py", line 924, in download_and_prepare
self._download_and_prepare(
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/builder.py", line 1000, in _download_and_prepare
self._prepare_split(split_generator, **prepare_split_kwargs)
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/builder.py", line 1741, in _prepare_split
for job_id, done, content in self._prepare_split_single(
File "/src/services/worker/.venv/lib/python3.9/site-packages/datasets/builder.py", line 1872, in _prepare_split_single
raise DatasetGenerationCastError.from_cast_error(
datasets.exceptions.DatasetGenerationCastError: An error occurred while generating the dataset
All the data files must have the same columns, but at some point there are 3 new columns ({'id', 'instances', 'name'}) and 2 missing columns ({'output', 'input'}).
This happened while the json dataset builder was generating data using
hf://datasets/BAAI/OPI/OPI_DATA/AP/Function/test/CASPSimilarSeq_function_test.jsonl (at revision 893d4028ed394a960d1730c82cb4694bba1efcb2)
Please either edit the data files to have matching columns, or separate them into different configurations (see docs at https://hf.co/docs/hub/datasets-manual-configuration#multiple-configurations)Need help to make the dataset viewer work? Make sure to review how to configure the dataset viewer, and open a discussion for direct support.
instruction
string | input
string | output
string |
|---|---|---|
Can you provide the functional description of the following protein sequence? | MRWQEMGYIFYPRKLR | Regulates insulin sensitivity and metabolic homeostasis. Inhibits the folate cycle, thereby reducing de novo purine biosynthesis which leads to the accumulation of the de novo purine synthesis intermediate 5-aminoimidazole-4-carboxamide (AICAR) and the activation of the metabolic regulator 5'-AMP-activated protein kinase (AMPK). Protects against age-dependent and diet-induced insulin resistance as well as diet-induced obesity. In response to metabolic stress, translocates to the nucleus where it binds to antioxidant response elements (ARE) present in the promoter regions of a number of genes and plays a role in regulating nuclear gene expression in an NFE2L2-dependent manner and increasing cellular resistance to metabolic stress. Increases mitochondrial respiration and levels of CPT1A and cytokines IL1B, IL6, IL8, IL10 and TNF in senescent cells. Increases activity of the serine/threonine protein kinase complex mTORC2 and reduces activity of the PTEN phosphatase, thus promoting phosphorylation of AKT. This promotes AKT-mediated phosphorylation of transcription factor FOXO1 which reduces FOXO1 activity, leading to reduced levels of MSTN and promotion of skeletal muscle growth. Promotes osteogenic differentiation of bone marrow mesenchymal stem cells via the TGFB/SMAD pathway. Promotes osteoblast proliferation and osteoblast synthesis of type I collagens COL1A1 and COL1A2 via the TGFB/SMAD pathway. |
Predict the functional description of the protein sequence below. | MLLLLLLLLLLPPLVLRVAASRCLHDETQKSVSLLRPPFSQLPSKSRSSSLTLPSSRDPQPLRIQSCYLGDHISDGAWDPEGEGMRGGSRALAAVREATQRIQAVLAVQGPLLLSRDPAQYCHAVWGDPDSPNYHRCSLLNPGYKGESCLGAKIPDTHLRGYALWPEQGPPQLVQPDGPGVQNTDFLLYVRVAHTSKCHQETVSLCCPGWSTAAQSQLTAALTSWAQRRGFVMLPRLCLKLLGSSNLPTLASQSIRITGPSVIAYAACCQLDSEDRPLAGTIVYCAQHLTSPSLSHSDIVMATLHELLHALGFSGQLFKKWRDCPSGFSVRENCSTRQLVTRQDEWGQLLLTTPAVSLSLAKHLGVSGASLGVPLEEEEGLLSSHWEARLLQGSLMTATFDGAQRTRLDPITLAAFKDSGWYQVNHSAAEELLWGQGSGPEFGLVTTCGTGSSDFFCTGSGLGCHYLHLDKGSCSSDPMLEGCRMYKPLANGSECWKKENGFPAGVDNPHGEIYHPQSRCFFANLTSQLLPGDKPRHPSLTPHLKEAELMGRCYLHQCTGRGAYKVQVEGSPWVPCLPGKVIQIPGYYGLLFCPRGRLCQTNEDINAVTSPPVSLSTPDPLFQLSLELAGPPGHSLGKEQQEGLAEAVLEALASKGGTGRCYFHGPSITTSLVFTVHMWKSPGCQGPSVATLHKALTLTLQKKPLEVYHGGANFTTQPSKLLVTSDHNPSMTHLRLSMGLCLMLLILVGVMGTTAYQKRATLPVRPSASYHSPELHSTRVPVRGIREV | Putative metalloproteinase that plays a role in left-right patterning process. |
Predict the functional description of the protein sequence below. | MAGIIKKQILKHLSRFTKNLSPDKINLSTLKGEGELKNLELDEEVLQNMLDLPTWLAINKVFCNKASIRIPWTKLKTHPICLSLDKVIMEMSTCEEPRSPNGPSPIATASGQSEYGFAEKVVEGISVSVNSIVIRIGAKAFNASFELSQLRIYSVNAHWEHGDLRFTRIQDPQRGEVLTFKEINWQMIRIEADATQSSHLEIMCAPVRLITNQSKIRVTLKRRLKDCNVIATKLVLILDDLLWVLTDSQLKAMVQYAKSLSEAIEKSTEQRKSMAPEPTQSSTVVASAQQVKTTQTSNAPDVNDAIVKLFNDFDVKETSHHLVISHLDLHICDDIHAKEKESNRRITGGAMQLSFTQLTIDYYPYHKAGDSCNHWMYFSDATKTKNGWANELLHEFECNVEMLKQAVKDHNVGSPPKSPTHASPQHTQTEKDYPLKGTCRTPSVLSQQSKAKLMSSSVVVRLADFNIYQVSTAEQCRSSPKSMICCNKKSLYLPQEMSAVYIEFTEYYYPDGKDFPIPSPNLYSQLNALQFTVDERSILWLNQFLLDLKQSLNQFMAVYKLNDNSKSDEHVDVRVDGLMLKFVIPSEVKSECHQDQPRAISIQSSEMIATNTRHCPNCRHSDLEALFQDFKDCDFFSKTYTSFPKSCDNFNLLHPIFQRHAHEQDTKMHEIYKGNITPQLNKNTLKTSAATDVWAVYFSQFWIDYEGMKSGKGRPISFVDSFPLSIWICQPTRYAESQKEPQTCNQVSLNTSQSESSDLAGRLKRKKLLKEYYSTESEPLTNGGQKPSSSDTFFRFSPSSSEADIHLLVHVHKHVSMQINHYQYLLLLFLHESLILLSENLRKDVEAVTGSPASQTSICIGILLRSAELALLLHPVDQANTLKSPVSESVSPVVPDYLPTENGDFLSSKRKQISRDINRIRSVTVNHMSDNRSMSVDLSHIPLKDPLLFKSASDTNLQKGISFMDYLSDKHLGKISEDESSGLVYKSGSGEIGSETSDKKDSFYTDSSSILNYREDSNILSFDSDGNQNILSSTLTSKGNETIESIFKAEDLLPEAASLSENLDISKEETPPVRTLKSQSSLSGKPKERCPPNLAPLCVSYKNMKRSSSQMSLDTISLDSMILEEQLLESDGSDSHMFLEKGNKKNSTTNYRGTAESVNAGANLQNYGETSPDAISTNSEGAQENHDDLMSVVVFKITGVNGEIDIRGEDTEICLQVNQVTPDQLGNISLRHYLCNRPVGSDQKAVIHSKSSPEISLRFESGPGAVIHSLLAEKNGFLQCHIENFSTEFLTSSLMNIQHFLEDETVATVMPMKIQVSNTKINLKDDSPRSSTVSLEPAPVTVHIDHLVVERSDDGSFHIRDSHMLNTGNDLKENVKSDSVLLTSGKYDLKKQRSVTQATQTSPGVPWPSQSANFPEFSFDFTREQLMEENESLKQELAKAKMALAEAHLEKDALLHHIKKMTVE | Tube-forming lipid transport protein which mediates the transfer of lipids between membranes at organelle contact sites. Required for retrograde traffic of vesicle clusters in the early endocytic pathway to the Golgi complex. |
Predict the functional description of the protein sequence below. | MPGWFKKAWYGLASLLSFSSFILIIVALVVPHWLSGKILCQTGVDLVNATDRELVKFIGDIYYGLFRGCKVRQCGLGGRQSQFTIFPHLVKELNAGLHVMILLLLFLALALALVSMGFAILNMIQVPYRAVSGPGGICLWNVLAGGVVALAIASFVAAVKFHDLTERIANFQEKLFQFVVVEEQYEESFWICVASASAHAANLVVVAISQIPLPEIKTKIEEATVTAEDILY | Plays a key role to hearing function. Required for normal organization and maintenance of the stereocilia bundle and for mechano-electrical transduction. |
Predict the general functional description of the protein sequence. | MGLQPLEFSDCYLDSPWFRERIRAHEAELERTNKFIKELIKDGKNLIAATKSLSVAQRKFAHSLRDFKFEFIGDAVTDDERCIDASLREFSNFLKNLEEQREIMALSVTETLIKPLEKFRKEQLGAVKEEKKKFDKETEKNYSLIDKHLNLSAKKKDSHLQEADIQVEQNRQHFYELSLEYVCKLQEIQERKKFEFVEPMLSFFQGMFTFYHQGHELAKDFNHYKMELQINIQNTRNRFEGTRSEVEELMNKIRQNPKDHKRASQFTAEGYLYVQEKRPAPFGSSWVKHYCMYRKAAKKFNMIPFEHRSGGKLGDGEVFFLKECTKRHTDSIDRRFCFDIEAADRPGVSLTMQAFSEEERKQWLEALGGKEALSHSFNTAIIPRPEGNAQLDKMGFTIIRKCISAVETRGINDQGLYRVVGVSSKVQRLLSMLMDVKTCNEVDLENSADWEVKTITSALKQYLRSLPEPLMTYELHGDFIVPAKSGSPESRVNAIHFLVHKLPEKNKEMLDILVKHLTNVSNHSKQNLMTVANLGVVFGPTLMRPQEETVAALMDLKFQNIVVEILIENHEKIFRTPPDTTFPEPTCLSASPPNAPPRQSKRQGQRTKRPVAVYNLCLELEDGDNPYPSKEDTPTSSLDSLSSPSPVTTAVPGPPGPDKNHLLADGGSFGDWASTIPGQTRSSMVQWLNPQSPTTTSSNSAVTPLSPGSSPFPFSPPATVADKPPESIRSRKARAVYPCEAEHSSELSFEIGAIFEDVQTSREPGWLEGTLNGKRGLIPQNYVKLL | GTPase activator for the small GTPases RhoA and Cdc42 by converting them to an inactive GDP-bound state. Essential for PTKB2 regulation of cytoskeletal organization via Rho family GTPases. Inhibits PAK2 proteolytic fragment PAK-2p34 kinase activity and changes its localization from the nucleus to the perinuclear region. Stabilizes PAK-2p34 thereby increasing stimulation of cell death (By similarity). |
Predict the functional description of the protein sequence below. | MKRRQKRKHLENEESQETAEKGGGMSKSQEDALQPGSTRVAKGWSQGVGEVTSTSEYCSCVSSSRKLIHSGIQRIHRDSPQPQSPLAQVQERGETPPRSQHVSLSSYSSYKTCVSSLCVNKEERGMKIYYMQVQMNKGVAVSWETEETLESLEKQPRMEEVTLSEVVRVGTPPSDVSTRNLLSDSEPSGEEKEHEERTESDSLPGSPTVEDTPRAKTPDWLVTMENGFRCMACCRVFTTMEALQEHVQFGIREGFSCHVFHLTMAQLTGNMESESTQDEQEEENGNEKEEEEKPEAKEEEGQPTEEDLGLRRSWSQCPGCVFHSPKDRNS | Acts as a nuclear transcription factor that positively regulates the expression of heat shock genes. Binds to heat shock promoter elements (HSE). |
Generate the functional description in free-text form based on the protein sequence. | MDDADPEERNYDNMLKMLSDLNKDLEKLLEEMEKISVQATWMAYDMVVMRTNPTLAESMRRLEDAFVNCKEEMEKNWQELLHETKQRL | Major component of the transverse central element of synaptonemal complexes (SCS), formed between homologous chromosomes during meiotic prophase. Required for chromosome loading of the central element-specific SCS proteins, and for initiating synapsis between homologous chromosomes. Chromosome loading appears to require SYCP1. Required for fertility. |
Predict the functional description of the protein sequence below. | MAFSDLTSRTVHLYDNWIKDADPRVEDWLLMSSPLPQTILLGFYVYFVTSLGPKLMENRKPFELKKAMITYNFFIVLFSVYMCYEFVMSGWGIGYSFRCDIVDYSRSPTALRMARTCWLYYFSKFIELLDTIFFVLRKKNSQVTFLHVFHHTIMPWTWWFGVKFAAGGLGTFHALLNTAVHVVMYSYYGLSALGPAYQKYLWWKKYLTSLQLVQFVIVAIHISQFFFMEDCKYQFPVFACIIMSYSFMFLLLFLHFWYRAYTKGQRLPKTVKNGTCKNKDN | Catalyzes the first and rate-limiting reaction of the four reactions that constitute the long-chain fatty acids elongation cycle. This endoplasmic reticulum-bound enzymatic process allows the addition of 2 carbons to the chain of long- and very long-chain fatty acids (VLCFAs) per cycle. Condensing enzyme with higher activity toward C18 acyl-CoAs, especially C18:3(n-3) acyl-CoAs and C18:3(n-6)-CoAs. Also active toward C20:4-, C18:0-, C18:1-, C18:2- and C16:0-CoAs, and weakly toward C20:0-CoA. Little or no activity toward C22:0-, C24:0-, or C26:0-CoAs. May participate in the production of saturated and polyunsaturated VLCFAs of different chain lengths that are involved in multiple biological processes as precursors of membrane lipids and lipid mediators. |
Generate the functional description in free-text form based on the protein sequence. | MPPRRSIVEVKVLDVQKRRVPNKHYVYIIRVTWSSGSTEAIYRRYSKFFDLQMQMLDKFPMEGGQKDPKQRIIPFLPGKILFRRSHIRDVAVKRLIPIDEYCKALIQLPPYISQCDEVLQFFETRPEDLNPPKEEHIGKKKSGGDQTSVDPMVLEQYVVVANYQKQESSEISLSVGQVVDIIEKNESGWWFVSTAEEQGWVPATCLEGQDGVQDEFSLQPEEEEKYTVIYPYTARDQDEMNLERGAVVEVIQKNLEGWWKIRYQGKEGWAPASYLKKNSGEPLPPKPGPGSPSHPGALDLDGVSRQQNAVGREKELLSSQRDGRFEGRPVPDGDAKQRSPKMRQRPPPRRDMTIPRGLNLPKPPIPPQVEEEYYTIAEFQTTIPDGISFQAGLKVEVIEKNLSGWWYIQIEDKEGWAPATFIDKYKKTSNASRPNFLAPLPHEVTQLRLGEAAALENNTGSEATGPSRPLPDAPHGVMDSGLPWSKDWKGSKDVLRKASSDMSASAGYEEISDPDMEEKPSLPPRKESIIKSEGELLERERERQRTEQLRGPTPKPPGVILPMMPAKHIPPARDSRRPEPKPDKSRLFQLKNDMGLECGHKVLAKEVKKPNLRPISKSKTDLPEEKPDATPQNPFLKSRPQVRPKPAPSPKTEPPQGEDQVDICNLRSKLRPAKSQDKSLLDGEGPQAVGGQDVAFSRSFLPGEGPGRAQDRTGKQDGLSPKEISCRAPPRPAKTTDPVSKSVPVPLQEAPQQRPVVPPRRPPPPKKTSSSSRPLPEVRGPQCEGHESRAAPTPGRALLVPPKAKPFLSNSLGGQDDTRGKGSLGPWGTGKIGENREKAAAASVPNADGLKDSLYVAVADFEGDKDTSSFQEGTVFEVREKNSSGWWFCQVLSGAPSWEGWIPSNYLRKKP | Adapter protein involved in invadopodia and podosome formation and extracellular matrix degradation. Binds matrix metalloproteinases (ADAMs), NADPH oxidases (NOXs) and phosphoinositides. Acts as an organizer protein that allows NOX1- or NOX3-dependent reactive oxygen species (ROS) generation and ROS localization. Plays a role in mitotic clonal expansion during the immediate early stage of adipocyte differentiation (By similarity). |
Can you provide the functional description of the following protein sequence? | MFASCHCVPRGRRTMKMIHFRSSSVKSLSQEMRCTIRLLDDSEISCHIQRETKGQFLIDHICNYYSLLEKDYFGIRYVDPEKQRHWLEPNKSIFKQMKTHPPYTMCFRVKFYPHEPLKIKEELTRYLLYLQIKRDIFHGRLLCSFSDAAYLGACIVQAELGDYDPDEHPENYISEFEIFPKQSQKLERKIVEIHKNELRGQSPPVAEFNLLLKAHTLETYGVDPHPCKDSTGTTTFLGFTAAGFVVFQGNKRIHLIKWPDVCKLKFEGKTFYVIGTQKEKKAMLAFHTSTPAACKHLWKCGVENQAFYKYAKSSQIKTVSSSKIFFKGSRFRYSGKVAKEVVEASSKIQREPPEVHRANITQSRSSHSLNKQLIINMEPLQPLLPSPSEQEEELPLGEGVPLPKEENISAPLISSSPVKAAREYEDPPSEEEDKIKEEPLTISELVYNPSASLLPTPVDDDEIDMLFDCPSRLELEREDTDSFEDLEADENAFLIAEEEELKEARRALSWSYDILTGHIRVNPLVKSFSRLLVVGLGLLLFVFPLLLLLLESGIDLSFLCEIRQTPEFEQFHYEYYCPLKEWVAGKVHLILYMLGCS | Putative tumor suppressor gene that may be implicated in the origin and progression of lung cancer. |
Generate the functional description in free-text form based on the protein sequence. | MAGTVLGVGAGVFILALLWVAVLLLCVLLSRASGAARFSVIFLFFGAVIITSVLLLFPRAGEFPAPEVEVKIVDDFFIGRYVLLAFLSAIFLGGLFLVLIHYVLEPIYAKPLHSY | May be involved in ciliary biogenesis or function. |
Predict the functional description of the protein sequence below. | MNLGDGLKLETELLDGKTKLILSPYEHKSKISVKMGNKAKIAKCPLRTKTGHILKSTQDTCIGSEKLLQKKPVGSETSQAKGEKNGMTFSSTKDLCKQCIDKDCLHIQKEISPATPNMQKTRNTVNTSLVGKQKPHKKHITAENMKSSLVCLTQDQLQQILMTVNQGNRSLSLTENGKEAKSQYSLYLNSISNQPKDENIMGLFKKTEMVSSVPAENKSVLNEHQETSKQCEQKIAIENEWKPADIFSTLGERECDRSSLEAKKAQWRKELDEQVALKKKEKEVSEKWNDPWKKSESDKIIWEKHQILDQSRETVLLEHPFSAVKQELQRKWIEELNKQIEDDRQRKIEEKIIYSKGEEHDRWAMHFDSLKSYPGSQSQLFSQSTHKQPEYFCVSPDTQELADVSSVCTPTTGSQVEPSEEEHIAKPIKDVVMANSKKTNFLRSMTALLDPAQIEERDRRRQKQLEHQKAITAQVEEKRRKKQLEEEQRKKEEQEEELRLAQEREEMQKQYEEDILKQKQKEEIMTLKTNELFQTMQRAQELAQRLKQEQRIRELAQKGHDTSRLIKNLGVDTIQMEYNASNISNSRHDSDEISGKMNTYMNSTTSKKDTGVQTDDLNIGIFTNAESHCGSLMERDITNCSSPEISAELIGQFSTKKNKQELTQDKGASLEKENNRCNDQCNQFTRIEKQTKHMKKYPKRPDWNINKPPKRYIPASEKYPKQLQKQREEKKVRRQMELLHLVEKNNPGHLSQNRGISPEIFHSSHQETESKLRWHLVKKEEEPLNIHSFSKERSPSSPVPVVKNRTQQTQNTLHLPLKNSSYERENLISGSNQTELSSGISESSHFIPYVRTNEIYYLDPDAPLSGPSTQDPQYQNSQDCGQKRQLFDSDCVRDPLLNPNMVKNRDRQQAILKGLSELRQGLLQKQKELESSLLPLAENQEESFGSSF | Microtubule-binding protein required for ciliogenesis. May function in ciliogenesis by mediating the transport of proteins like BBS4 to the cilium, but also through the organization of the centriolar satellites. Plays a role in retina morphogenesis and/or homeostasis (By similarity). |
What is the functional description of the protein sequence? | MAGQHLPVPRLEGVSREQFMQHLYPQRKPLVLEGIDLGPCTSKWTVDYLSQVGGKKEVKIHVAAVAQMDFISKNFVYRTLPFDQLVQRAAEEKHKEFFVSEDEKYYLRSLGEDPRKDVADIRKQFPLLKGDIKFPEFFKEEQFFSSVFRISSPGLQLWTHYDVMDNLLIQVTGKKRVVLFSPRDAQYLYLKGTKSEVLNIDNPDLAKYPLFSKARRYECSLEAGDVLFIPALWFHNVISEEFGVGVNIFWKHLPSECYDKTDTYGNKDPTAASRAAQILDRALKTLAELPEEYRDFYARRMVLHIQDKAYSKNSE | tRNA hydroxylase that acts as a component of the wybutosine biosynthesis pathway. Wybutosine is a hyper modified guanosine with a tricyclic base found at the 3'-position adjacent to the anticodon of eukaryotic phenylalanine tRNA. Catalyzes the hydroxylation of 7-(a-amino-a-carboxypropyl)wyosine (yW-72) into undermodified hydroxywybutosine (OHyW*). OHyW* being further transformed into hydroxywybutosine (OHyW) by LCMT2/TYW4. OHyW is a derivative of wybutosine found in higher eukaryotes. |
What is the functional description of the protein sequence? | MALRLVADFDLGKDVLPWLRAQRAVSEASGAGSGGADVLENDYESLHVLNVERNGNIIYTYKDDKGNVVFGLYDCQTRQNELLYTFEKDLQVFSCSVNSERTLLAASLVQSTKEGKRNELQPGSKCLTLLVEIHPVNNVKVLKAVDSYIWVQFLYPHIESHPLPENHLLLISEEKYIEQFRIHVAQEDGNRVVIKNSGHLPRDRIAEDFVWAQWDMSEQRLYYIDLKKSRSILKCIQFYADESYNLMFEVPLDISLSNSGFKLVNFGCDYHQYRDKFSKHLTLCVFTNHTGSLCVCYSPKCASWGQITYSVFYIHKGHSKTFTTSLENVGSHMTKGITFLNLDYYVAVYLPGHFFHLLNVQHPDLICHNLFLTGNNEMIDMLPHCPLQSLSGSLVLDCCSGKLYRALLSQSSLLQLLQNTCLDCEKMAALHCALYCGQGAQFLEAQIIQWISENVSACHSFDLIQEFIIASSYWSVYSETSNMDKLLPHSSVLTWNTEIPGITLVTEDIALPLMKVLSFKGYWEKLNSNLEYVKYAKPHFHYNNSVVRREWHNLISEEKTGKRRSAAYVRNILDNAVKVISNLEARNLGPRLTPLLQEEDSHQRLLMGLMVSELKDHFLRHLQGVEKKKIEQMVLDYISKLLDLICHIVETNWRKHNLHSWVLHFNSRGSAAEFAVFHIMTRILEATNSLFLPLPPGFHTLHTILGVQCLPLHNLLHCIDSGVLLLTETAVIRLMKDLDNTEKNEKLKFSIIVRLPPLIGQKICRLWDHPMSSNIISRNHVTRLLQNYKKQPRNSMINKSSFSVEFLPLNYFIEILTDIESSNQALYPFEGHDNVDAEFVEEAALKHTAMLLGL | Regulator of gamma-secretase activity, which specifically activates the production of amyloid-beta protein (amyloid-beta protein 40 and amyloid-beta protein 42), without affecting the cleavage of other gamma-secretase targets such has Notch. The gamma-secretase complex is an endoprotease complex that catalyzes the intramembrane cleavage of integral membrane proteins such as Notch receptors and APP (amyloid-beta precursor protein). Specifically promotes the gamma-cleavage of APP CTF-alpha (also named APP-CTF) by the gamma-secretase complex to generate amyloid-beta, while it reduces the epsilon-cleavage of APP CTF-alpha, leading to a low production of AICD. |
Predict the functional description of the protein sequence below. | MARVGPGRAGVSCQGRGRGRGGSGQRRPPTWEISDSDAEDSAGSEAAARARDPAGERRAAAEALRLLRPEQVLKRLAVCVDTAILEDAGADVLMEALEALGCECRIEPQRPARSLRWTRASPDPCPRSLPPEVWAAGEQELLLLLEPEEFLQGVATLTQISGPTHWVPWISPETTARPHLAVIGLDAYLWSRQHVSRGTQQPESPKVAGAEVAVSWPEVEEALVLLQLWANLDVLLVASWQELSRHVCAVTKALAQYPLKQYRESQAFSFCTAGRWAAGEPVARDGAGLQAAWRRQIRQFSRVSPAVADAVVTAFPSPRLLQQALEACSTERERMGLLADLPVPPSEGGRPRRVGPDLSRRICLFLTTANPDLLLDLGS | Interacts with MUS81 to form a DNA structure-specific endonuclease which cleaves substrates such as 3'-flap structures. |
Generate the functional description in free-text form based on the protein sequence. | MTSPLCRAASANALPPQDQASTPSSRVKGREASGKPSHLRGKGTAQAWTPGRSKGGSFHRGAGKPSVHSQVAELHKKIQLLEGDRKAFFESSQWNIKKNQETISQLRKETKALELKLLDLLKGDEKVVQAVIREWKWEKPYLKNRTGQALEHLDHRLREKVKQQNALRHQVVLRQRRLEELQLQHSLRLLEMAEAQNRHTEVAKTMRNLENRLEKAQMKAQEAEHITSVYLQLKAYLMDESLNLENRLDSMEAEVVRTKHELEALHVVNQEALNARDIAKNQLQYLEETLVRERKKRERYISECKKRAEEKKLENERMERKTHREHLLLQSDDTIQDSLHAKEEELRQRWSMYQMEVIFGKVKDATGTDETHSLVRRFLAQGDTFAQLETLKSENEQTLVRLKQEKQQLQRELEDLKYSGEATLVSQQKLQAEAQERLKKEERRHAEAKDQLERALRAMQVAKDSLEHLASKLIHITVEDGRFAGKELDPQADNYVPNLLGLVEEKLLKLQAQLQGHDVQEMLCHIANREFLASLEGRLPEYNTRIALPLATSKDKFFDEESEEEDNEVVTRASLKIRSQKLIESHKKHRRSRRS | Component of the outer dynein arm-docking complex (ODA-DC) that mediates outer dynein arms (ODA) binding onto the doublet microtubule. Involved in mediating assembly of both ODAs and their axonemal docking complex onto ciliary microtubules. |
Generate the functional description in free-text form based on the protein sequence. | MAGAENWPGQQLELDEDEASCCRWGAQHAGARELAALYSPGKRLQEWCSVILCFSLIAHNLVHLLLLARWEDTPLVILGVVAGALIADFLSGLVHWGADTWGSVELPIVGKAFIRPFREHHIDPTAITRHDFIETNGDNCLVTLLPLLNMAYKFRTHSPEALEQLYPWECFVFCLIIFGTFTNQIHKWSHTYFGLPRWVTLLQDWHVILPRKHHRIHHVSPHETYFCITTGWLNYPLEKIGFWRRLEDLIQGLTGEKPRADDMKWAQKIK | Plasmanylethanolamine desaturase involved in plasmalogen biogenesis in the endoplasmic reticulum membrane. Plasmalogens are glycerophospholipids with a hydrocarbon chain linked by a vinyl ether bond at the glycerol sn-1 position, and are involved in antioxidative and signaling mechanisms. |
Predict the functional description of the protein sequence below. | MVLSLTGLIAFSFLQATLALNPEDPNVCSHWESYAVTVQESYAHPFDQIYYTRCTDILNWFKCTRHRISYKTAYRRGLRTMYRRRSQCCPGYYESGDFCIPLCTEECVHGRCVSPDTCHCEPGWGGPDCSSGCDSDHWGPHCSNRCQCQNGALCNPITGACVCAAGFRGWRCEELCAPGTHGKGCQLPCQCRHGASCDPRAGECLCAPGYTGVYCEELCPPGSHGAHCELRCPCQNGGTCHHITGECACPPGWTGAVCAQPCPPGTFGQNCSQDCPCHHGGQCDHVTGQCHCTAGYMGDRCQEECPFGSFGFQCSQHCDCHNGGQCSPTTGACECEPGYKGPRCQERLCPEGLHGPGCTLPCPCDADNTISCHPVTGACTCQPGWSGHHCNESCPVGYYGDGCQLPCTCQNGADCHSITGGCTCAPGFMGEVCAVSCAAGTYGPNCSSICSCNNGGTCSPVDGSCTCKEGWQGLDCTLPCPSGTWGLNCNESCTCANGAACSPIDGSCSCTPGWLGDTCELPCPDGTFGLNCSEHCDCSHADGCDPVTGHCCCLAGWTGIRCDSTCPPGRWGPNCSVSCSCENGGSCSPEDGSCECAPGFRGPLCQRICPPGFYGHGCAQPCPLCVHSSRPCHHISGICECLPGFSGALCNQVCAGGYFGQDCAQLCSCANNGTCSPIDGSCQCFPGWIGKDCSQACPPGFWGPACFHACSCHNGASCSAEDGACHCTPGWTGLFCTQRCPAAFFGKDCGRVCQCQNGASCDHISGKCTCRTGFTGQHCEQRCAPGTFGYGCQQLCECMNNSTCDHVTGTCYCSPGFKGIRCDQAALMMEELNPYTKISPALGAERHSVGAVTGIMLLLFLIVVLLGLFAWHRRRQKEKGRDLAPRVSYTPAMRMTSTDYSLSGACGMDRRQNTYIMDKGFKDYMKESVCSSSTCSLNSSENPYATIKDPPILTCKLPESSYVEMKSPVHMGSPYTDVPSLSTSNKNIYEVEPTVSVVQEGCGHNSSYIQNAYDLPRNSHIPGHYDLLPVRQSPANGPSQDKQS | May regulate the mosaic spacing of specific neuron subtypes in the retina through homotypic retinal neuron repulsion. Mosaics provide a mechanism to distribute each cell type evenly across the retina, ensuring that all parts of the visual field have access to a full set of processing elements (By similarity). |
Predict the functional description of the protein sequence below. | MFRRARLSVKPNVRPGVGARGSTASNPQRGRESPRPPDPATDSASKPAEPTDVPTVDFGGAEPQEKAPRSSTEKTGGDNDVEESSRSSSTVSQRRKRISSTSSLVKSSVSVPSESHPLSTINQEAPQPTATSTKEKQPCSDRYRIYKAQKLREMLKEELRKEKKQWKNKYAINESQRPPDRSKMTMRDFIYYLPDNNPMTSSLEQEKKTEKPSTPVQTREQEGKSTPNAEDNEMEEETDDGPLLVPRVKVAEDGSIILDEESLTVEVLRTKGPCVVEENDPIFERGSTTTYSSFRKNYYSKPWSNKETDMFFLAISMVGTDFSMIGQLFPHRARIEIKNKFKREEKTNGWRIDKAFQEKRPFDFDFFAHLLQKVLAEEEKRKQKSVKNHSLKEKKSTKPRKNVKVKKVACEGVNNDPDESMSSRISDTERSQKDAQTVEEESLTLSREDAEQVALEVDLNQKKRRRKKQDGANELGVNNLLENATVQAGPSKGEKHKNKCQAIRPELKEGECSKEQMLSCTQNIDGIVGFASTEKVEKRTDPILSLSNQQDATSVATESSESSTSDLPSFEVGIRALCEVNNAEGSCIEERNVDLKNNSLEIDQTENVKPMLRGRFQRPKPNLSRAGKKSVLSQGKTESESKNSHSKTSVEKNHVEKDKMNTLDILRMETTERENPEAETVSVLGEKNCLQEGSQLKALRPVQVRGRLQKPKPNAGKAAERKEILISQEEIGANVEKNENESCADRDTPQHMEDQSRKDFEEEDVILQPEKNDSFQNVQPDEPKVLNECLSVQENNKANKLNQVPILRTRFQKPKPNIGRGTGRREISSKEEVLEKILVSGEMAAALRETVRLDTSPKEMVPAEINTKEMQSDLKETGRRAISPREKILDVIDDTIEMETGLKAMGREICLREKTPEVIDATEEIDKDLEEAGRREISPQKNGPEEVKPLGEVETDLKATGNESSPREKTPEVTDATEEIDKNLEETGRRKISPRENGPEEVKPVDEMETDLNATGRESSPREKTPEVIDATEEIDLEETEREVSPQENGLEEVKPLGEMETDLKATGRDSFPRGKTPEVIDAIEEIEIDLEETEREISPQENGLEEVKPLGEMQTDLKATGREISPREKTPEVIDATEEIDKDLEETGRREISPEENGPEEVKPVDEMETDLKTTGREGSSREKTREVIDAAEVIETDLEETEREISPQENGPEEVKPVGKMETDLKEIREEISQREKVLAEFSAIREKEIDLKETGKRDIPIMEKVSGKMAVVEEMEADLKETGKENFRERGSEEICVTEEKVAELKQTGKTDISPRENELEETSTSRQTDTHLMQSGSNDFSAVPSLDIQNISSEVLSMMHTPVEEKRNSEKEVSSHFSHFKISSQTHESDKTEVQGIQSPDVPEQFSDINLSKSLPQEQKPLEIKPAPFVRSRFKRPKPNLARAALKRETTESEKYIYEKKSETKKMETIVMQENNEQTDTLPSQHDEASLMISREKDTLGHRNEEAVILPCTQTERNLSPSNSCEPKEESQSAPVQKNDSVVSVGTNNVNTFQQEMKESVIQTARQVRGRLQRPRPNIRKTGQRQIVDKGEAKGIIKEGRTILPKDETEKKVLTVSNSQIETEIEVPSSAVPEHRMYENQSQVVLVENLHVNKTNETIRHENKPYVPSSAQMTRRKFQKAKPNLGRAHSKKEEPVLEKVTTDQSKEGKPEDHLLQKGASNTQLLLKEKAELLTSLEVSARKDCVGSKESALAKIDAELEEVGPSRRVGEETVGDNSPSSVVEEQYLNKLTSCPQPLNETSYSKIALDGKTTISSTSEYERNRGERRSHKKFKPNVTRGRGSKRVRGKTSKKEPRASKAMLVTLRASQEEDDDADDFESDYEEESYHLAPEEVNKAPVFVPVGLRSPEPVSAQIEETMEELEITVNVPDVGCIAVVEHELPNTDVTTEEMKQEENLSVPFEMTTSEHIQDEPGTNDGSTEAAITLLTMGDLVLQSEISSEQGDVGVCIIPHVHSKDKSHIPSSLDNVNHKIVHECQELSSPVITTSPASFEENKIVLEEQSSREEISLMEKVKENATPTRNTISKVTSNLRIRSRLAKPKPNLEKTLGTNRLDDYQEVSSLCVTKGAEMETQRETEKNASKATELENKNLGPVTTAENKDQSKLACVHGIKGTSISSEVNLTERNENQEESSQEVHMLSVAPVASSETGPCTLGLDRGLGENSVEEPQIKDSKGDSVLTLPVPEYTPTSIPEVQQENIINPQDLTVNLVANVPQDGEDEQAFILTLVEIPANAVEEFTDATAQFMPNPLLPAPILVKSVNTEERGDMSICLPATSVGQDAMGLSISGRDNSKKPPDNLDLVSRKRFQCRLDKNDHIPPAKKRSLTLRDDCQEYTTEVHSKELTNVFEETGESHKGQDIFLTSGSTLTTPEPQRQQVEAAFQSRGSRSPDACMDKNVPQLPQDEMIVSDKEERTDAAPKSQQMDSRTSSSKASLSRPGRRPLGFLSLICSKNSLESDEPMQVHSKKRLKPLIPGLRKKLKRSNPFNESQEKNRESSDLLPSPSVITTQSENISSSATQVSCDQPLLKEGYKSAQKRAPQGEATTVSEYFFNDIFIEVDETE | General activator of RNA polymerase III transcription. Requires for transcription from all three types of polymerase III promoters. Requires for transcription of genes with internal promoter elements and with promoter elements upstream of the initiation site. |
Generate the functional description in free-text form based on the protein sequence. | MLLSPVTSTPFSVKDILRLERERSCPAASPHPRVRKSPENFQYLRMDAEPRGSEVHNAGGGGGDRKLDGSEPPGGPCEAVLEMDAERMGEPQPGLNAASPLGGGTRVPERGVGNSGDSVRGGRSEQPKARQRRKPRVLFSQAQVLALERRFKQQRYLSAPEREHLASALQLTSTQVKIWFQNRRYKCKRQRQDKSLELAGHPLTPRRVAVPVLVRDGKPCLGPGPGAPAFPSPYSAAVSPYSCYGGYSGAPYGAGYGTCYAGAPSGPAPHTPLASAGFGHGGQNATPQGHLAATLQGVRAW | Acts as a transcriptional activator. In conjunction with NKX2-5, may play a role in both pharyngeal and cardiac embryonic development. |
What is the functional description of the protein sequence? | MSRHSRLQRQVLSLYRDLLRAGRGKPGAEARVRAEFRQHAGLPRSDVLRIEYLYRRGRRQLQLLRSGHATAMGAFVRPRAPTGEPGGVGCQPDDGDSPRNPHDSTGAPETRPDGR | Plays an essential role in the assembly of succinate dehydrogenase (SDH), an enzyme complex (also referred to as respiratory complex II) that is a component of both the tricarboxylic acid (TCA) cycle and the mitochondrial electron transport chain, and which couples the oxidation of succinate to fumarate with the reduction of ubiquinone (coenzyme Q) to ubiquinol. Promotes maturation of the iron-sulfur protein subunit SDHB of the SDH catalytic dimer, protecting it from the deleterious effects of oxidants. May act together with SDHAF3. Contributes to iron-sulfur cluster incorporation into SDHB by binding to SDHB and recruiting the iron-sulfur transfer complex formed by HSC20, HSPA9 and ISCU through direct binding to HSC20. |
Predict the functional description of the protein sequence below. | MGCTPSHSDLVNSVAKSGIQFLKKPKAIRPGCQGGSERGSIPLLVKNSTCYDAGEGLAEEQPSPRRNQTTAKGLCQLMGDPASGKRKDMEGLIPGTKTSSSQLNKSQSHMAKDIPFKTQGSHGSQGADFSGDESEESSTQDTSKWKRTAKCHTSSTQSHCYQTIHPAHEPEGKVDFPEPLVKAHQQAYTYLHSSLSKYEAILCIIHQATQTRELLQPMVSFLLLCFEEISQLLGEISKDGEVLLQEVREDLAWPLKKREPQEQPNLLQQLLQYTVSKLQVLNGTVASLTGSFLEGSSSYLHSTATHLENKLSTKRNVDERLLRALRQLESLASGCGDPGVQGLPLCSEDSGIGADNESVQSVDKLGKQTSWDLAPEPEEWKSVTSPHTEARQSGHTWQQSPFCLGSGRPQDCLLSGAPMAKVQPRAQDEARSPCLSSTSPENITSPPLKLGTSTPCDSFGIGVSVEPHLSKTSRPMDASSLSDSEDSSPEEEEEDKMSSMSLCAWQEKTPHSRPQSSPADRESPFQARTRRLRSLQAQEMILKMKESISERIKFVPVPCGHQDWSEEEEGRTVVPPRPSTVSGSRRAPERQTRSQSESCLQSHVEDPTFQELRRVQRDLSQKLEAFYALGAKGQGQSQEQILQPRAAAVWPNGTCRVSPSNTTSRLKASLTKNFSILPSQDKSILQKCNPHPEDEQGKAGKLPNAIPSGEVSEAAKATDWNVRGCPTRTSVKKLIETFSPTESLRMLGDSKDAGASPCLRNCIMPPRFPKYTGLAPLYPKPQISPASGRESLKMGIGWKPLAPIFPPLPKAEAAKSEELSCEMEGNLEHLPPPPMEVLMDKSFASLESPESSKSTENSPKETQEPGPGEAGPTRRTWASPKLRASVSPLDLLPSKSTASLTKPHSTGPGSGRSSCQPRKPALDLSSPPATSQSPEVKGGTWSQAEKATSLYRQPRKAIAWHHSGPPSGQNRTSESSLARPRQSRERSPPVGRKASPTRTHWVPQADKRRRSLPSSYRPAQPSPSAVQTPPSPPVSPRVLSPPTTKRRTSPPHQPKLPNPPPESAPAQCKVPSPPTQHPEASPPFSIPSPSPPMSPSQEHKETRDSEDSQAVIAKVSGNTHSIFCPATSSLFEAKPPLSTAHPLTPPSLPPEAGGPLGNPAECWKNSSGPWLRADSQRRAALCALNPLPFLRRTASDRQPGGRPQPPTLDPTSTSYESQLGQNSSSEESPKKDTEPGSSPCSPELQGGTRRASPPEFCVLGHGLQPEPRTGHIQDKSQPEAQPQQEEVS | Plays an essential role for normal photoreceptor cell maintenance and vision. |
What is the functional description of the protein sequence? | MGTLVAKLLLPTLSSLAFLPTVSIAAKRRFHMEAMVYLFTLFFVALHHACNGPGLSVLCFMRHDILEYFSVYGTALSMWVSLMALADFDEPKRSTFVMFGVLTIAVRIYHDRWGYGVYSGPIGTAILIIAAKWLQKMKEKKGLYPDKSVYTQQIGPGLCFGALALMLRFFFEDWDYTYVHSFYHCALAMSFVLLLPKVNKKAGSPGTPAKLDCSTLCCACV | Myoblast-specific protein that mediates myoblast fusion, an essential step for the formation of multi-nucleated muscle fibers. Actively participates in the membrane fusion reaction by mediating the mixing of cell membrane lipids (hemifusion) upstream of MYMX. Acts independently of MYMX (By similarity). Involved in skeletal muscle regeneration in response to injury by mediating the fusion of satellite cells, a population of muscle stem cells, with injured myofibers (By similarity). Also involved in skeletal muscle hypertrophy, probably by mediating the fusion of satellite cells with myofibers (By similarity). |
Predict the general functional description of the protein sequence. | MVSQVLQLLRQGVWAALTGGWYHDPEQSKFTNSCHLYLWLFLLLLPLALHLAFPPNAIIVFFYCSAVTIFFTIIKLVSYRLHLMFDKGEVIQQKPSRKEEKPNKDKEAKGEHITNHRNPSNNRQIHNGKKEEASRNLSTPPLRCSSRGQSITSHHSSGPLELSAQETVEDLKGVILLEDHPIAPVSSTSPGIKVESLPASQAHMLETTTKSVIPVKPVATETLINGKGKERGGKGQPPLRHRSEGGLVDKGPLKKLPHLSLSQYDLLETDVSFQPWGSENSVLIPEPVSCPRGSIRERVQSKSPQDSLSSSCPQCDTIVAKPVEEPADTSCQVDTSCQGDLPLHQEVDSSDSEVAVTLIDTSQPGDPLSLHEPIKIVITMSSTPNSMTDLESSLHLRVVGTEKTSVKSDAEPTNPGAAGSPNAEQISIPVITLDLPEGGGGGVPCPEGNGSERTPERLKTRVSTNQCSGYGSGEGGNAIKDHSSSSREPWESVSRLTPDTGSESKVGKEGQTNLDPSSCKSSHEKRHARVLSVDSGTDVFLSKSSAEIVNDTEKTMPTSKSDLEAKEGQMPNESNFLEFVSLLESINTSKMTASSQLNGSAEQNEESGLLRDNCSQEKKEEILENEKPSGHSSKQGKPDLQSQDHTSTGPACTQPAKTTAFFQGNRQRQIIYRVTSQQDSSVLQVISGPETSVQEEISVDAMHVFIDEHGEIRSCYLKSGNQKEGPLQPLPSNNDCLSQAREMQVSSSSTTTSESQDPSSGDPAVSALQQQLLLMVARRTQSETPRHVSQDLEASSCSSTQGKFNREQFYKFIIFPGKWIKVWYDRLTLLALLDRTEDIKENVLAILLIVLVSLLGFLTLSQGFCKDMWVLLFCLVMASCQYSLLKSVQPDPASPIHGHNQIITYSRPIYFCVLCGLILLLDTGAKARHPPSYVVYGLKLFSPVFLQSARDYLIVFLYCFPAISLLGLFPQINTFCTYLLEQIDMLFFGGSAVSGITSAVYSVARSVLAAALLHAVCFSAVKEPWSMQHIPALFSAFCGLLVALSYHLSRQSSDPSVLMSFIQCRLFPKFLHQNLAESAADPLPKKMKDSVTDVLKWDLIVCAVVAVLSFAVSASTVFLSLRPFLSIVLFALAGAVGFVTHYVLPQLRKHHPWMWISHPILKNKEYHQREVRDVAHLMWFERLYVWLQCFEKYILYPALILNALTIDAFLISNHRRLGTHWDIFLMIIAGMKLLRTSFCNPVYQFINLSFTVIFFHFDYKDISESFLLDFFMVSILFSKLGDLLHKLQFVLTYVAPWQMAWGSSFHVFAQLFAIPHSAMLFFQTIATSIFSTPLSPFLGSVIFITSYVRPVKFWEKNYNTRRVDNSNTRLAVQIERDPGNDDNNLNSIFYEHLTRTLQESLCGDLVLGRWGNYSSGDCFILASDDLNAFVHLIEIGNGLVTFQLRGLEFRGTYCQQREVEAIMEGDEEDRGCCCCKPGHLPHLLSCNAAFHLRWLTWEITQTQYILEGYSILDNNAATMLQVFDLRRILIRYYIKSIIYYMVTSPKLLSWIKNESLLKSLQPFAKWHYIERDLAMFNINIDDDYVPCLQGITRASFCNVYLEWIQHCARKRQEPSTTLDSDEDSPLVTLSFALCTLGRRALGTAAHNMAISLDSFLYGLHVLFKGDFRITARDEWVFADMDLLHKVVAPAIRMSLKLHQDQFTCPDEYEDPAVLYEAIQSFEKKVVICHEGDPAWRGAVLSNKEELLTLRHVVDEGADEYKVIMLHRSFLSFKVIKVNKECVRGLWAGQQQELIFLRNRNPERGSIQNNKQVLRNLINSSCDQPLGYPMYVSPLTTSYLGTHRQLKNIWGGPITLDRIRTWFWTKWVRMRKDCNARQHSGGNIEDVDGGGAPTTGGNNAPSGGSQESSAEQPRKGGAQHGVSSCEGTQRTGRRKGRSQSVQAHSALSQRPPMLSSSGPILESRQTFLQTSTSVHELAQRLSGSRLSLHASATSLHSQPPPVTTTGHLSVRERAEALIRSSLGSSTSSTLSFLFGKRSFSSALVISGLSAAEGGNTSDTQSSSSVNIVMGPSARAASQATRHLSEPCEPPDATEQGQLHDRCLAEAVADTLGVVCRRASQEDMGLDDTASQQSVSDEQ | May play a role in tumorigenesis of colorectal carcinomas with high microsatellite instability (MSI-H). |
Predict the general functional description of the protein sequence. | MDTAYPREDTRAPTPSKAGAHTALTLGAPHPPPRDHLIWSVFSTLYLNLCCLGFLALAYSIKARDQKVVGDLEAARRFGSKAKCYNILAAMWTLVPPLLLLGLVVTGALHLARLAKDSAAFFSTKFDDADYD | Required for normal bone mineralization. |
Can you provide the functional description of the following protein sequence? | MELMFAEWEDGERFSFEDSDRFEEDSLCSFISEAESLCQNWRGWRKQSAGPNSPTGGGGGGGSGGTRMRDGLVIPLVELSAKQVAFHIPFEVVEKVYPPVPEQLQLRIAFWSFPENEEDIRLYSCLANGSADEFQRGDQLFRMRAVKDPLQIGFHLSATVVPPQMVPPKGAYNVAVMFDRCRVTSCSCTCGAGAKWCTHVVALCLFRIHNASAVCLRAPVSESLSRLQRDQLQKFAQYLISELPQQILPTAQRLLDELLSSQSTAINTVCGAPDPTAGPSASDQSTWYLDESTLTDNIKKTLHKFCGPSPVVFSDVNSMYLSSTEPPAAAEWACLLRPLRGREPEGVWNLLSIVREMFKRRDSNAAPLLEILTDQCLTYEQITGWWYSVRTSASHSSASGHTGRSNGQSEVAAHACASMCDEMVTLWRLAVLDPALSPQRRRELCTQLRQWQLKVIENVKRGQHKKTLERLFPGFRPAVEACYFNWEEAYPLPGVTYSGTDRKLALCWARALPSRPGASRSGGLEESRDRPRPLPTEPAVRPKEPGTKRKGLGEGVPSSQRGPRRLSAEGGDKALHKMGPGGGKAKALGGAGSGSKGSAGGGSKRRLSSEDSSLEPDLAEMSLDDSSLALGAEASTFGGFPESPPPCPLHGGSRGPSTFLPEPPDTYEEDGGVYFSEGPEPPTASVGPPGLLPGDVCTQDDLPSTDESGNGLPKTKEAAPAVGEEDDDYQAYYLNAQDGAGGEEEKAEGGAGEEHDLFAGLKPLEQESRMEVLFACAEALHAHGYSSEASRLTVELAQDLLANPPDLKVEPPPAKGKKNKVSTSRQTWVATNTLSKAAFLLTVLSERPEHHNLAFRVGMFALELQRPPASTKALEVKLAYQESEVAALLKKIPLGPSEMSTMRCRAEELREGTLCDYRPVLPLMLASFIFDVLCAPGSRPPSRNWNSETPGDEELGFEAAVAALGMKTTVSEAEHPLLCEGTRREKGDLALALMITYKDDQAKLKKILDKLLDRESQTHKPQTLSSFYSSSRPTTASQRSPSKHGGPSAPGALQPLTSGSAGPAQPGSVAGAGPGPTEGFTEKNVPESSPHSPCEGLPSEAALTPRPEGKVPSRLALGSRGGYNGRGWGSPGRPKKKHTGMASIDSSAPETTSDSSPTLSRRPLRGGWAPTSWGRGQDSDSISSSSSDSLGSSSSSGSRRASASGGARAKTVEVGRYKGRRPESHAPHVPNQPSEAAAHFYFELAKTVLIKAGGNSSTSIFTHPSSSGGHQGPHRNLHLCAFEIGLYALGLHNFVSPNWLSRTYSSHVSWITGQAMEIGSAALTILVECWDGHLTPPEVASLADRASRARDSNMVRAAAELALSCLPHAHALNPNEIQRALVQCKEQDNLMLEKACMAVEEAAKGGGVYPEVLFEVAHQWFWLYEQTAGGSSTAREGATSCSASGIRAGGEAGRGMPEGRGGPGTEPVTVAAAAVTAAATVVPVISVGSSLYPGPGLGHGHSPGLHPYTALQPHLPCSPQYLTHPAHPAHPMPHMPRPAVFPVPSSAYPQGVHPAFLGAQYPYSVTPPSLAATAVSFPVPSMAPITVHPYHTEPGLPLPTSVACELWGQGTVSSVHPASTFPAIQGASLPALTTQPSPLVSGGFPPPEEETHSQPVNPHSLHHLHAAYRVGMLALEMLGRRAHNDHPNNFSRSPPYTDDVKWLLGLAAKLGVNYVHQFCVGAAKGVLSPFVLQEIVMETLQRLSPAHAHNHLRAPAFHQLVQRCQQAYMQYIHHRLIHLTPADYDDFVNAIRSARSAFCLTPMGMMQFNDILQNLKRSKQTKELWQRVSLEMATFSP | Substrate recognition component of a SCF-like E3 ubiquitin-protein ligase complex that promotes target-directed microRNA degradation (TDMD), a process that mediates degradation of microRNAs (miRNAs). The SCF-like E3 ubiquitin-protein ligase complex acts by catalyzing ubiquitination and subsequent degradation of AGO proteins (AGO1, AGO2, AGO3 and/or AGO4), thereby exposing miRNAs for degradation. Specifically recognizes and binds AGO proteins when they are engaged with a TDMD target. May also act as a regulator of axon guidance: specifically recognizes misfolded ROBO3 and promotes its ubiquitination and subsequent degradation. |
Can you provide the functional description of the following protein sequence? | MESMFEDDISILTQEALGPSEVWLDSPGDPSLGGDMCSASHFALITAYGDIKERLGGLERENATLRRRLKVYEIKYPLISDFGEEHGFSLYEIKDGSLLEVEKVSLQQRLNQFQHELQKNKEQEEQLGEMIQAYEKLCVEKSDLETELREMRALVETHLRQICGLEQQLRQQQGLQDAAFSNLSPPPAPAPPCTDLDLHYLALRGGSGLSHAGWPGSTPSVSDLERRRLEEALEAAQGEARGAQLREEQLQAECERLQGELKQLQETRAQDLASNQSERDMAWVKRVGDDQVNLALAYTELTEELGRLRELSSLQGRILRTLLQEQARSGGQRHSPLSQRHSPAPQCPSPSPPARAAPPCPPCQSPVPQRRSPVPPCPSPQQRRSPASPSCPSPVPQRRSPVPPSCQSPSPQRRSPVPPSCPAPQPRPPPPPPPGERTLAERAYAKPPSHHVKAGFQGRRSYSELAEGAAYAGASPPWLQAEAATLPKPRAYGSELYGPGRPLSPRRAFEGIRLRFEKQPSEEDEWAVPTSPPSPEVGTIRCASFCAGFPIPESPAATAYAHAEHAQSWPSINLLMETVGSDIRSCPLCQLGFPVGYPDDALIKHIDSHLENSKI | Adapter protein which constitutively binds TBK1 and IKBKE playing a role in antiviral innate immunity. |
What is the functional description of the protein sequence? | MASVQQGEKQLFEKFWRGTFKAVATPRPESIIVASITARKPLPRTEPQNNPVVPAQDGPSEKLGQHLATEPLGTNSWERDKTCRELGATRGHSASHDKDLTPPPSSRGKKKKKKSTRKKRRRSSSYSPSPVKKKKKKSSKKHKRRRSFSKKRRHSSSSPKSKRRDEKRHKKQSRSRPRKSHRHRHHRCPSRSQSSESRPSSCESRHRGRSPEEGQKSRRRHSRRCSKTLCKDSPEAQSSRPPSQPLQMLGYLSARGVITGSGSAADLFTKTASPLTTSRGRSQEYDSGNDTSSPPSTQTSSARSRGQEKGSPSGGLSKSRELNSGNTSDSGNSFTTSSPQNKGAMLENLSPTSRGRESRGFQSPCLECAEVKKSSLVPSTARSSPMKGCSRSSSYASTRSSSHSSRSPNPRASPRYTQSRSTSSEKRSYSRSPSYSSKSGKRSPPSRSSRSRRSPSYSRYSPSRERDPKYSEKDSQQRERERARRRRRSYSPMRKRRRDSPSHLEARRITSARKRPIPYYRPSPSSSGSLSSTSSWYSSSSSRSASRSYSRSRSRSRSRRRSRTRTSSSSSSRSPSPGSRSRSRSRSRSRSRSRSQSRSYSSADSYSSTRR | Splicing factor specifically required for neural cell differentiation. Acts in conjunction with nPTB/PTBP2 by binding directly to its regulated target transcripts and promotes neural-specific exon inclusion in many genes that function in neural cell differentiation. Required to promote the inclusion of neural-specific exon 10 in nPTB/PTBP2, leading to increased expression of neural-specific nPTB/PTBP2. Also promotes the inclusion of exon 16 in DAAM1 in neuron extracts (By similarity). Promotes alternative splicing of REST transcripts to produce REST isoform 3 (REST4) with greatly reduced repressive activity, thereby activating expression of REST targets in neural cells. Plays an important role during embryonic development as well as in the proper functioning of the adult nervous system. Regulates alternative splicing events in genes with important neuronal functions (By similarity). |
What is the functional description of the protein sequence? | MAEMEKEGRPPENKRSRKPAHPVKREINEEMKNFAENTMNELLGWYGYDKVELKDGEDIEFRSYPTDGESRQHISVLKENSLPKPKLPEDSVISPYNISTGYSGLATGNGLSDSPAGSKDHGSVPIIVPLIPPPFIKPPAEDDVSNVQIMCAWCQKVGIKRYSLSMGSEVKSFCSEKCFAACRRAYFKRNKARDEDGHAENFPQQHYAKETPRLAFKNNCELLVCDWCKHIRHTKEYLDFGDGERRLQFCSAKCLNQYKMDIFYKETQANLPAGLCSTLHPPMENKAEGTGVQLLTPDSWNIPLTDARRKAPSPVATAGQSQGPGPSASTTVSPSDTANCSVTKIPTPVPKSIPISETPNIPPVSVQPPASIGPPLGVPPRSPPMVMTNRGPVPLPIFMEQQIMQQIRPPFIRGPPHHASNPNSPLSNPMLPGIGPPPGGPRNLGPTSSPMHRPMLSPHIHPPSTPTMPGNPPGLLPPPPPGAPLPSLPFPPVSMMPNGPMPVPQMMNFGLPSLAPLVPPPTLLVPYPVIVPLPVPIPIPIPIPHVSDSKPPNGFSSNGENFIPNAPGDSAAAGGKPSGHSLSPRDSKQGSSKSADSPPGCSGQALSLAPTPAEHGRSEVVDLTRRAGSPPGPPGAGGQLGFPGVLQGPQDGVIDLTVGHRARLHNVIHRALHAHVKAEREPSAAERRTCGGCRDGHCSPPAAGDPGPGAPAGPEAAAACNVIVNGTRGAAAEGAKSAEPPPEQPPPPPPPAPPKKLLSPEEPAVSELESVKENNCASNCHLDGEAAKKLMGEEALAGGDKSDPNLNNPADEDHAYALRMLPKTGCVIQPVPKPAEKAAMAPCIISSPMLSAGPEDLEPPLKRRCLRIRNQNK | Implicated in development of the cochlea. |
Predict the functional description of the protein sequence below. | MDVQKWYLRTSKLALALAIIRSKPADKSSREYTEHLAMLLSEEQSKWRSKVEILEAEVMQLRQKLLVSRLCSGSFKSGYVSSQLEAQEPKSSESTLTSMEDSGCDLSNEQRTESSDLSQHFVESCTPTHFPPLPLVKRPCAILQNPLSSHMQFLQYLLELKNLTESGNLKRDLTHFEKDSSTVSDSVFQLLDGLITFYRNPKLPFSRFWTEAVGTLASLISDYNLSSHILKKCSKKLEEFEKTLLHAILGNNHINQFQVQHYVSQSLVTLGNCSLLRKSIISLLLSEVNGFADDLGAINQEQASYDVSRYENIFYLFWVLEQLLQKETEEGNTSSIGHDDQEIKKFLQKHDETIFQLSDAFPLFTFYLWRVGILLSSAQIETLRK | Required for DNA double-strand breaks (DSBs) formation in unsynapsed regions during meiotic recombination. Probably acts by forming a complex with IHO1 and REC114, which activates DSBs formation in unsynapsed regions, an essential step to ensure completion of synapsis. |
Can you provide the functional description of the following protein sequence? | MGEPSREEYKIQSFDAETQQLLKTALKDPGAVDLEKVANVIVDHSLQDCVFSKEAGRMCYAIIQAESKQAGQSVFRRGLLNRLQQEYQAREQLRARSLQGWVCYVTFICNIFDYLRVNNMPMMALVNPVYDCLFRLAQPDSLSKEEEVDCLVLQLHRVGEQLEKMNGQRMDELFVLIRDGFLLPTGLSSLAQLLLLEIIEFRAAGWKTTPAAHKYYYSEVSD | Functions in replication-dependent translation of histone mRNAs which differ from other eukaryotic mRNAs in that they do not end with a poly-A tail but a stem-loop. May participate in circularizing those mRNAs specifically enhancing their translation. |
Predict the functional description of the protein sequence below. | MALEQALQAARQGELDVLRSLHAAGLLGPSLRDPLDALPVHHAARAGKLHCLRFLVEEAALPAAARARNGATPAHDASATGHLACLQWLLSQGGCRVQDKDNSGATVLHLAARFGHPEVVNWLLHHGGGDPTAATDMGALPIHYAAAKGDFPSLRLLVEHYPEGVNAQTKNGATPLYLACQEGHLEVTQYLVQECGADPHARAHDGMTPLHAAAQMGHSPVIVWLVSCTDVSLSEQDKDGATAMHFAASRGHTKVLSWLLLHGGEISADLWGGTPLHDAAENGELECCQILVVNGAELDVRDRDGYTAADLSDFNGHSHCTRYLRTVENLSVEHRVLSRDPSAELEAKQPDSGMSSPNTTVSVQPLNFDLSSPTSTLSNYDSCSSSHSSIKGQHPPCGLSSARAADIQSYMDMLNPELGLPRGTIGKPTPPPPPPSFPPPPPPPGTQLPPPPPGYPAPKPPVGPQAADIYMQTKNKLRHVETEALKKELSSCDGHDGLRRQDSSRKPRAFSKQPSTGDYYRQLGRCPGETLAARPGMAHSEEVRARQPARAGCPRLGPAARGSLEGPSAPPQAALLPGNHVPNGCAADPKASRELPPPPPPPPPPLPEAASSPPPAPPLPLESAGPGCGQRRSSSSTGSTKSFNMMSPTGDNSELLAEIKAGKSLKPTPQSKGLTTVFSGIGQPAFQPDSPLPSVSPALSPVRSPTPPAAGFQPLLNGSLVPVPPTTPAPGVQLDVEALIPTHDEQGRPIPEWKRQVMVRKMQLKMQEEEEQRRKEEEEEARLASMPAWRRDLLRKKLEEEREQKRKEEERQKQEELRREKEQSEKLRTLGYDESKLAPWQRQVILKKGDIAKY | Multifunctional actin-bundling protein. Plays a major role in regulating the organization, dimension, dynamics and signaling capacities of the actin filament-rich microvilli in the mechanosensory and chemosensory cells. Required for the assembly and stabilization of the stereociliary parallel actin bundles. Plays a crucial role in the formation and maintenance of inner ear hair cell stereocilia (By similarity). Involved in the elongation of actin in stereocilia. In extrastriolar hair cells, required for targeting MYO3B to stereocilia tips, and for regulation of stereocilia diameter and staircase formation. |
What is the functional description of the protein sequence? | MALHIHEACILLLVIPGLVTSAAISHEDYPADEGDQISSNDNLIFDDYRGKGCVDDSGFVYKLGERFFPGHSNCPCVCALDGPVCDQPECPKIHPKCTKVEHNGCCPECKEVKNFCEYHGKNYKILEEFKPSPCEWCRCEPSNEVHCVVADCAVPECVNPVYEPEQCCPVCKNGPNCFAGTTIIPAGIEVKVDECNICHCHNGDWWKPAQCSKRECQGKQTV | May play a role in neurogenesis. May play a role in bone differentiation and matrix mineralization. |
Generate the functional description in free-text form based on the protein sequence. | MRLEELKRLQNPLEQVNDGKYSFENHQLAMDAENNIEKYPLNLQPLESKVKIIQRAWREYLQRQEPLGKRSPSPPSVSSEKLSSSVSMNTFSDSSTPDYREDGMDLGSDAGSSSSSSRASSQSNSTKVTPCSECKSSSSPGGSLDLVSALEDYEEPFPVYQKKVIDEWAPEEDGEEEEEEDERDQRGYRDDRSPAREPGDVSARTRSGGGGGRSATTAMPPPVPNGNLHQHDPQDLRHNGNVVVAGRPSCSRGPRRAIQKPQPAGGRRSGRGPAAGGLCLQPPDGGTCVPEEPPVPPMDWEALEKHLAGLQFREQEVRNQGQARTNSTSAQKNERESIRQKLALGSFFDDGPGIYTSCSKSGKPSLSSRLQSGMNLQICFVNDSGSDKDSDADDSKTETSLDTPLSPMSKQSSSYSDRDTTEEESESLDDMDFLTRQKKLQAEAKMALAMAKPMAKMQVEVEKQNRKKSPVADLLPHMPHISECLMKRSLKPTDLRDMTIGQLQVIVNDLHSQIESLNEELVQLLLIRDELHTEQDAMLVDIEDLTRHAESQQKHMAEKMPAK | May play a role in action potential conduction in myelinated cells through the organization of molecular complexes at nodes of Ranvier and axon initial segments. May also play a role in axon outgrowth and guidance (By similarity). |
Predict the general functional description of the protein sequence. | MIPKLLSLLCFRLCVGQGDTRGDGSLPKPSLSAWPSSVVPANSNVTLRCWTPARGVSFVLRKGGIILESPKPLDSTEGAAEFHLNNLKVRNAGEYTCEYYRKASPHILSQRSDVLLLLVTGHLSKPFLRTYQRGTVTAGGRVTLQCQKRDQLFVPIMFALLKAGTPSPIQLQSPAGKEIDFSLVDVTAGDAGNYSCMYYQTKSPFWASEPSDQLEILVTVPPGTTSSNYSLGNFVRLGLAAVIVVIMGAFLVEAWYSRNVSPGESEAFKPE | May act as receptor (By similarity). Negatively regulates TCR-mediated CD4(+) T cell proliferation and activation, possibly by binding an unknown ligand on the T cell surface. Enhances Toll-like receptor-mediated production of pro-inflammatory cytokines by macrophages and neutrophils (By similarity). |
Can you provide the functional description of the following protein sequence? | MTAASRANPYSIVSLEEDGLHLVTMSGANGFGNGKVHTRRRCRNRFVKKNGQCNIAFANMDEKSQRYLADMFTTCVDIRWRYMLLIFSLAFLASWLLFGVIFWVIAVAHGDLEPAEGHGRTPCVMQVHGFMAAFLFSIETQTTIGYGLRCVTEECLVAVFMVVAQSIVGCIIDSFMIGAIMAKMARPKKRAQTLLFSHNAVVALRDGKLCLMWRVGNLRKSHIVEAHVRAQLIKPRVTEEGEYIPLDQIDIDVGFDKGLDRIFLVSPITILHEIDEASPLFGISRQDLETDDFEIVVILEGMVEATAMTTQARSSYLANEILWGHRFEPVLFEEKNQYKIDYSHFHKTYEVPSTPRCSAKDLVENKFLLPSANSFCYENELAFLSRDEEDEADGDQDGRSRDGLSPQARHDFDRLQAGGGVLEQRPYRRGSEI | Inward rectifier potassium channels are characterized by a greater tendency to allow potassium to flow into the cell rather than out of it. Their voltage dependence is regulated by the concentration of extracellular potassium; as external potassium is raised, the voltage range of the channel opening shifts to more positive voltages. The inward rectification is mainly due to the blockage of outward current by internal magnesium. |
Generate the functional description in free-text form based on the protein sequence. | MTSSPVSRVVYNGKRTSSPRSPPSSSEIFTPAHEENVRFIYEAWQGVERDLRGQVPGGERGLVEEYVEKVPNPSLKTFKPIDLSDLKRRSTQDAKKS | The phosphorylation status of MCRIP1 functions as a molecular switch to regulate epithelial-mesenchymal transition. Unphosphorylated MCRIP1 binds to and inhibits the transcriptional corepressor CTBP(s). When phosphorylated by MAPK/ERK, MCRIP1 releases CTBP(s) resulting in transcriptional silencing of the E-cadherin gene and induction of epithelial-mesenchymal transition. |
Generate the functional description in free-text form based on the protein sequence. | MATFVELSTKAKMPIVGLGTWRSLLGKVKEAVKVAIDAEYRHIDCAYFYENQHEVGEAIQEKIQEKAVMREDLFIVSKVWPTFFERPLVRKAFEKTLKDLKLSYLDVYLIHWPQGFKTGDDFFPKDDKGNMISGKGTFLDAWEAMEELVDEGLVKALGVSNFNHFQIERLLNKPGLKYKPVTNQVECHPYLTQEKLIQYCHSKGITVTAYSPLGSPDRPWAKPEDPSLLEDPKIKEIAAKHKKTTAQVLIRFHIQRNVTVIPKSMTPAHIVENIQVFDFKLSDEEMATILSFNRNWRAFDFKEFSHLEDFPFDAEY | [Isoform 1]: Catalyzes the NADPH-dependent reduction of a variety of carbonyl substrates, like aromatic aldehydes, alkenals, ketones and alpha-dicarbonyl compounds. In addition, catalyzes the reduction of androgens and estrogens with high positional selectivity (shows 17-beta-hydroxysteroid dehydrogenase activity) as well as 3-keto-acyl-CoAs. Displays strong enzymatic activity toward all-trans-retinal and 9-cis-retinal. May play a physiological role in retinoid metabolism. |
Generate the functional description in free-text form based on the protein sequence. | MENEDGYMTLSFKNRCKSKQKSKDFSLYPQYYCLLLIFGCIVILIFIMTGIDLKFWHKKMDFSQNVNVSSLSGHNYLCPNDWLLNEGKCYWFSTSFKTWKESQRDCTQLQAHLLVIQNLDELEFIQNSLKPGHFGWIGLYVTFQGNLWMWIDEHFLVPELFSVIGPTDDRSCAVITGNWVYSEDCSSTFKGICQRDAILTHNGTSGV | C-type lectin-like receptor involved in natural killer cell mediated cytotoxicity and cytokine secretion in keratinocytes via its interaction with CLEC2A. |
Can you provide the functional description of the following protein sequence? | MLQFLLGFTLGNVVGMYLAQNYDIPNLAKKLEEIKKDLDAKKKPPSA | Microprotein involved in mitochondrial respiratory chain complex III (ubiquinol-cytochrome c oxidoreductase) and complex IV (mitochondrial cytochrome c oxidase complex) assembly. Required for the formation of mitochondrial supercomplexes (SCs). Also required for the activation of the NLRP3 inflammasome (By similarity). |
Can you provide the functional description of the following protein sequence? | MIPLLLAALLCVPAGALTCYGDSGQPVDWFVVYKLPALRGSGEAAQRGLQYKYLDESSGGWRDGRALINSPEGAVGRSLQPLYRSNTSQLAFLLYNDQPPQPSKAQDSSMRGHTKGVLLLDHDGGFWLVHSVPNFPPPASSAAYSWPHSACTYGQTLLCVSFPFAQFSKMGKQLTYTYPWVYNYQLEGIFAQEFPDLENVVKGHHVSQEPWNSSITLTSQAGAVFQSFAKFSKFGDDLYSGWLAAALGTNLQVQFWHKTVGILPSNCSDIWQVLNVNQIAFPGPAGPSFNSTEDHSKWCVSPKGPWTCVGDMNRNQGEEQRGGGTLCAQLPALWKAFQPLVKNYQPCNGMARKPSRAYKI | Hydrolyzes DNA under acidic conditions with a preference for double-stranded DNA. Plays a major role in the clearance of nucleic acids generated through apoptosis, hence preventing autoinflammation. Necessary for proper fetal development and for definitive erythropoiesis in fetal liver and bone marrow, where it degrades nuclear DNA expelled from erythroid precursor cells. |
Predict the general functional description of the protein sequence. | MAEAAAAAGGTGLGAGASYGSAADRDRDPDPDRAGRRLRVLSGHLLGRPREALSTNECKARRAASAATAAPTATPAAQESGTIPKKRQEVMKWNGWGYNDSKFIFNKKGQIELTGKRYPLSGMGLPTFKEWIQNTLGVNVEHKTTSKASLNPSDTPPSVVNEDFLHDLKETNISYSQEADDRVFRAHGHCLHEIFLLREGMFERIPDIVLWPTCHDDVVKIVNLACKYNLCIIPIGGGTSVSYGLMCPADETRTIISLDTSQMNRILWVDENNLTAHVEAGITGQELERQLKESGYCTGHEPDSLEFSTVGGWVSTRASGMKKNIYGNIEDLVVHIKMVTPRGIIEKSCQGPRMSTGPDIHHFIMGSEGTLGVITEATIKIRPVPEYQKYGSVAFPNFEQGVACLREIAKQRCAPASIRLMDNKQFQFGHALKPQVSSIFTSFLDGLKKFYITKFKGFDPNQLSVATLLFEGDREKVLQHEKQVYDIAAKFGGLAAGEDNGQRGYLLTYVIAYIRDLALEYYVLGESFETSAPWDRVVDLCRNVKERITRECKEKGVQFAPFSTCRVTQTYDAGACIYFYFAFNYRGISDPLTVFEQTEAAAREEILANGGSLSHHHGVGKLRKQWLKESISDVGFGMLKSVKEYVDPNNIFGNRNLL | Catalyzes the exchange of the acyl chain in acyl-dihydroxyacetonephosphate (acyl-DHAP) for a long chain fatty alcohol, yielding the first ether linked intermediate, i.e. alkyl-dihydroxyacetonephosphate (alkyl-DHAP), in the pathway of ether lipid biosynthesis. |
Predict the general functional description of the protein sequence. | MALQVELVPTGEIIRVVHPHRPCKLALGSDGVRVTMESALTARDRVGVQDFVLLENFTSEAAFIENLRRRFRENLIYTYIGPVLVSVNPYRDLQIYSRQHMERYRGVSFYEVPPHLFAVADTVYRALRTERRDQAVMISGESGAGKTEATKRLLQFYAETCPAPERGGAVRDRLLQSNPVLEAFGNAKTLRNDNSSRFGKYMDVQFDFKGAPVGGHILSYLLEKSRVVHQNHGERNFHIFYQLLEGGEEETLRRLGLERNPQSYLYLVKGQCAKVSSINDKSDWKVVRKALTVIDFTEDEVEDLLSIVASVLHLGNIHFAANEESNAQVTTENQLKYLTRLLSVEGSTLREALTHRKIIAKGEELLSPLNLEQAAYARDALAKAVYSRTFTWLVGKINRSLASKDVESPSWRSTTVLGLLDIYGFEVFQHNSFEQFCINYCNEKLQQLFIELTLKSEQEEYEAEGIAWEPVQYFNNKIICDLVEEKFKGIISILDEECLRPGEATDLTFLEKLEDTVKHHPHFLTHKLADQRTRKSLGRGEFRLLHYAGEVTYSVTGFLDKNNDLLFRNLKETMCSSKNPIMSQCFDRSELSDKKRPETVATQFKMSLLQLVEILQSKEPAYVRCIKPNDAKQPGRFDEVLIRHQVKYLGLLENLRVRRAGFAYRRKYEAFLQRYKSLCPETWPTWAGRPQDGVAVLVRHLGYKPEEYKMGRTKIFIRFPKTLFATEDALEVRRQSLATKIQAAWRGFHWRQKFLRVKRSAICIQSWWRGTLGRRKAAKRKWAAQTIRRLIRGFVLRHAPRCPENAFFLDHVRTSFLLNLRRQLPQNVLDTSWPTPPPALREASELLRELCIKNMVWKYCRSISPEWKQQLQQKAVASEIFKGKKDNYPQSVPRLFISTRLGTDEISPRVLQALGSEPIQYAVPVVKYDRKGYKPRSRQLLLTPNAVVIVEDAKVKQRIDYANLTGISVSSLSDSLFVLHVQRADNKQKGDVVLQSDHVIETLTKTALSANRVNSININQGSITFAGGPGRDGTIDFTPGSELLITKAKNGHLAVVAPRLNSR | Myosins are actin-based motor molecules with ATPase activity. Unconventional myosins serve in intracellular movements. Their highly divergent tails are presumed to bind to membranous compartments, which would be moved relative to actin filaments. Involved in glucose transporter recycling in response to insulin by regulating movement of intracellular GLUT4-containing vesicles to the plasma membrane. Component of the hair cell's (the sensory cells of the inner ear) adaptation-motor complex. Acts as a mediator of adaptation of mechanoelectrical transduction in stereocilia of vestibular hair cells. Binds phosphoinositides and links the actin cytoskeleton to cellular membranes. |
Generate the functional description in free-text form based on the protein sequence. | MDNLSSEEIQQRAHQITDESLESTRRILGLAIESQDAGIKTITMLDEQKEQLNRIEEGLDQINKDMRETEKTLTELNKCCGLCVCPCNRTKNFESGKAYKTTWGDGGENSPCNVVSKQPGPVTNGQLQQPTTGAASGGYIKRITNDAREDEMEENLTQVGSILGNLKDMALNIGNEIDAQNPQIKRITDKADTNRDRIDIANARAKKLIDS | Essential component of the high affinity receptor for the general membrane fusion machinery and an important regulator of transport vesicle docking and fusion. |
Predict the general functional description of the protein sequence. | MSLFDLFRGFFGFPGPRSHRDPFFGGMTRDEDDDEEEEEEGGSWGRGNPRFHSPQHPPEEFGFGFSFSPGGGIRFHDNFGFDDLVRDFNSIFSDMGAWTLPSHPPELPGPESETPGERLREGQTLRDSMLKYPDSHQPRIFGGVLESDARSESPQPAPDWGSQRPFHRFDDVWPMDPHPRTREDNDLDSQVSQEGLGPVLQPQPKSYFKSISVTKITKPDGIVEERRTVVDSEGRTETTVTRHEADSSPRGDPESPRPPALDDAFSILDLFLGRWFRSR | Recruits the Arp2/3 complex to the cell cortex and regulates reorganization of the cortical actin cytoskeleton via its interaction with KCNC3 and the Arp2/3 complex. Slows down the rate of inactivation of KCNC3 channels. Promotes GNA13-mediated cell migration. Involved in the clathrin-mediated endocytosis pathway. May be involved in internalization of ABC transporters such as ABCB11. May inhibit CASP9 and CASP3. Promotes cell survival. May regulate intracellular calcium pools. |
Predict the general functional description of the protein sequence. | MASLGHILVFCVGLLTMAKAESPKEHDPFTYDYQSLQIGGLVIAGILFILGILIVLSRRCRCKFNQQQRTGEPDEEEGTFRSSIRRLSTRRR | Associates with and regulates the activity of the sodium/potassium-transporting ATPase (NKA) which transports Na(+) out of the cell and K(+) into the cell. Inhibits NKA activity in its unphosphorylated state and stimulates activity when phosphorylated. Reduces glutathionylation of the NKA beta-1 subunit ATP1B1, thus reversing glutathionylation-mediated inhibition of ATP1B1. Contributes to female sexual development by maintaining the excitability of neurons which secrete gonadotropin-releasing hormone. |
Predict the functional description of the protein sequence below. | MMLSLNNLQNIIYNPVIPFVGTIPDQLDPGTLIVIRGHVPSDADRFQVDLQNGSSMKPRADVAFHFNPRFKRAGCIVCNTLINEKWGREEITYDTPFKREKSFEIVIMVLKDKFQVAVNGKHTLLYGHRIGPEKIDTLGIYGKVNIHSIGFSFSSDLQSTQASSLELTEISRENVPKSGTPQLRLPFAARLNTPMGPGRTVVVKGEVNANAKSFNVDLLAGKSKDIALHLNPRLNIKAFVRNSFLQESWGEEERNITSFPFSPGMYFEMIIYCDVREFKVAVNGVHSLEYKHRFKELSSIDTLEINGDIHLLEVRSW | Beta-galactoside-binding lectin that acts as a sensor of membrane damage caused by infection and restricts the proliferation of infecting pathogens by targeting them for autophagy. Detects membrane rupture by binding beta-galactoside ligands located on the lumenal side of the endosome membrane; these ligands becoming exposed to the cytoplasm following rupture. Restricts infection by initiating autophagy via interaction with CALCOCO2/NDP52. Required to restrict infection of bacterial invasion such as S.typhimurium. Also required to restrict infection of Picornaviridae viruses. Has a marked preference for 3'-O-sialylated and 3'-O-sulfated glycans. |
Can you provide the functional description of the following protein sequence? | MKALIFAAAGLLLLLPTFCQSGMENDTNNLAKPTLPIKTFRGAPPNSFEEFPFSALEGWTGATITVKIKCPEESASHLHVKNATMGYLTSSLSTKLIPAIYLLVFVVGVPANAVTLWMLFFRTRSICTTVFYTNLAIADFLFCVTLPFKIAYHLNGNNWVFGEVLCRATTVIFYGNMYCSILLLACISINRYLAIVHPFTYRGLPKHTYALVTCGLVWATVFLYMLPFFILKQEYYLVQPDITTCHDVHNTCESSSPFQLYYFISLAFFGFLIPFVLIIYCYAAIIRTLNAYDHRWLWYVKASLLILVIFTICFAPSNIILIIHHANYYYNNTDGLYFIYLIALCLGSLNSCLDPFLYFLMSKTRNHSTAYLTK | Receptor for activated thrombin coupled to G proteins that stimulate phosphoinositide hydrolysis. |
What is the functional description of the protein sequence? | MAAGSDLLDEVFFNSEVDEKVVSDLVGSLESQLAASAAHHHHLAPRTPEVRAAAAGALGNHVVSGSPAGAAGAGPAAPAEGAPGAAPEPPPAGRARPGGGGPQRPGPPSPRRPLVPAGPAPPAAKLRPPPEGSAGSCAPVPAAAAVAAGPEPAPAGPAKPAGPAALAARAGPGPGPGPGPGPGPGPGKPAGPGAAQTLNGSAALLNSHHAAAPAVSLVNNGPAALLPLPKPAAPGTVIQTPPFVGAAAPPAPAAPSPPAAPAPAAPAAAPPPPPPAPATLARPPGHPAGPPTAAPAVPPPAAAQNGGSAGAAPAPAPAAGGPAGVSGQPGPGAAAAAPAPGVKAESPKRVVQAAPPAAQTLAASGPASTAASMVIGPTMQGALPSPAAVPPPAPGTPTGLPKGAAGAVTQSLSRTPTATTSGIRATLTPTVLAPRLPQPPQNPTNIQNFQLPPGMVLVRSENGQLLMIPQQALAQMQAQAHAQPQTTMAPRPATPTSAPPVQISTVQAPGTPIIARQVTPTTIIKQVSQAQTTVQPSATLQRSPGVQPQLVLGGAAQTASLGTATAVQTGTPQRTVPGATTTSSAATETMENVKKCKNFLSTLIKLASSGKQSTETAANVKELVQNLLDGKIEAEDFTSRLYRELNSSPQPYLVPFLKRSLPALRQLTPDSAAFIQQSQQQPPPPTSQATTALTAVVLSSSVQRTAGKTAATVTSALQPPVLSLTQPTQVGVGKQGQPTPLVIQQPPKPGALIRPPQVTLTQTPMVALRQPHNRIMLTTPQQIQLNPLQPVPVVKPAVLPGTKALSAVSAQAAAAQKNKLKEPGGGSFRDDDDINDVASMAGVNLSEESARILATNSELVGTLTRSCKDETFLLQAPLQRRILEIGKKHGITELHPDVVSYVSHATQQRLQNLVEKISETAQQKNFSYKDDDRYEQASDVRAQLKFFEQLDQIEKQRKDEQEREILMRAAKSRSRQEDPEQLRLKQKAKEMQQQELAQMRQRDANLTALAAIGPRKKRKVDCPGPGSGAEGSGPGSVVPGSSGVGTPRQFTRQRITRVNLRDLIFCLENERETSHSLLLYKAFLK | The TFIID basal transcription factor complex plays a major role in the initiation of RNA polymerase II (Pol II)-dependent transcription. TFIID recognizes and binds promoters with or without a TATA box via its subunit TBP, a TATA-box-binding protein, and promotes assembly of the pre-initiation complex (PIC). The TFIID complex consists of TBP and TBP-associated factors (TAFs), including TAF1, TAF2, TAF3, TAF4, TAF5, TAF6, TAF7, TAF8, TAF9, TAF10, TAF11, TAF12 and TAF13. TAF4 may maintain an association between the TFIID and TFIIA complexes, while bound to the promoter, together with TBP, during PIC assembly. Potentiates transcriptional activation by the AF-2S of the retinoic acid, vitamin D3 and thyroid hormone. |
Generate the functional description in free-text form based on the protein sequence. | MDRMASSMKQVPNPLPKVLSRRGVGAGLEAAERESFERTQTVSINKAINTQEVAVKEKHARTCILGTHHEKGAQTFWSVVNRLPLSSNAVLCWKFCHVFHKLLRDGHPNVLKDSLRYRNELSDMSRMWGHLSEGYGQLCSIYLKLLRTKMEYHTKNPRFPGNLQMSDRQLDEAGESDVNNFFQLTVEMFDYLECELNLFQTVFNSLDMSRSVSVTAAGQCRLAPLIQVILDCSHLYDYTVKLLFKLHSCLPADTLQGHRDRFMEQFTKLKDLFYRSSNLQYFKRLIQIPQLPENPPNFLRASALSEHISPVVVIPAEASSPDSEPVLEKDDLMDMDASQQNLFDNKFDDIFGSSFSSDPFNFNSQNGVNKDEKDHLIERLYREISGLKAQLENMKTESQRVVLQLKGHVSELEADLAEQQHLRQQAADDCEFLRAELDELRRQREDTEKAQRSLSEIERKAQANEQRYSKLKEKYSELVQNHADLLRKNAEVTKQVSMARQAQVDLEREKKELEDSLERISDQGQRKTQEQLEVLESLKQELATSQRELQVLQGSLETSAQSEANWAAEFAELEKERDSLVSGAAHREEELSALRKELQDTQLKLASTEESMCQLAKDQRKMLLVGSRKAAEQVIQDALNQLEEPPLISCAGSADHLLSTVTSISSCIEQLEKSWSQYLACPEDISGLLHSITLLAHLTSDAIAHGATTCLRAPPEPADSLTEACKQYGRETLAYLASLEEEGSLENADSTAMRNCLSKIKAIGEELLPRGLDIKQEELGDLVDKEMAATSAAIETATARIEEMLSKSRAGDTGVKLEVNERILGCCTSLMQAIQVLIVASKDLQREIVESGRGTASPKEFYAKNSRWTEGLISASKAVGWGATVMVDAADLVVQGRGKFEELMVCSHEIAASTAQLVAASKVKADKDSPNLAQLQQASRGVNQATAGVVASTISGKSQIEETDNMDFSSMTLTQIKRQEMDSQVRVLELENELQKERQKLGELRKKHYELAGVAEGWEEGTEASPPTLQEVVTEKE | Plays a role in clathrin-mediated endocytosis and trafficking. Involved in regulating AMPA receptor trafficking in the central nervous system in an NMDA-dependent manner (By similarity). Regulates presynaptic nerve terminal activity (By similarity). Enhances androgen receptor (AR)-mediated transcription. May act as a proapoptotic protein that induces cell death by acting through the intrinsic apoptosis pathway. Binds 3-phosphoinositides (via ENTH domain). May act through the ENTH domain to promote cell survival by stabilizing receptor tyrosine kinases following ligand-induced endocytosis. May play a functional role in the cell filament networks. May be required for differentiation, proliferation, and/or survival of somatic and germline progenitors. |
Predict the functional description of the protein sequence below. | MNNLLCCALVFLDISIKWTTQETFPPKYLHYDEETSHQLLCDKCPPGTYLKQHCTAKWKTVCAPCPDHYYTDSWHTSDECLYCSPVCKELQYVKQECNRTHNRVCECKEGRYLEIEFCLKHRSCPPGFGVVQAGTPERNTVCKRCPDGFFSNETSSKAPCRKHTNCSVFGLLLTQKGNATHDNICSGNSESTQKCGIDVTLCEEAFFRFAVPTKFTPNWLSVLVDNLPGTKVNAESVERIKRQHSSQEQTFQLLKLWKHQNKDQDIVKKIIQDIDLCENSVQRHIGHANLTFEQLRSLMESLPGKKVGAEDIEKTIKACKPSDQILKLLSLWRIKNGDQDTLKGLMHALKHSKTYHFPKTVTQSLKKTIRFLHSFTMYKLYQKLFLEMIGNQVQSVKISCL | Acts as decoy receptor for TNFSF11/RANKL and thereby neutralizes its function in osteoclastogenesis. Inhibits the activation of osteoclasts and promotes osteoclast apoptosis in vitro. Bone homeostasis seems to depend on the local ratio between TNFSF11 and TNFRSF11B. May also play a role in preventing arterial calcification. May act as decoy receptor for TNFSF10/TRAIL and protect against apoptosis. TNFSF10/TRAIL binding blocks the inhibition of osteoclastogenesis. |
Can you provide the functional description of the following protein sequence? | MASAAAAEAEKGSPVVVGLLVVGNIIILLSGLSLFAETIWVTADQYRVYPLMGVSGKDDVFAGAWIAIFCGFSFFMVASFGVGAALCRRRSMVLTYLVLMLIVYIFECASCITSYTHRDYMVSNPSLITKQMLTFYSADTDQGQELTRLWDRVMIEQECCGTSGPMDWVNFTSAFRAATPEVVFPWPPLCCRRTGNFIPLNEEGCRLGHMDYLFTKGCFEHIGHAIDSYTWGISWFGFAILMWTLPVMLIAMYFYTML | Component of the asymmetric unit membrane (AUM); a highly specialized biomembrane elaborated by terminally differentiated urothelial cells. May play an important role in normal bladder epithelial physiology, possibly in regulating membrane permeability of superficial umbrella cells or in stabilizing the apical membrane through AUM/cytoskeletal interactions (By similarity). |
Generate the functional description in free-text form based on the protein sequence. | MPPGVDCPMEFWTKEENQSVVVDFLLPTGVYLNFPVSRNANLSTIKQLLWHRAQYEPLFHMLSGPEAYVFTCINQTAEQQELEDEQRRLCDVQPFLPVLRLVAREGDRVKKLINSQISLLIGKGLHEFDSLCDPEVNDFRAKMCQFCEEAAARRQQLGWEAWLQYSFPLQLEPSAQTWGPGTLRLPNRALLVNVKFEGSEESFTFQVSTKDVPLALMACALRKKATVFRQPLVEQPEDYTLQVNGRHEYLYGSYPLCQFQYICSCLHSGLTPHLTMVHSSSILAMRDEQSNPAPQVQKPRAKPPPIPAKKPSSVSLWSLEQPFRIELIQGSKVNADERMKLVVQAGLFHGNEMLCKTVSSSEVSVCSEPVWKQRLEFDINICDLPRMARLCFALYAVIEKAKKARSTKKKSKKADCPIAWANLMLFDYKDQLKTGERCLYMWPSVPDEKGELLNPTGTVRSNPNTDSAAALLICLPEVAPHPVYYPALEKILELGRHSECVHVTEEEQLQLREILERRGSGELYEHEKDLVWKLRHEVQEHFPEALARLLLVTKWNKHEDVAQMLYLLCSWPELPVLSALELLDFSFPDCHVGSFAIKSLRKLTDDELFQYLLQLVQVLKYESYLDCELTKFLLDRALANRKIGHFLFWHLRSEMHVPSVALRFGLILEAYCRGSTHHMKVLMKQGEALSKLKALNDFVKLSSQKTPKPQTKELMHLCMRQEAYLEALSHLQSPLDPSTLLAEVCVEQCTFMDSKMKPLWIMYSNEEAGSGGSVGIIFKNGDDLRQDMLTLQMIQLMDVLWKQEGLDLRMTPYGCLPTGDRTGLIEVVLRSDTIANIQLNKSNMAATAAFNKDALLNWLKSKNPGEALDRAIEEFTLSCAGYCVATYVLGIGDRHSDNIMIRESGQLFHIDFGHFLGNFKTKFGINRERVPFILTYDFVHVIQQGKTNNSEKFERFRGYCERAYTILRRHGLLFLHLFALMRAAGLPELSCSKDIQYLKDSLALGKTEEEALKHFRVKFNEALRESWKTKVNWLAHNVSKDNRQ | Phosphoinositide-3-kinase (PI3K) phosphorylates phosphatidylinositol (PI) and its phosphorylated derivatives at position 3 of the inositol ring to produce 3-phosphoinositides. Uses ATP and PtdIns(4,5)P2 (phosphatidylinositol 4,5-bisphosphate) to generate phosphatidylinositol 3,4,5-trisphosphate (PIP3). PIP3 plays a key role by recruiting PH domain-containing proteins to the membrane, including AKT1 and PDPK1, activating signaling cascades involved in cell growth, survival, proliferation, motility and morphology. Mediates immune responses. Plays a role in B-cell development, proliferation, migration, and function. Required for B-cell receptor (BCR) signaling. Mediates B-cell proliferation response to anti-IgM, anti-CD40 and IL4 stimulation. Promotes cytokine production in response to TLR4 and TLR9. Required for antibody class switch mediated by TLR9. Involved in the antigen presentation function of B-cells. Involved in B-cell chemotaxis in response to CXCL13 and sphingosine 1-phosphate (S1P). Required for proliferation, signaling and cytokine production of naive, effector and memory T-cells. Required for T-cell receptor (TCR) signaling. Mediates TCR signaling events at the immune synapse. Activation by TCR leads to antigen-dependent memory T-cell migration and retention to antigenic tissues. Together with PIK3CG participates in T-cell development. Contributes to T-helper cell expansion and differentiation. Required for T-cell migration mediated by homing receptors SELL/CD62L, CCR7 and S1PR1 and antigen dependent recruitment of T-cells. Together with PIK3CG is involved in natural killer (NK) cell development and migration towards the sites of inflammation. Participates in NK cell receptor activation. Plays a role in NK cell maturation and cytokine production. Together with PIK3CG is involved in neutrophil chemotaxis and extravasation. Together with PIK3CG participates in neutrophil respiratory burst. Plays important roles in mast-cell development and mast cell mediated allergic response. Involved in stem cell factor (SCF)-mediated proliferation, adhesion and migration. Required for allergen-IgE-induced degranulation and cytokine release. The lipid kinase activity is required for its biological function. Isoform 2 may be involved in stabilizing total RAS levels, resulting in increased ERK phosphorylation and increased PI3K activity. |
Generate the functional description in free-text form based on the protein sequence. | MAASWRLGCDPRLLRYLVGFPGRRSVGLVKGALGWSVSRGANWRWFHSTQWLRGDPIKILMPSLSPTMEEGNIVKWLKKEGEAVSAGDALCEIETDKAVVTLDASDDGILAKIVVEEGSKNIRLGSLIGLIVEEGEDWKHVEIPKDVGPPPPVSKPSEPRPSPEPQISIPVKKEHIPGTLRFRLSPAARNILEKHSLDASQGTATGPRGIFTKEDALKLVQLKQTGKITESRPTPAPTATPTAPSPLQATAGPSYPRPVIPPVSTPGQPNAVGTFTEIPASNIRRVIAKRLTESKSTVPHAYATADCDLGAVLKVRQDLVKDDIKVSVNDFIIKAAAVTLKQMPDVNVSWDGEGPKQLPFIDISVAVATDKGLLTPIIKDAAAKGIQEIADSVKALSKKARDGKLLPEEYQGGSFSISNLGMFGIDEFTAVINPPQACILAVGRFRPVLKLTEDEEGNAKLQQRQLITVTMSSDSRVVDDELATRFLKSFKANLENPIRLA | Required for anchoring dihydrolipoamide dehydrogenase (E3) to the dihydrolipoamide transacetylase (E2) core of the pyruvate dehydrogenase complexes of eukaryotes. This specific binding is essential for a functional PDH complex. |
Generate the functional description in free-text form based on the protein sequence. | MENDPSRRRESISLTPVAKGLENMGADFLESLEEGQLPRSDLSPAEIRSSWSEAAPKPFSRWRNLQPALRARSFCREHMQLFRWIGTGLLCTGLSAFLLVACLLDFQRALALFVLTCVVLTFLGHRLLKRLLGPKLRRFLKPQGHPRLLLWFKRGLALAAFLGLVLWLSLDTSQRPEQLVSFAGICVFVALLFACSKHHCAVSWRAVSWGLGLQFVLGLLVIRTEPGFIAFEWLGEQIRIFLSYTKAGSSFVFGEALVKDVFAFQVLPIIVFFSCVISVLYHVGLMQWVILKIAWLMQVTMGTTATETLSVAGNIFVSQTEAPLLIRPYLADMTLSEVHVVMTGGYATIAGSLLGAYISFGIDATSLIAASVMAAPCALALSKLVYPEVEESKFRREEGVKLTYGDAQNLIEAASTGAAISVKVVANIAANLIAFLAVLDFINAALSWLGDMVDIQGLSFQLICSYILRPVAFLMGVAWEDCPVVAELLGIKLFLNEFVAYQDLSKYKQRRLAGAEEWVGDRKQWISVRAEVLTTFALCGFANFSSIGIMLGGLTSMVPQRKSDFSQIVLRALFTGACVSLVNACMAGILYMPRGAEVDCMSLLNTTLSSSSFEIYQCCREAFQSVNPEFSPEALDNCCRFYNHTICAQ | Sodium-dependent and pyrimidine-selective transporter. Exhibits the transport characteristics of the nucleoside transport system cit or N2 subtype (N2/cit) (selective for pyrimidine nucleosides and adenosine). Transports uridine, cytidine, thymidine, and nucleoside-derived drugs. Transports the antiviral pyrimidine nucleoside analogs 3'-azido-3'-deoxythymidine (AZT) and 2',3'-dideoxycytidine (ddC). It may be involved in the intestinal absorption and renal handling of pyrimidine nucleoside analogs used to treat acquired immunodeficiency syndrome (AIDS). It has the following selective inhibition: adenosine, thymidine, cytidine, uridine >> guanosine, inosine. |
Can you provide the functional description of the following protein sequence? | MSPTISHKDSSRQRRPGNFSHSLDMKSGPLPPGGWDDSHLDSAGREGDREALLGDTGTGDFLKAPQSFRAELSSILLLLFLYVLQGIPLGLAGSIPLILQSKNVSYTDQAFFSFVFWPFSLKLLWAPLVDAVYVKNFGRRKSWLVPTQYILGLFMIYLSTQVDRLLGNTDDRTPDVIALTVAFFLFEFLAATQDIAVDGWALTMLSRENVGYASTCNSVGQTAGYFLGNVLFLALESADFCNKYLRFQPQPRGIVTLSDFLFFWGTVFLITTTLVALLKKENEVSVVKEETQGITDTYKLLFAIIKMPAVLTFCLLILTAKIGFSAADAVTGLKLVEEGVPKEHLALLAVPMVPLQIILPLIISKYTAGPQPLNTFYKAMPYRLLLGLEYALLVWWTPKVEHQGGFPIYYYIVVLLSYALHQVTVYSMYVSIMAFNAKVSDPLIGGTYMTLLNTVSNLGGNWPSTVALWLVDPLTVKECVGASNQNCRTPDAVELCKKLGGSCVTALDGYYVESIICVFIGFGWWFFLGPKFKKLQDEGSSSWKCKRNN | Probable acetyl-CoA transporter necessary for O-acetylation of gangliosides. Negatively regulates BMP signaling. |
Generate the functional description in free-text form based on the protein sequence. | MGPVMPPSKKPESSGISVSSGLSQCYGGSGFSKALQEDDDLDFSLPDIRLEEGAMEDEELTNLNWLHESKNLLKSFGESVLRSVSPVQDLDDDTPPSPAHSDMPYDARQNPNCKPPYSFSCLIFMAIEDSPTKRLPVKDIYNWILEHFPYFANAPTGWKNSVRHNLSLNKCFKKVDKERSQSIGKGSLWCIDPEYRQNLIQALKKTPYHPHPHVFNTPPTCPQAYQSTSGPPIWPGSTFFKRNGALLQDPDIDAASAMMLLNTPPEIQAGFPPGVIQNGARVLSRGLFPGVRPLPITPIGVTAAMRNGITSCRMRTESEPSCGSPVVSGDPKEDHNYSSAKSSNARSTSPTSDSISSSSSSADDHYEFATKGSQEGSEGSEGSFRSHESPSDTEEDDRKHSQKEPKDSLGDSGYASQHKKRQHFAKARKVPSDTLPLKKRRTEKPPESDDEEMKEAAGSLLHLAGIRSCLNNITNRTAKGQKEQKETTKN | Acts as a transcriptional repressor. May be involved in DNA damage-inducible cell cycle arrests (checkpoints). |
Predict the functional description of the protein sequence below. | MAVESRVTQEEIKKEPEKPIDREKTCPLLLRVFTTNNGRHHRMDEFSRGNVPSSELQIYTWMDATLKELTSLVKEVYPEARKKGTHFNFAIVFTDVKRPGYRVKEIGSTMSGRKGTDDSMTLQSQKFQIGDYLDIAITPPNRAPPPSGRMRPY | Component of the SIN3-repressing complex. Enhances the ability of SIN3-HDAC1-mediated transcriptional repression. When tethered to the promoter, it can direct the formation of a repressive complex to core histone proteins. Auxiliary component of the splicing-dependent multiprotein exon junction complex (EJC) deposited at splice junction on mRNAs. The EJC is a dynamic structure consisting of core proteins and several peripheral nuclear and cytoplasmic associated factors that join the complex only transiently either during EJC assembly or during subsequent mRNA metabolism. Component of the ASAP and PSAP complexes which bind RNA in a sequence-independent manner and are proposed to be recruited to the EJC prior to or during the splicing process and to regulate specific excision of introns in specific transcription subsets. The ASAP complex can inhibit mRNA processing during in vitro splicing reactions. The ASAP complex promotes apoptosis and is disassembled after induction of apoptosis. Involved in the splicing modulation of BCL2L1/Bcl-X (and probably other apoptotic genes); specifically inhibits the formation of proapoptotic isoforms such as Bcl-X(S); the activity is different from the established EJC assembly and function. |
Generate the functional description in free-text form based on the protein sequence. | MATCIGEKIEDFKVGNLLGKGSFAGVYRAESIHTGLEVAIKMIDKKAMYKAGMVQRVQNEVKIHCQLKHPSILELYNYFEDSNYVYLVLEMCHNGEMNRYLKNRVKPFSENEARHFMHQIITGMLYLHSHGILHRDLTLSNLLLTRNMNIKIADFGLATQLKMPHEKHYTLCGTPNYISPEIATRSAHGLESDVWSLGCMFYTLLIGRPPFDTDTVKNTLNKVVLADYEMPSFLSIEAKDLIHQLLRRNPADRLSLSSVLDHPFMSRNSSTKSKDLGTVEDSIDSGHATISTAITASSSTSISGSLFDKRRLLIGQPLPNKMTVFPKNKSSTDFSSSGDGNSFYTQWGNQETSNSGRGRVIQDAEERPHSRYLRRAYSSDRSGTSNSQSQAKTYTMERCHSAEMLSVSKRSGGGENEERYSPTDNNANIFNFFKEKTSSSSGSFERPDNNQALSNHLCPGKTPFPFADPTPQTETVQQWFGNLQINAHLRKTTEYDSISPNRDFQGHPDLQKDTSKNAWTDTKVKKNSDASDNAHSVKQQNTMKYMTALHSKPEIIQQECVFGSDPLSEQSKTRGMEPPWGYQNRTLRSITSPLVAHRLKPIRQKTKKAVVSILDSEEVCVELVKEYASQEYVKEVLQISSDGNTITIYYPNGGRGFPLADRPPSPTDNISRYSFDNLPEKYWRKYQYASRFVQLVRSKSPKITYFTRYAKCILMENSPGADFEVWFYDGVKIHKTEDFIQVIEKTGKSYTLKSESEVNSLKEEIKMYMDHANEGHRICLALESIISEEERKTRSAPFFPIIIGRKPGSTSSPKALSPPPSVDSNYPTRERASFNRMVMHSAASPTQAPILNPSMVTNEGLGLTTTASGTDISSNSLKDCLPKSAQLLKSVFVKNVGWATQLTSGAVWVQFNDGSQLVVQAGVSSISYTSPNGQTTRYGENEKLPDYIKQKLQCLSSILLMFSNPTPNFH | Serine/threonine-protein kinase that plays a central role in centriole duplication. Able to trigger procentriole formation on the surface of the parental centriole cylinder, leading to the recruitment of centriole biogenesis proteins such as SASS6, CENPJ/CPAP, CCP110, CEP135 and gamma-tubulin. When overexpressed, it is able to induce centrosome amplification through the simultaneous generation of multiple procentrioles adjoining each parental centriole during S phase. Phosphorylates 'Ser-151' of FBXW5 during the G1/S transition, leading to inhibit FBXW5 ability to ubiquitinate SASS6. Its central role in centriole replication suggests a possible role in tumorigenesis, centrosome aberrations being frequently observed in tumors. Also involved in deuterosome-mediated centriole amplification in multiciliated that can generate more than 100 centrioles. Also involved in trophoblast differentiation by phosphorylating HAND1, leading to disrupt the interaction between HAND1 and MDFIC and activate HAND1. Phosphorylates CDC25C and CHEK2. Required for the recruitment of STIL to the centriole and for STIL-mediated centriole amplification. Phosphorylates CEP131 at 'Ser-78' and PCM1 at 'Ser-372' which is essential for proper organization and integrity of centriolar satellites. |
Predict the functional description of the protein sequence below. | MLSRNDDICIYGGLGLGGLLLLAVVLLSACLCWLHRRVKRLERSWAQGSSEQELHYASLQRLPVPSSEGPDLRGRDKRGTKEDPRADYACIAENKPT | Possible role in modulating immune responses. Induces morphological changes including production of filopodia and microspikes when overexpressed in a variety of cell types and may be involved in dendritic cell maturation. Isoform 1 and isoform 2 have an inhibitory effect on lymphocyte proliferation. |
Can you provide the functional description of the following protein sequence? | MRLHLLLLLALCGAGTTAAELSYSLRGNWSICNGNGSLELPGAVPGCVHSALFQQGLIQDSYYRFNDLNYRWVSLDNWTYSKEFKIPFEISKWQKVNLILEGVDTVSKILFNEVTIGETDNMFNRYSFDITNVVRDVNSIELRFQSAVLYAAQQSKAHTRYQVPPDCPPLVQKGECHVNFVRKEQCSFSWDWGPSFPTQGIWKDVRIEAYNICHLNYFTFSPIYDKSAQEWNLEIESTFDVVSSKPVGGQVIVAIPKLQTQQTYSIELQPGKRIVELFVNISKNITVETWWPHGHGNQTGYNMTVLFELDGGLNIEKSAKVYFRTVELIEEPIKGSPGLSFYFKINGFPIFLKGSNWIPADSFQDRVTSELLRLLLQSVVDANMNTLRVWGGGIYEQDEFYELCDELGIMVWQDFMFACALYPTDQGFLDSVTAEVAYQIKRLKSHPSIIIWSGNNENEEALMMNWYHISFTDRPIYIKDYVTLYVKNIRELVLAGDKSRPFITSSPTNGAETVAEAWVSQNPNSNYFGDVHFYDYISDCWNWKVFPKARFASEYGYQSWPSFSTLEKVSSTEDWSFNSKFSLHRQHHEGGNKQMLYQAGLHFKLPQSTDPLRTFKDTIYLTQVMQAQCVKTETEFYRRSRSEIVDQQGHTMGALYWQLNDIWQAPSWASLEYGGKWKMLHYFAQNFFAPLLPVGFENENTFYIYGVSDLHSDYSMTLSVRVHTWSSLEPVCSRVTERFVMKGGEAVCLYEEPVSELLRRCGNCTRESCVVSFYLSADHELLSPTNYHFLSSPKEAVGLCKAQITAIISQQGDIFVFDLETSAVAPFVWLDVGSIPGRFSDNGFLMTEKTRTILFYPWEPTSKNELEQSFHVTSLTDIY | Exoglycosidase that cleaves the single beta-linked mannose residue from the non-reducing end of all N-linked glycoprotein oligosaccharides. |
What is the functional description of the protein sequence? | MKMASSLAFLLLNFHVSLFLVQLLTPCSAQFSVLGPSGPILAMVGEDADLPCHLFPTMSAETMELRWVSSSLRQVVNVYADGKEVEDRQSAPYRGRTSILRDGITAGKAALRIHNVTASDSGKYLCYFQDGDFYEKALVELKVAALGSDLHIEVKGYEDGGIHLECRSTGWYPQPQIKWSDTKGENIPAVEAPVVADGVGLYAVAASVIMRGSSGGGVSCIIRNSLLGLEKTASISIADPFFRSAQPWIAALAGTLPISLLLLAGASYFLWRQQKEKIALSRETEREREMKEMGYAATEQEISLREKLQEELKWRKIQYMARGEKSLAYHEWKMALFKPADVILDPDTANAILLVSEDQRSVQRAEEPRDLPDNPERFEWRYCVLGCENFTSGRHYWEVEVGDRKEWHIGVCSKNVERKKGWVKMTPENGYWTMGLTDGNKYRALTEPRTNLKLPEPPRKVGIFLDYETGEISFYNATDGSHIYTFPHASFSEPLYPVFRILTLEPTALTICPIPKEVESSPDPDLVPDHSLETPLTPGLANESGEPQAEVTSLLLPAHPGAEVSPSATTNQNHKLQARTEALY | Plays a role in T-cell responses in the adaptive immune response. |
Predict the functional description of the protein sequence below. | MDRLLRLGGGMPGLGQGPPTDAPAVDTAEQVYISSLALLKMLKHGRAGVPMEVMGLMLGEFVDDYTVRVIDVFAMPQSGTGVSVEAVDPVFQAKMLDMLKQTGRPEMVVGWYHSHPGFGCWLSGVDINTQQSFEALSERAVAVVVDPIQSVKGKVVIDAFRLINANMMVLGHEPRQTTSNLGHLNKPSIQALIHGLNRHYYSITINYRKNELEQKMLLNLHKKSWMEGLTLQDYSEHCKHNESVVKEMLELAKNYNKAVEEEDKMTPEQLAIKNVGKQDPKRHLEEHVDVLMTSNIVQCLAAMLDTVVFK | Component of the 26S proteasome, a multiprotein complex involved in the ATP-dependent degradation of ubiquitinated proteins. This complex plays a key role in the maintenance of protein homeostasis by removing misfolded or damaged proteins, which could impair cellular functions, and by removing proteins whose functions are no longer required. Therefore, the proteasome participates in numerous cellular processes, including cell cycle progression, apoptosis, or DNA damage repair. The PSMD14 subunit is a metalloprotease that specifically cleaves 'Lys-63'-linked polyubiquitin chains within the complex. Plays a role in response to double-strand breaks (DSBs): acts as a regulator of non-homologous end joining (NHEJ) by cleaving 'Lys-63'-linked polyubiquitin, thereby promoting retention of JMJD2A/KDM4A on chromatin and restricting TP53BP1 accumulation. Also involved in homologous recombination repair by promoting RAD51 loading. |
Predict the functional description of the protein sequence below. | MAENPSLENHRIKSFKNKGRDVETMRRHRNEVTVELRKNKRDEHLLKKRNVPQEESLEDSDVDADFKAQNVTLEAILQNATSDNPVVQLSAVQAARKLLSSDRNPPIDDLIKSGILPILVKCLERDDNPSLQFEAAWALTNIASGTSAQTQAVVQSNAVPLFLRLLRSPHQNVCEQAVWALGNIIGDGPQCRDYVISLGVVKPLLSFISPSIPITFLRNVTWVIVNLCRNKDPPPPMETVQEILPALCVLIYHTDINILVDTVWALSYLTDGGNEQIQMVIDSGVVPFLVPLLSHQEVKVQTAALRAVGNIVTGTDEQTQVVLNCDVLSHFPNLLSHPKEKINKEAVWFLSNITAGNQQQVQAVIDAGLIPMIIHQLAKGDFGTQKEAAWAISNLTISGRKDQVEYLVQQNVIPPFCNLLSVKDSQVVQVVLDGLKNILIMAGDEASTIAEIIEECGGLEKIEVLQQHENEDIYKLAFEIIDQYFSGDDIDEDPCLIPEATQGGTYNFDPTANLQTKEFNF | Functions in nuclear protein import as an adapter protein for nuclear receptor KPNB1. Binds specifically and directly to substrates containing either a simple or bipartite NLS motif. Docking of the importin/substrate complex to the nuclear pore complex (NPC) is mediated by KPNB1 through binding to nucleoporin FxFG repeats and the complex is subsequently translocated through the pore by an energy requiring, Ran-dependent mechanism. At the nucleoplasmic side of the NPC, Ran binds to importin-beta and the three components separate and importin-alpha and -beta are re-exported from the nucleus to the cytoplasm where GTP hydrolysis releases Ran from importin. The directionality of nuclear import is thought to be conferred by an asymmetric distribution of the GTP- and GDP-bound forms of Ran between the cytoplasm and nucleus. In vitro, mediates the nuclear import of human cytomegalovirus UL84 by recognizing a non-classical NLS. Recognizes NLSs of influenza A virus nucleoprotein probably through ARM repeats 7-9. |
Predict the functional description of the protein sequence below. | MAHLRGFANQHSRVDPEELFTKLDRIGKGSFGEVYKGIDNHTKEVVAIKIIDLEEAEDEIEDIQQEITVLSQCDSPYITRYFGSYLKSTKLWIIMEYLGGGSALDLLKPGPLEETYIATILREILKGLDYLHSERKIHRDIKAANVLLSEQGDVKLADFGVAGQLTDTQIKRNTFVGTPFWMAPEVIKQSAYDFKADIWSLGITAIELAKGEPPNSDLHPMRVLFLIPKNSPPTLEGQHSKPFKEFVEACLNKDPRFRPTAKELLKHKFITRYTKKTSFLTELIDRYKRWKSEGHGEESSSEDSDIDGEAEDGEQGPIWTFPPTIRPSPHSKLHKGTALHSSQKPAEPVKRQPRSQCLSTLVRPVFGELKEKHKQSGGSVGALEELENAFSLAEESCPGISDKLMVHLVERVQRFSHNRNHLTSTR | Oxidant stress-activated serine/threonine kinase that may play a role in the response to environmental stress. Targets to the Golgi apparatus where it appears to regulate protein transport events, cell adhesion, and polarity complexes important for cell migration. |
Generate the functional description in free-text form based on the protein sequence. | MTAITHGSPVGGNDSQGQVLDGQSQHLFQQNQTSSPDSSNENSVATPPPEEQGQGDAPPQHEDEEPAFPHTELANLDDMINRPRWVVPVLPKGELEVLLEAAIDLSVKGLDVKSEACQRFFRDGLTISFTKILMDEAVSGWKFEIHRCIINNTHRLVELCVAKLSQDWFPLLELLAMALNPHCKFHIYNGTRPCELISSNAQLPEDELFARSSDPRSPKGWLVDLINKFGTLNGFQILHDRFFNGSALNIQIIAALIKPFGQCYEFLSQHTLKKYFIPVIEIVPHLLENLTDEELKKEAKNEAKNDALSMIIKSLKNLASRISGQDETIKNLEIFRLKMILRLLQISSFNGKMNALNEINKVISSVSYYTHRHSNPEEEEWLTAERMAEWIQQNNILSIVLQDSLHQPQYVEKLEKILRFVIKEKALTLQDLDNIWAAQAGKHEAIVKNVHDLLAKLAWDFSPGQLDHLFDCFKASWTNASKKQREKLLELIRRLAEDDKDGVMAHKVLNLLWNLAQSDDVPVDIMDLALSAHIKILDYSCSQDRDAQKIQWIDHFIEELRTNDKWVIPALKQIREICSLFGEASQNLSQTQRSPHIFYRHDLINQLQQNHALVTLVAENLATYMNSIRLYAGDHEDYDPQTVRLGSRYSHVQEVQERLNFLRFLLKDGQLWLCAPQAKQIWKCLAENAVYLCDREACFKWYSKLMGDEPDLDPDINKDFFESNVLQLDPSLLTENGMKCFERFFKAVNCRERKLIAKRRSYMMDDLELIGLDYLWRVVIQSSDEIANRAIDLLKEIYTNLGPRLKANQVVIHEDFIQSCFDRLKASYDTLCVFDGDKNSINCARQEAIRMVRVLTVIKEYINECDSDYHKERMILPMSRAFRGKHLSLIVRFPNQGRQVDELDIWSHTNDTIGSVRRCIVNRIKANVAHKKIELFVGGELIDSEDDRKLIGQLNLKDKSLITAKLTQINFNMPSSPDSSSDSSTASPGNHRNHYNDGPNLEVESCLPGVIMSVHPRYISFLWQVADLGSNLNMPPLRDGARVLMKLMPPDRTAVEKLRAVCLDHAKLGEGKLSPPLDSLFFGPSASQVLYLTEVVYALLMPAGVPLTDGSSDFQVHFLKSGGLPLVLSMLIRNNFLPNTDMETRRGAYLNALKIAKLLLTAIGYGHVRAVAEACQPVVDGTDPITQINQVTHDQAVVLQSALQSIPNPSSECVLRNESILLAQEISNEASRYMPDICVIRAIQKIIWASACGALGLVFSPNEEITKIYQMTTNGSNKLEVEDEQVCCEALEVMTLCFALLPTALDALSKEKAWQTFIIDLLLHCPSKTVRQLAQEQFFLMCTRCCMGHRPLLFFITLLFTILGSTAREKGKYSGDYFTLLRHLLNYAYNGNINIPNAEVLLVSEIDWLKRIRDNVKNTGETGVEEPILEGHLGVTKELLAFQTSEKKYHFGCEKGGANLIKELIDDFIFPASKVYLQYLRSGELPAEQAIPVCSSPVTINAGFELLVALAIGCVRNLKQIVDCLTEMYYMGTAITTCEALTEWEYLPPVGPRPPKGFVGLKNAGATCYMNSVIQQLYMIPSIRNSILAIEGTGSDLHDDMFGDEKQDSESNVDPRDDVFGYPHQFEDKPALSKTEDRKEYNIGVLRHLQVIFGHLAASQLQYYVPRGFWKQFRLWGEPVNLREQHDALEFFNSLVDSLDEALKALGHPAILSKVLGGSFADQKICQGCPHRYECEESFTTLNVDIRNHQNLLDSLEQYIKGDLLEGANAYHCEKCDKKVDTVKRLLIKKLPRVLAIQLKRFDYDWERECAIKFNDYFEFPRELDMGPYTVAGVANLERDNVNSENELIEQKEQSDNETAGGTKYRLVGVLVHSGQASGGHYYSYIIQRNGKDDQTDHWYKFDDGDVTECKMDDDEEMKNQCFGGEYMGEVFDHMMKRMSYRRQKRWWNAYILFYEQMDMIDEDDEMIRYISELTIARPHQIIMSPAIERSVRKQNVKFMHNRLQYSLEYFQFVKKLLTCNGVYLNPAPGQDYLLPEAEEITMISIQLAARFLFTTGFHTKKIVRGPASDWYDALCVLLRHSKNVRFWFTHNVLFNVSNRFSEYLLECPSAEVRGAFAKLIVFIAHFSLQDGSCPSPFASPGPSSQACDNLSLSDHLLRATLNLLRREVSEHGHHLQQYFNLFVMYANLGVAEKTQLLKLNVPATFMLVSLDEGPGPPIKYQYAELGKLYSVVSQLIRCCNVSSTMQSSINGNPPLPNPFGDLNLSQPIMPIQQNVLDILFVRTSYVKKIIEDCSNSEDTIKLLRFCSWENPQFSSTVLSELLWQVAYSYTYELRPYLDLLFQILLIEDSWQTHRIHNALKGIPDDRDGLFDTIQRSKNHYQKRAYQCIKCMVALFSSCPVAYQILQGNGDLKRKWTWAVEWLGDELERRPYTGNPQYSYNNWSPPVQSNETANGYFLERSHSARMTLAKACELCPEEEPDDQDAPDEHEPSPSEDAPLYPHSPASQYQQNNHVHGQPYTGPAAHHLNNPQKTGQRTQENYEGNEEVSSPQMKDQ | May function as a ubiquitin-protein or polyubiquitin hydrolase involved both in the processing of ubiquitin precursors and of ubiquitinated proteins. May therefore play an important regulatory role at the level of protein turnover by preventing degradation of proteins through the removal of conjugated ubiquitin. Essential component of TGF-beta/BMP signaling cascade. Deubiquitinates monoubiquitinated SMAD4, opposing the activity of E3 ubiquitin-protein ligase TRIM33. Monoubiquitination of SMAD4 hampers its ability to form a stable complex with activated SMAD2/3 resulting in inhibition of TGF-beta/BMP signaling cascade. Deubiquitination of SMAD4 by USP9X re-empowers its competence to mediate TGF-beta signaling (By similarity). |
Generate the functional description in free-text form based on the protein sequence. | MSLYPSLEDLKVDKVIQAQTAFSANPANPAILSEASAPIPHDGNLYPRLYPELSQYMGLSLNEEEIRANVAVVSGAPLQGQLVARPSSINYMVAPVTGNDVGIRRAEIKQGIREVILCKDQDGKIGLRLKSIDNGIFVQLVQANSPASLVGLRFGDQVLQINGENCAGWSSDKAHKVLKQAFGEKITMTIRDRPFERTITMHKDSTGHVGFIFKNGKITSIVKDSSAARNGLLTEHNICEINGQNVIGLKDSQIADILSTSGTVVTITIMPAFIFEHIIKRMAPSIMKSLMDHTIPEV | Multifunctional adapter protein involved in diverse array of functions including trafficking of transmembrane proteins, neuro and immunomodulation, exosome biogenesis, and tumorigenesis. Positively regulates TGFB1-mediated SMAD2/3 activation and TGFB1-induced epithelial-to-mesenchymal transition (EMT) and cell migration in various cell types. May increase TGFB1 signaling by enhancing cell-surface expression of TGFR1 by preventing the interaction between TGFR1 and CAV1 and subsequent CAV1-dependent internalization and degradation of TGFR1. In concert with SDC1/4 and PDCD6IP, regulates exosome biogenesis. Regulates migration, growth, proliferation, and cell cycle progression in a variety of cancer types. In adherens junctions may function to couple syndecans to cytoskeletal proteins or signaling components. Seems to couple transcription factor SOX4 to the IL-5 receptor (IL5RA). May also play a role in vesicular trafficking. Seems to be required for the targeting of TGFA to the cell surface in the early secretory pathway. |
Generate the functional description in free-text form based on the protein sequence. | MNYSLHLAFVCLSLFTERMCIQGSQFNVEVGRSDKLSLPGFENLTAGYNKFLRPNFGGEPVQIALTLDIASISSISESNMDYTATIYLRQRWMDQRLVFEGNKSFTLDARLVEFLWVPDTYIVESKKSFLHEVTVGNRLIRLFSNGTVLYALRITTTVACNMDLSKYPMDTQTCKLQLESWGYDGNDVEFTWLRGNDSVRGLEHLRLAQYTIERYFTLVTRSQQETGNYTRLVLQFELRRNVLYFILETYVPSTFLVVLSWVSFWISLDSVPARTCIGVTTVLSMTTLMIGSRTSLPNTNCFIKAIDVYLGICFSFVFGALLEYAVAHYSSLQQMAAKDRGTTKEVEEVSITNIINSSISSFKRKISFASIEISSDNVDYSDLTMKTSDKFKFVFREKMGRIVDYFTIQNPSNVDHYSKLLFPLIFMLANVFYWAYYMYF | GABA, the major inhibitory neurotransmitter in the vertebrate brain, mediates neuronal inhibition by binding to the GABA/benzodiazepine receptor and opening an integral chloride channel. In the uterus, the function of the receptor appears to be related to tissue contractility. The binding of this pI subunit with other GABA(A) receptor subunits alters the sensitivity of recombinant receptors to modulatory agents such as pregnanolone. |
What is the functional description of the protein sequence? | MSSRIARALALVVTLLHLTRLALSTCPAACHCPLEAPKCAPGVGLVRDGCGCCKVCAKQLNEDCSKTQPCDHTKGLECNFGASSTALKGICRAQSEGRPCEYNSRIYQNGESFQPNCKHQCTCIDGAVGCIPLCPQELSLPNLGCPNPRLVKVTGQCCEEWVCDEDSIKDPMEDQDGLLGKELGFDASEVELTRNNELIAVGKGSSLKRLPVFGMEPRILYNPLQGQKCIVQTTSWSQCSKTCGTGISTRVTNDNPECRLVKETRICEVRPCGQPVYSSLKKGKKCSKTKKSPEPVRFTYAGCLSVKKYRPKYCGSCVDGRCCTPQLTRTVKMRFRCEDGETFSKNVMMIQSCKCNYNCPHANEAAFPFYRLFNDIHKFRD | Promotes cell proliferation, chemotaxis, angiogenesis and cell adhesion. Appears to play a role in wound healing by up-regulating, in skin fibroblasts, the expression of a number of genes involved in angiogenesis, inflammation and matrix remodeling including VEGA-A, VEGA-C, MMP1, MMP3, TIMP1, uPA, PAI-1 and integrins alpha-3 and alpha-5. CCN1-mediated gene regulation is dependent on heparin-binding. Down-regulates the expression of alpha-1 and alpha-2 subunits of collagen type-1. Promotes cell adhesion and adhesive signaling through integrin alpha-6/beta-1, cell migration through integrin alpha-v/beta-5 and cell proliferation through integrin alpha-v/beta-3. |
Predict the functional description of the protein sequence below. | MASTTSTKKMMEEATCSICLSLMTNPVSINCGHSYCHLCITDFFKNPSQKQLRQETFCCPQCRAPFHMDSLRPNKQLGSLIEALKETDQEMSCEEHGEQFHLFCEDEGQLICWRCERAPQHKGHTTALVEDVCQGYKEKLQKAVTKLKQLEDRCTEQKLSTAMRITKWKEKVQIQRQKIRSDFKNLQCFLHEEEKSYLWRLEKEEQQTLSRLRDYEAGLGLKSNELKSHILELEEKCQGSAQKLLQNVNDTLSRSWAVKLETSEAVSLELHTMCNVSKLYFDVKKMLRSHQVSVTLDPDTAHHELILSEDRRQVTRGYTQENQDTSSRRFTAFPCVLGCEGFTSGRRYFEVDVGEGTGWDLGVCMENVQRGTGMKQEPQSGFWTLRLCKKKGYVALTSPPTSLHLHEQPLLVGIFLDYEAGVVSFYNGNTGCHIFTFPKASFSDTLRPYFQVYQYSPLFLPPPGD | E3 ubiquitin-protein and E3 SUMO-protein ligase that acts as a regulator of innate immunity. Acts as a negative regulator of type I interferon IFN-beta production by catalyzing 'Lys-48'-linked polyubiquitination of AZI2/NAP1, leading to its degradation (By similarity). Mediates 'Lys-48'-linked polyubiquitination and proteasomal degradation of the critical TLR adapter TICAM1, inhibiting TLR3-mediated type I interferon signaling. Acts as positive regulator of the cGAS-STING pathway by acting as a E3 SUMO-protein ligase: mediates sumoylation of CGAS and STING, preventing their degradation and thereby activating the innate immune response to DNA virus (By similarity). Also acts as a negative regulator of NF-kappa-B signaling independently of its E3 protein ligase activity by promoting lysosome-dependent degradation of TAB2 and TAB3 adapters. |
Predict the functional description of the protein sequence below. | MMYSPICLTQDEFHPFIEALLPHVRAIAYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLSEKPEIKQKWASRLLAKLRKDIRQEYREDFVLTVTGKKHPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLMKSPHCTNPALCVQPHHITVSVKELDLFLAYYVQEQDSGQSGSPSHNDPAKNPPGYLEDSFVKSGVFNVSELVRVSRTPITQGTGVNFPIGEIPSQPYYHDMNSGVNLQRSLSSPPSSKRPKTISIDENMEPSPTGDFYPSPSSPAAGSRTWHERDQDMSSPTTMKKPEKPLFSSASPQDSSPRLSTFPQHHHPGIPGVAHSVISTRTPPPPSPLPFPTQAILPPAPSSYFSHPTIRYPPHLNPQDTLKNYVPSYDPSSPQTSQSWYLG | Transcriptional activator of GFAP, essential for proper brain development. Recognizes and binds the palindromic sequence 5'-TTGGCNNNNNGCCAA-3' present in viral and cellular promoters and in the origin of replication of adenovirus type 2. These proteins are individually capable of activating transcription and replication. |
Generate the functional description in free-text form based on the protein sequence. | MGGLFWRSALRGLRCGPRAPGPSLLVRHGSGGPSWTRERTLVAVKPDGVQRRLVGDVIQRFERRGFTLVGMKMLQAPESVLAEHYQDLRRKPFYPALIRYMSSGPVVAMVWEGYNVVRASRAMIGHTDSAEAAPGTIRGDFSVHISRNVIHASDSVEGAQREIQLWFQSSELVSWADGGQHSSIHPA | Major role in the synthesis of nucleoside triphosphates other than ATP. The ATP gamma phosphate is transferred to the NDP beta phosphate via a ping-pong mechanism, using a phosphorylated active-site intermediate. Through the catalyzed exchange of gamma-phosphate between di- and triphosphonucleosides participates in regulation of intracellular nucleotide homeostasis. Binds to anionic phospholipids, predominantly to cardiolipin; the binding inhibits its phosphotransfer activity. Acts as mitochondria-specific NDK; its association with cardiolipin-containing mitochondrial inner membrane is coupled to respiration suggesting that ADP locally regenerated in the mitochondrion innermembrane space by its activity is directly taken up via ANT ADP/ATP translocase into the matrix space to stimulate respiratory ATP regeneration. Proposed to increase GTP-loading on dynamin-related GTPase OPA1 in mitochondria. In vitro can induce liposome cross-linking suggesting that it can cross-link inner and outer membranes to form contact sites, and promotes intermembrane migration of anionic phosphoplipids. Promotes the redistribution of cardiolipin between the mitochondrial inner membrane and outer membrane which is implicated in pro-apoptotic signaling. |
Generate the functional description in free-text form based on the protein sequence. | MRLHRLRARLSAVACGLLLLLVRGQGQDSASPIRTTHTGQVLGSLVHVKGANAGVQTFLGIPFAKPPLGPLRFAPPEPPESWSGVRDGTTHPAMCLQDLTAVESEFLSQFNMTFPSDSMSEDCLYLSIYTPAHSHEGSNLPVMVWIHGGALVFGMASLYDGSMLAALENVVVVIIQYRLGVLGFFSTGDKHATGNWGYLDQVAALRWVQQNIAHFGGNPDRVTIFGESAGGTSVSSLVVSPISQGLFHGAIMESGVALLPGLIASSADVISTVVANLSACDQVDSEALVGCLRGKSKEEILAINKPFKMIPGVVDGVFLPRHPQELLASADFQPVPSIVGVNNNEFGWLIPKVMRIYDTQKEMDREASQAALQKMLTLLMLPPTFGDLLREEYIGDNGDPQTLQAQFQEMMADSMFVIPALQVAHFQCSRAPVYFYEFQHQPSWLKNIRPPHMKADHGDELPFVFRSFFGGNYIKFTEEEEQLSRKMMKYWANFARNGNPNGEGLPHWPLFDQEEQYLQLNLQPAVGRALKAHRLQFWKKALPQKIQELEEPEERHTEL | Involved in the detoxification of xenobiotics and in the activation of ester and amide prodrugs. Shows high catalytic efficiency for hydrolysis of cocaine, 4-methylumbelliferyl acetate, heroin and 6-monoacetylmorphine. Hydrolyzes aspirin, substrates with large alcohol group and small acyl group and endogenous lipids such as triacylglycerol. Converts monoacylglycerides to free fatty acids and glycerol. Hydrolyzes of 2-arachidonoylglycerol and prostaglandins. |
Can you provide the functional description of the following protein sequence? | MASQNRDPAATSVAAARKGAEPSGGAARGPVGKRLQQELMTLMMSGDKGISAFPESDNLFKWVGTIHGAAGTVYEDLRYKLSLEFPSGYPYNAPTVKFLTPCYHPNVDTQGNICLDILKEKWSALYDVRTILLSIQSLLGEPNIDSPLNTHAAELWKNPTAFKKYLQETYSKQVTSQEP | Accepts ubiquitin from the E1 complex and catalyzes its covalent attachment to other proteins. In vitro catalyzes 'Lys-11'- and 'Lys-48'-linked polyubiquitination. Acts as an essential factor of the anaphase promoting complex/cyclosome (APC/C), a cell cycle-regulated ubiquitin ligase that controls progression through mitosis. Acts by initiating 'Lys-11'-linked polyubiquitin chains on APC/C substrates, leading to the degradation of APC/C substrates by the proteasome and promoting mitotic exit. |
What is the functional description of the protein sequence? | MAAQVAPAAASSLGNPPPPPPSELKKAEQQQREEAGGEAAAAAAAERGEMKAAAGQESEGPAVGPPQPLGKELQDGAESNGGGGGGGAGSGGGPGAEPDLKNSNGNAGPRPALNNNLTEPPGGGGGGSSDGVGAPPHSAAAALPPPAYGFGQPYGRSPSAVAAAAAAVFHQQHGGQQSPGLAALQSGGGGGLEPYAGPQQNSHDHGFPNHQYNSYYPNRSAYPPPAPAYALSSPRGGTPGSGAAAAAGSKPPPSSSASASSSSSSFAQQRFGAMGGGGPSAAGGGTPQPTATPTLNQLLTSPSSARGYQGYPGGDYSGGPQDGGAGKGPADMASQCWGAAAAAAAAAAASGGAQQRSHHAPMSPGSSGGGGQPLARTPQPSSPMDQMGKMRPQPYGGTNPYSQQQGPPSGPQQGHGYPGQPYGSQTPQRYPMTMQGRAQSAMGGLSYTQQIPPYGQQGPSGYGQQGQTPYYNQQSPHPQQQQPPYSQQPPSQTPHAQPSYQQQPQSQPPQLQSSQPPYSQQPSQPPHQQSPAPYPSQQSTTQQHPQSQPPYSQPQAQSPYQQQQPQQPAPSTLSQQAAYPQPQSQQSQQTAYSQQRFPPPQELSQDSFGSQASSAPSMTSSKGGQEDMNLSLQSRPSSLPDLSGSIDDLPMGTEGALSPGVSTSGISSSQGEQSNPAQSPFSPHTSPHLPGIRGPSPSPVGSPASVAQSRSGPLSPAAVPGNQMPPRPPSGQSDSIMHPSMNQSSIAQDRGYMQRNPQMPQYSSPQPGSALSPRQPSGGQIHTGMGSYQQNSMGSYGPQGGQYGPQGGYPRQPNYNALPNANYPSAGMAGGINPMGAGGQMHGQPGIPPYGTLPPGRMSHASMGNRPYGPNMANMPPQVGSGMCPPPGGMNRKTQETAVAMHVAANSIQNRPPGYPNMNQGGMMGTGPPYGQGINSMAGMINPQGPPYSMGGTMANNSAGMAASPEMMGLGDVKLTPATKMNNKADGTPKTESKSKKSSSSTTTNEKITKLYELGGEPERKMWVDRYLAFTEEKAMGMTNLPAVGRKPLDLYRLYVSVKEIGGLTQVNKNKKWRELATNLNVGTSSSAASSLKKQYIQCLYAFECKIERGEDPPPDIFAAADSKKSQPKIQPPSPAGSGSMQGPQTPQSTSSSMAEGGDLKPPTPASTPHSQIPPLPGMSRSNSVGIQDAFNDGSDSTFQKRNSMTPNPGYQPSMNTSDMMGRMSYEPNKDPYGSMRKAPGSDPFMSSGQGPNGGMGDPYSRAAGPGLGNVAMGPRQHYPYGGPYDRVRTEPGIGPEGNMSTGAPQPNLMPSNPDSGMYSPSRYPPQQQQQQQQRHDSYGNQFSTQGTPSGSPFPSQQTTMYQQQQQNYKRPMDGTYGPPAKRHEGEMYSVPYSTGQGQPQQQQLPPAQPQPASQQQAAQPSPQQDVYNQYGNAYPATATAATERRPAGGPQNQFPFQFGRDRVSAPPGTNAQQNMPPQMMGGPIQASAEVAQQGTMWQGRNDMTYNYANRQSTGSAPQGPAYHGVNRTDEMLHTDQRANHEGSWPSHGTRQPPYGPSAPVPPMTRPPPSNYQPPPSMQNHIPQVSSPAPLPRPMENRTSPSKSPFLHSGMKMQKAGPPVPASHIAPAPVQPPMIRRDITFPPGSVEATQPVLKQRRRLTMKDIGTPEAWRVMMSLKSGLLAESTWALDTINILLYDDNSIMTFNLSQLPGLLELLVEYFRRCLIEIFGILKEYEVGDPGQRTLLDPGRFSKVSSPAPMEGGEEEEELLGPKLEEEEEEEVVENDEEIAFSGKDKPASENSEEKLISKFDKLPVKIVQKNDPFVVDCSDKLGRVQEFDSGLLHWRIGGGDTTEHIQTHFESKTELLPSRPHAPCPPAPRKHVTTAEGTPGTTDQEGPPPDGPPEKRITATMDDMLSTRSSTLTEDGAKSSEAIKESSKFPFGISPAQSHRNIKILEDEPHSKDETPLCTLLDWQDSLAKRCVCVSNTIRSLSFVPGNDFEMSKHPGLLLILGKLILLHHKHPERKQAPLTYEKEEEQDQGVSCNKVEWWWDCLEMLRENTLVTLANISGQLDLSPYPESICLPVLDGLLHWAVCPSAEAQDPFSTLGPNAVLSPQRLVLETLSKLSIQDNNVDLILATPPFSRLEKLYSTMVRFLSDRKNPVCREMAVVLLANLAQGDSLAARAIAVQKGSIGNLLGFLEDSLAATQFQQSQASLLHMQNPPFEPTSVDMMRRAARALLALAKVDENHSEFTLYESRLLDISVSPLMNSLVSQVICDVLFLIGQS | Involved in transcriptional activation and repression of select genes by chromatin remodeling (alteration of DNA-nucleosome topology). Component of SWI/SNF chromatin remodeling complexes that carry out key enzymatic activities, changing chromatin structure by altering DNA-histone contacts within a nucleosome in an ATP-dependent manner. Binds DNA non-specifically. Belongs to the neural progenitors-specific chromatin remodeling complex (npBAF complex) and the neuron-specific chromatin remodeling complex (nBAF complex). During neural development a switch from a stem/progenitor to a postmitotic chromatin remodeling mechanism occurs as neurons exit the cell cycle and become committed to their adult state. The transition from proliferating neural stem/progenitor cells to postmitotic neurons requires a switch in subunit composition of the npBAF and nBAF complexes. As neural progenitors exit mitosis and differentiate into neurons, npBAF complexes which contain ACTL6A/BAF53A and PHF10/BAF45A, are exchanged for homologous alternative ACTL6B/BAF53B and DPF1/BAF45B or DPF3/BAF45C subunits in neuron-specific complexes (nBAF). The npBAF complex is essential for the self-renewal/proliferative capacity of the multipotent neural stem cells. The nBAF complex along with CREST plays a role regulating the activity of genes essential for dendrite growth (By similarity). |
Predict the general functional description of the protein sequence. | MERIPSAQPPPACLPKAPGLEHGDLPGMYPAHMYQVYKSRRGIKRSEDSKETYKLPHRLIEKKRRDRINECIAQLKDLLPEHLKLTTLGHLEKAVVLELTLKHVKALTNLIDQQQQKIIALQSGLQAGELSGRNVETGQEMFCSGFQTCAREVLQYLAKHENTRDLKSSQLVTHLHRVVSELLQGGTSRKPSDPAPKVMDFKEKPSSPAKGSEGPGKNCVPVIQRTFAHSSGEQSGSDTDTDSGYGGESEKGDLRSEQPCFKSDHGRRFTMGERIGAIKQESEEPPTKKNRMQLSDDEGHFTSSDLISSPFLGPHPHQPPFCLPFYLIPPSATAYLPMLEKCWYPTSVPVLYPGLNASAAALSSFMNPDKISAPLLMPQRLPSPLPAHPSVDSSVLLQALKPIPPLNLETKD | Transcriptional repressor involved in the regulation of the circadian rhythm by negatively regulating the activity of the clock genes and clock-controlled genes. Acts as the negative limb of a novel autoregulatory feedback loop (DEC loop) which differs from the one formed by the PER and CRY transcriptional repressors (PER/CRY loop). Both these loops are interlocked as it represses the expression of PER1/2 and in turn is repressed by PER1/2 and CRY1/2. Represses the activity of the circadian transcriptional activator: CLOCK-BMAL1|BMAL2 heterodimer by competing for the binding to E-box elements (5'-CACGTG-3') found within the promoters of its target genes. Negatively regulates its own expression and the expression of DBP and BHLHE41/DEC2. Acts as a corepressor of RXR and the RXR-LXR heterodimers and represses the ligand-induced RXRA and NR1H3/LXRA transactivation activity. May be involved in the regulation of chondrocyte differentiation via the cAMP pathway. Represses the transcription of NR0B2 and attentuates the transactivation of NR0B2 by the CLOCK-BMAL1 complex. Drives the circadian rhythm of blood pressure through transcriptional repression of ATP1B1 in the cardiovascular system. |
Predict the functional description of the protein sequence below. | MTLRCLEPSGNGGEGTRSQWGTAGSAEEPSPQAARLAKALRELGQTGWYWGSMTVNEAKEKLKEAPEGTFLIRDSSHSDYLLTISVKTSAGPTNLRIEYQDGKFRLDSIICVKSKLKQFDSVVHLIDYYVQMCKDKRTGPEAPRNGTVHLYLTKPLYTSAPSLQHLCRLTINKCTGAIWGLPLPTRLKDYLEEYKFQV | SOCS family proteins form part of a classical negative feedback system that regulates cytokine signal transduction. SOCS2 appears to be a negative regulator in the growth hormone/IGF1 signaling pathway. Probable substrate recognition component of a SCF-like ECS (Elongin BC-CUL2/5-SOCS-box protein) E3 ubiquitin-protein ligase complex which mediates the ubiquitination and subsequent proteasomal degradation of target proteins. |
Predict the functional description of the protein sequence below. | MSYKPNLAAHMPAAALNAAGSVHSPSTSMATSSQYRQLLSDYGPPSLGYTQGTGNSQVPQSKYAELLAIIEELGKEIRPTYAGSKSAMERLKRGIIHARGLVRECLAETERNARS | Inhibitor of cyclin-dependent kinase CDK2 (By similarity). Also acts as a component of the histone deacetylase NuRD complex which participates in the remodeling of chromatin. |
What is the functional description of the protein sequence? | MAGGMKVAVSPAVGPGPWGSGVGGGGTVRLLLILSGCLVYGTAETDVNVVMLQESQVCEKRASQQFCYTNVLIPKWHDIWTRIQIRVNSSRLVRVTQVENEEKLKELEQFSIWNFFSSFLKEKLNDTYVNVGLYSTKTCLKVEIIEKDTKYSVIVIRRFDPKLFLVFLLGLMLFFCGDLLSRSQIFYYSTGMTVGIVASLLIIIFILSKFMPKKSPIYVILVGGWSFSLYLIQLVFKNLQEIWRCYWQYLLSYVLTVGFMSFAVCYKYGPLENERSINLLTWTLQLMGLCFMYSGIQIPHIALAIIIIALCTKNLEHPIQWLYITCRKVCKGAEKPVPPRLLTEEEYRIQGEVETRKALEELREFCNSPDCSAWKTVSRIQSPKRFADFVEGSSHLTPNEVSVHEQEYGLGSIIAQDEIYEEASSEEEDSYSRCPAITQNNFLT | Together with EMD, contributes to nuclear envelope stiffness in germ cells. Required for female fertility (By similarity). |
Predict the general functional description of the protein sequence. | MASRVLSAYVSRLPAAFAPLPRVRMLAVARPLSTALCSAGTQTRLGTLQPALVLAQVPGRVTQLCRQYSDMPPLTLEGIQDRVLYVLKLYDKIDPEKLSVNSHFMKDLGLDSLDQVEIIMAMEDEFGFEIPDIDAEKLMCPQEIVDYIADKKDVYE | Carrier of the growing fatty acid chain in fatty acid biosynthesis (By similarity). Accessory and non-catalytic subunit of the mitochondrial membrane respiratory chain NADH dehydrogenase (Complex I), which functions in the transfer of electrons from NADH to the respiratory chain. Accessory protein, of the core iron-sulfur cluster (ISC) assembly complex, that regulates, in association with LYRM4, the stability and the cysteine desulfurase activity of NFS1 and participates in the [2Fe-2S] clusters assembly on the scaffolding protein ISCU. The core iron-sulfur cluster (ISC) assembly complex is involved in the de novo synthesis of a [2Fe-2S] cluster, the first step of the mitochondrial iron-sulfur protein biogenesis. This process is initiated by the cysteine desulfurase complex (NFS1:LYRM4:NDUFAB1) that produces persulfide which is delivered on the scaffold protein ISCU in a FXN-dependent manner. Then this complex is stabilized by FDX2 which provides reducing equivalents to accomplish the [2Fe-2S] cluster assembly. Finally, the [2Fe-2S] cluster is transferred from ISCU to chaperone proteins, including HSCB, HSPA9 and GLRX5 (By similarity). |
Can you provide the functional description of the following protein sequence? | MSTKVPIYLKRGSRKGKKEKLRDLLSSDMISPPLGDFRHTIHIGSGGGSDMFGDISFLQGKFHLLPGTMVEGPEEDGTFDLPFQFTRTATVCGRELPDGPSPLLKNAISLPVIGGPQALTLPTAQAPPKPPRLHLETPQPSPQEGGSVDIWRIPETGSPNSGLTPESGAEEPFLSNASSLLSLHVDLGPSILDDVLQIMDQDLDSMQIPT | Probably involved in the organization of the actin cytoskeleton. May act downstream of CDC42 to induce actin filament assembly leading to cell shape changes. Induces pseudopodia formation in fibroblasts in a CDC42-dependent manner. |
Can you provide the functional description of the following protein sequence? | MASDSGNQGTLCTLEFAVQMTCQSCVDAVRKSLQGVAGVQDVEVHLEDQMVLVHTTLPSQEVQALLEGTGRQAVLKGMGSGQLQNLGAAVAILGGPGTVQGVVRFLQLTPERCLIEGTIDGLEPGLHGLHVHQYGDLTNNCNSCGNHFNPDGASHGGPQDSDRHRGDLGNVRADADGRAIFRMEDEQLKVWDVIGRSLIIDEGEDDLGRGGHPLSKITGNSGERLACGIIARSAGLFQNPKQICSCDGLTIWEERGRPIAGKGRKESAQPPAHL | Delivers copper to copper zinc superoxide dismutase (SOD1). |
What is the functional description of the protein sequence? | MTLLTFRDVAIEFSLEEWKCLDLAQQNLYRDVMLENYRNLFSVGLTVCKPGLITCLEQRKEPWNVKRQEAADGHPEMGFHHATQACLELLGSSDLPASASQSAGITGVNHRAQPGLNVSVDKFTALCSPGVLQTVKWFLEFRCIFSLAMSSHFTQDLLPEQGIQDAFPKRILRGYGNCGLDNLYLRKDWESLDECKLQKDYNGLNQCSSTTHSKIFQYNKYVKIFDNFSNLHRRNISNTGEKPFKCQECGKSFQMLSFLTEHQKIHTGKKFQKCGECGKTFIQCSHFTEPENIDTGEKPYKCQECNNVIKTCSVLTKNRIYAGGEHYRCEEFGKVFNQCSHLTEHEHGTEEKPCKYEECSSVFISCSSLSNQQMILAGEKLSKCETWYKGFNHSPNPSKHQRNEIGGKPFKCEECDSIFKWFSDLTKHKRIHTGEKPYKCDECGKAYTQSSHLSEHRRIHTGEKPYQCEECGKVFRTCSSLSNHKRTHSEEKPYTCEECGNIFKQLSDLTKHKKTHTGEKPYKCDECGKNFTQSSNLIVHKRIHTGEKPYKCEECGRVFMWFSDITKHKKTHTGEKPYKCDECGKNFTQSSNLIVHKRIHTGEKPYKCEKCGKAFTQFSHLTVHESIHT | May be involved in transcriptional regulation. |
What is the functional description of the protein sequence? | MESTLTLATEQPVKKNTLKKYKIACIVLLALLVIMSLGLGLGLGLRKLEKQGSCRKKCFDASFRGLENCRCDVACKDRGDCCWDFEDTCVESTRIWMCNKFRCGETRLEASLCSCSDDCLQRKDCCADYKSVCQGETSWLEENCDTAQQSQCPEGFDLPPVILFSMDGFRAEYLYTWDTLMPNINKLKTCGIHSKYMRAMYPTKTFPNHYTIVTGLYPESHGIIDNNMYDVNLNKNFSLSSKEQNNPAWWHGQPMWLTAMYQGLKAATYFWPGSEVAINGSFPSIYMPYNGSVPFEERISTLLKWLDLPKAERPRFYTMYFEEPDSSGHAGGPVSARVIKALQVVDHAFGMLMEGLKQRNLHNCVNIILLADHGMDQTYCNKMEYMTDYFPRINFFYMYEGPAPRIRAHNIPHDFFSFNSEEIVRNLSCRKPDQHFKPYLTPDLPKRLHYAKNVRIDKVHLFVDQQWLAVRSKSNTNCGGGNHGYNNEFRSMEAIFLAHGPSFKEKTEVEPFENIEVYNLMCDLLRIQPAPNNGTHGSLNHLLKVPFYEPSHAEEVSKFSVCGFANPLPTESLDCFCPHLQNSTQLEQVNQMLNLTQEEITATVKVNLPFGRPRVLQKNVDHCLLYHREYVSGFGKAMRMPMWSSYTVPQLGDTSPLPPTVPDCLRADVRVPPSESQKCSFYLADKNITHGFLYPPASNRTSDSQYDALITSNLVPMYEEFRKMWDYFHSVLLIKHATERNGVNVVSGPIFDYNYDGHFDAPDEITKHLANTDVPIPTHYFVVLTSCKNKSHTPENCPGWLDVLPFIIPHRPTNVESCPEGKPEALWVEERFTAHIARVRDVELLTGLDFYQDKVQPVSEILQLKTYLPTFETTI | Hydrolase that metabolizes extracellular nucleotides, including ATP, GTP, UTP and CTP. Limits mast cell and basophil responses during inflammation and during the chronic phases of allergic responses by eliminating the extracellular ATP that functions as signaling molecule and activates basophils and mast cells and induces the release of inflammatory cytokines. Metabolizes extracellular ATP in the lumen of the small intestine, and thereby prevents ATP-induced apoptosis of intestinal plasmacytoid dendritic cells (By similarity). Has also alkaline phosphodiesterase activity. |
What is the functional description of the protein sequence? | MAETKIIYHMDEEETPYLVKLPVAPERVTLADFKNVLSNRPVHAYKFFFKSMDQDFGVVKEEIFDDNAKLPCFNGRVVSWLVLAEGAHSDAGSQGTDSHTDLPPPLERTGGIGDSRPPSFHPNVASSRDGMDNETGTESMVSHRRERARRRNREEAARTNGHPRGDRRRDVGLPPDSASTALSSELESSSFVDSDEDGSTSRLSSSTEQSTSSRLIRKHKRRRRKQRLRQADRASSFSSITDSTMSLNIVTVTLNMERHHFLGISIVGQSNDRGDGGIYIGSIMKGGAVAADGRIEPGDMLLQVNDVNFENMSNDDAVRVLREIVSQTGPISLTVAKCWDPTPRSYFTVPRADPVRPIDPAAWLSHTAALTGALPRYGTSPCSSAVTRTSSSSLTSSVPGAPQLEEAPLTVKSDMSAVVRVMQLPDSGLEIRDRMWLKITIANAVIGADVVDWLYTHVEGFKERREARKYASSLLKHGFLRHTVNKITFSEQCYYVFGDLCSNLATLNLNSGSSGTSDQDTLAPLPHPAAPWPLGQGYPYQYPGPPPCFPPAYQDPGFSYGSGSTGSQQSEGSKSSGSTRSSRRAPGREKERRAAGAGGSGSESDHTAPSGVGSSWRERPAGQLSRGSSPRSQASATAPGLPPPHPTTKAYTVVGGPPGGPPVRELAAVPPELTGSRQSFQKAMGNPCEFFVDIM | Participates in Wnt signaling by binding to the cytoplasmic C-terminus of frizzled family members and transducing the Wnt signal to down-stream effectors. Plays a role both in canonical and non-canonical Wnt signaling. Plays a role in the signal transduction pathways mediated by multiple Wnt genes. Required for LEF1 activation upon WNT1 and WNT3A signaling. DVL1 and PAK1 form a ternary complex with MUSK which is important for MUSK-dependent regulation of AChR clustering during the formation of the neuromuscular junction (NMJ). |
Generate the functional description in free-text form based on the protein sequence. | MLRAGWLRGAAALALLLAARVVAAFEPITVGLAIGAASAITGYLSYNDIYCRFAECCREERPLNASALKLDLEEKLFGQHLATEVIFKALTGFRNNKNPKKPLTLSLHGWAGTGKNFVSQIVAENLHPKGLKSNFVHLFVSTLHFPHEQKIKLYQDQLQKWIRGNVSACANSVFIFDEMDKLHPGIIDAIKPFLDYYEQVDGVSYRKAIFIFLSNAGGDLITKTALDFWRAGRKREDIQLKDLEPVLSVGVFNNKHSGLWHSGLIDKNLIDYFIPFLPLEYRHVKMCVRAEMRARGSAIDEDIVTRVAEEMTFFPRDEKIYSDKGCKTVQSRLDFH | May serve as a molecular chaperone assisting in the proper folding of secreted and/or membrane proteins. Plays a role in non-neural cells nuclear envelope and endoplasmic reticulum integrity. May have a redundant function with TOR1A in non-neural tissues. |
What is the functional description of the protein sequence? | MAAMAVGGAGGSRVSSGRDLNCVPEIADTLGAVAKQGFDFLCMPVFHPRFKREFIQEPAKNRPGPQTRSDLLLSGRDWNTLIVGKLSPWIRPDSKVEKIRRNSEAAMLQELNFGAYLGLPAFLLPLNQEDNTNLARVLTNHIHTGHHSSMFWMRVPLVAPEDLRDDIIENAPTTHTEEYSGEEKTWMWWHNFRTLCDYSKRIAVALEIGADLPSNHVIDRWLGEPIKAAILPTSIFLTNKKGFPVLSKMHQRLIFRLLKLEVQFIITGTNHHSEKEFCSYLQYLEYLSQNRPPPNAYELFAKGYEDYLQSPLQPLMDNLESQTYEVFEKDPIKYSQYQQAIYKCLLDRVPEEEKDTNVQVLMVLGAGRGPLVNASLRAAKQADRRIKLYAVEKNPNAVVTLENWQFEEWGSQVTVVSSDMREWVAPEKADIIVSELLGSFADNELSPECLDGAQHFLKDDGVSIPGEYTSFLAPISSSKLYNEVRACREKDRDPEAQFEMPYVVRLHNFHQLSAPQPCFTFSHPNRDPMIDNNRYCTLEFPVEVNTVLHGFAGYFETVLYQDITLSIRPETHSPGMFSWFPILFPIKQPITVREGQTICVRFWRCSNSKKVWYEWAVTAPVCSAIHNPTGRSYTIGL | Arginine methyltransferase that can both catalyze the formation of omega-N monomethylarginine (MMA) and symmetrical dimethylarginine (sDMA), with a preference for the formation of MMA. Specifically mediates the symmetrical dimethylation of arginine residues in the small nuclear ribonucleoproteins Sm D1 (SNRPD1) and Sm D3 (SNRPD3); such methylation being required for the assembly and biogenesis of snRNP core particles. Methylates SUPT5H and may regulate its transcriptional elongation properties. May methylate the N-terminal region of MBD2. Mono- and dimethylates arginine residues of myelin basic protein (MBP) in vitro. May play a role in cytokine-activated transduction pathways. Negatively regulates cyclin E1 promoter activity and cellular proliferation. Methylates histone H2A and H4 'Arg-3' during germ cell development (By similarity). Methylates histone H3 'Arg-8', which may repress transcription (By similarity). Methylates the Piwi proteins (PIWIL1, PIWIL2 and PIWIL4), methylation of Piwi proteins being required for the interaction with Tudor domain-containing proteins and subsequent localization to the meiotic nuage (By similarity). Methylates RPS10. Attenuates EGF signaling through the MAPK1/MAPK3 pathway acting at 2 levels. First, monomethylates EGFR; this enhances EGFR 'Tyr-1197' phosphorylation and PTPN6 recruitment, eventually leading to reduced SOS1 phosphorylation. Second, methylates RAF1 and probably BRAF, hence destabilizing these 2 signaling proteins and reducing their catalytic activity. Required for induction of E-selectin and VCAM-1, on the endothelial cells surface at sites of inflammation. Methylates HOXA9. Methylates and regulates SRGAP2 which is involved in cell migration and differentiation. Acts as a transcriptional corepressor in CRY1-mediated repression of the core circadian component PER1 by regulating the H4R3 dimethylation at the PER1 promoter (By similarity). Methylates GM130/GOLGA2, regulating Golgi ribbon formation. Methylates H4R3 in genes involved in glioblastomagenesis in a CHTOP- and/or TET1-dependent manner. Symmetrically methylates POLR2A, a modification that allows the recruitment to POLR2A of proteins including SMN1/SMN2 and SETX. This is required for resolving RNA-DNA hybrids created by RNA polymerase II, that form R-loop in transcription terminal regions, an important step in proper transcription termination. Along with LYAR, binds the promoter of gamma-globin HBG1/HBG2 and represses its expression. Symmetrically methylates NCL. Methylates p53/TP53; methylation might possibly affect p53/TP53 target gene specificity. Involved in spliceosome maturation and mRNA splicing in prophase I spermatocytes through the catalysis of the symmetrical arginine dimethylation of SNRPB (small nuclear ribonucleoprotein-associated protein) and the interaction with tudor domain-containing protein TDRD6 (By similarity). |
Can you provide the functional description of the following protein sequence? | MWLYLAAFVGLYYLLHWYRERQVVSHLQDKYVFITGCDSGFGNLLARQLDARGLRVLAACLTEKGAEQLRGQTSDRLETVTLDVTKMESIAAATQWVKEHVGDRGLWGLVNNAGILTPITLCEWLNTEDSMNMLKVNLIGVIQVTLSMLPLVRRARGRIVNVSSILGRVAFFVGGYCVSKYGVEAFSDILRREIQHFGVKISIVEPGYFRTGMTNMTQSLERMKQSWKEAPKHIKETYGQQYFDALYNIMKEGLLNCSTNLNLVTDCMEHALTSVHPRTRYSAGWDAKFFFIPLSYLPTSLADYILTRSWPKPAQAV | NAD-dependent oxidoreductase with broad substrate specificity that shows both oxidative and reductive activity (in vitro). Has 17-beta-hydroxysteroid dehydrogenase activity towards various steroids (in vitro). Converts 5-alpha-androstan-3-alpha,17-beta-diol to androsterone and estradiol to estrone (in vitro). Has 3-alpha-hydroxysteroid dehydrogenase activity towards androsterone (in vitro). Has retinol dehydrogenase activity towards all-trans-retinol (in vitro). Can convert androsterone to epi-androsterone. Androsterone is first oxidized to 5-alpha-androstane-3,17-dione and then reduced to epi-andosterone. Can act on both C-19 and C-21 3-alpha-hydroxysteroids. |
Predict the general functional description of the protein sequence. | MAVPFVEDWDLVQTLGEGAYGEVQLAVNRVTEEAVAVKIVDMKRAVDCPENIKKEICINKMLNHENVVKFYGHRREGNIQYLFLEYCSGGELFDRIEPDIGMPEPDAQRFFHQLMAGVVYLHGIGITHRDIKPENLLLDERDNLKISDFGLATVFRYNNRERLLNKMCGTLPYVAPELLKRREFHAEPVDVWSCGIVLTAMLAGELPWDQPSDSCQEYSDWKEKKTYLNPWKKIDSAPLALLHKILVENPSARITIPDIKKDRWYNKPLKKGAKRPRVTSGGVSESPSGFSKHIQSNLDFSPVNSASSEENVKYSSSQPEPRTGLSLWDTSPSYIDKLVQGISFSQPTCPDHMLLNSQLLGTPGSSQNPWQRLVKRMTRFFTKLDADKSYQCLKETCEKLGYQWKKSCMNQVTISTTDRRNNKLIFKVNLLEMDDKILVDFRLSKGDGLEFKRHFLKIKGKLIDIVSSQKIWLPAT | Serine/threonine-protein kinase which is required for checkpoint-mediated cell cycle arrest and activation of DNA repair in response to the presence of DNA damage or unreplicated DNA. May also negatively regulate cell cycle progression during unperturbed cell cycles. This regulation is achieved by a number of mechanisms that together help to preserve the integrity of the genome. Recognizes the substrate consensus sequence [R-X-X-S/T]. Binds to and phosphorylates CDC25A, CDC25B and CDC25C. Phosphorylation of CDC25A at 'Ser-178' and 'Thr-507' and phosphorylation of CDC25C at 'Ser-216' creates binding sites for 14-3-3 proteins which inhibit CDC25A and CDC25C. Phosphorylation of CDC25A at 'Ser-76', 'Ser-124', 'Ser-178', 'Ser-279' and 'Ser-293' promotes proteolysis of CDC25A. Phosphorylation of CDC25A at 'Ser-76' primes the protein for subsequent phosphorylation at 'Ser-79', 'Ser-82' and 'Ser-88' by NEK11, which is required for polyubiquitination and degradation of CDCD25A. Inhibition of CDC25 leads to increased inhibitory tyrosine phosphorylation of CDK-cyclin complexes and blocks cell cycle progression. Also phosphorylates NEK6. Binds to and phosphorylates RAD51 at 'Thr-309', which promotes the release of RAD51 from BRCA2 and enhances the association of RAD51 with chromatin, thereby promoting DNA repair by homologous recombination. Phosphorylates multiple sites within the C-terminus of TP53, which promotes activation of TP53 by acetylation and promotes cell cycle arrest and suppression of cellular proliferation. Also promotes repair of DNA cross-links through phosphorylation of FANCE. Binds to and phosphorylates TLK1 at 'Ser-743', which prevents the TLK1-dependent phosphorylation of the chromatin assembly factor ASF1A. This may enhance chromatin assembly both in the presence or absence of DNA damage. May also play a role in replication fork maintenance through regulation of PCNA. May regulate the transcription of genes that regulate cell-cycle progression through the phosphorylation of histones (By similarity). Phosphorylates histone H3.1 (to form H3T11ph), which leads to epigenetic inhibition of a subset of genes (By similarity). May also phosphorylate RB1 to promote its interaction with the E2F family of transcription factors and subsequent cell cycle arrest. Phosphorylates SPRTN, promoting SPRTN recruitment to chromatin. Reduces replication stress and activates the G2/M checkpoint, by phosphorylating and inactivating PABIR1/FAM122A and promoting the serine/threonine-protein phosphatase 2A-mediated dephosphorylation and stabilization of WEE1 levels and activity. |
Predict the functional description of the protein sequence below. | MEQRGQNAPAASGARKRHGPGPREARGARPGPRVPKTLVLVVAAVLLLVSAESALITQQDLAPQQRAAPQQKRSSPSEGLCPPGHHISEDGRDCISCKYGQDYSTHWNDLLFCLRCTRCDSGEVELSPCTTTRNTVCQCEEGTFREEDSPEMCRKCRTGCPRGMVKVGDCTPWSDIECVHKESGTKHSGEVPAVEETVTSSPGTPASPCSLSGIIIGVTVAAVVLIVAVFVCKSLLWKKVLPYLKGICSGGGGDPERVDRSSQRPGAEDNVLNEIVSILQPTQVPEQEMEVQEPAEPTGVNMLSPGESEHLLEPAEAERSQRRRLLVPANEGDPTETLRQCFDDFADLVPFDSWEPLMRKLGLMDNEIKVAKAEAAGHRDTLYTMLIKWVNKTGRDASVHTLLDALETLGERLAKQKIEDHLLSSGKFMYLEGNADSAMS | Receptor for the cytotoxic ligand TNFSF10/TRAIL. The adapter molecule FADD recruits caspase-8 to the activated receptor. The resulting death-inducing signaling complex (DISC) performs caspase-8 proteolytic activation which initiates the subsequent cascade of caspases (aspartate-specific cysteine proteases) mediating apoptosis. Promotes the activation of NF-kappa-B. Essential for ER stress-induced apoptosis. |
What is the functional description of the protein sequence? | MRRASRDYTKYLRGSEEMGGGPGAPHEGPLHAPPPPAPHQPPAASRSMFVALLGLGLGQVVCSVALFFYFRAQMDPNRISEDGTHCIYRILRLHENADFQDTTLESQDTKLIPDSCRRIKQAFQGAVQKELQHIVGSQHIRAEKAMVDGSWLDLAKRSKLEAQPFAHLTINATDIPSGSHKVSLSSWYHDRGWAKISNMTFSNGKLIVNQDGFYYLYANICFRHHETSGDLATEYLQLMVYVTKTSIKIPSSHTLMKGGSTKYWSGNSEFHFYSINVGGFFKLRSGEEISIEVSNPSLLDPDQDATYFGAFKVRDID | Cytokine that binds to TNFRSF11B/OPG and to TNFRSF11A/RANK. Osteoclast differentiation and activation factor. Augments the ability of dendritic cells to stimulate naive T-cell proliferation. May be an important regulator of interactions between T-cells and dendritic cells and may play a role in the regulation of the T-cell-dependent immune response. May also play an important role in enhanced bone-resorption in humoral hypercalcemia of malignancy. Induces osteoclastogenesis by activating multiple signaling pathways in osteoclast precursor cells, chief among which is induction of long lasting oscillations in the intracellular concentration of Ca (2+) resulting in the activation of NFATC1, which translocates to the nucleus and induces osteoclast-specific gene transcription to allow differentiation of osteoclasts. During osteoclast differentiation, in a TMEM64 and ATP2A2-dependent manner induces activation of CREB1 and mitochondrial ROS generation necessary for proper osteoclast generation (By similarity). |
What is the functional description of the protein sequence? | MRAVFIQGAEEHPAAFCYQVNGSCPRTVHTLGIQLVIYLACAAGMLIIVLGNVFVAFAVSYFKALHTPTNFLLLSLALADMFLGLLVLPLSTIRSVESCWFFGDFLCRLHTYLDTLFCLTSIFHLCFISIDRHCAICDPLLYPSKFTVRVALRYILAGWGVPAAYTSLFLYTDVVETRLSQWLEEMPCVGSCQLLLNKFWGWLNFPLFFVPCLIMISLYVKIFVVATRQAQQITTLSKSLAGAAKHERKAAKTLGIAVGIYLLCWLPFTIDTMVDSLLHFITPPLVFDIFIWFAYFNSACNPIIYVFSYQWFRKALKLTLSQKVFSPQTRTVDLYQE | Olfactory receptor specific for trimethylamine, a trace amine. Also activated at lower level by dimethylethylamine. Trimethylamine is a bacterial metabolite found in some animal odors, and to humans it is a repulsive odor associated with bad breath and spoiled food. This receptor is probably mediated by the G(s)-class of G-proteins which activate adenylate cyclase. |
Generate the functional description in free-text form based on the protein sequence. | MDYSYLNSYDSCVAAMEASAYGDFGACSQPGGFQYSPLRPAFPAAGPPCPALGSSNCALGALRDHQPAPYSAVPYKFFPEPSGLHEKRKQRRIRTTFTSAQLKELERVFAETHYPDIYTREELALKIDLTEARVQVWFQNRRAKFRKQERAASAKGAAGAAGAKKGEARCSSEDDDSKESTCSPTPDSTASLPPPPAPGLASPRLSPSPLPVALGSGPGPGPGPQPLKGALWAGVAGGGGGGPGAGAAELLKAWQPAESGPGPFSGVLSSFHRKPGPALKTNLF | May be involved in regulating the specificity of expression of the catecholamine biosynthetic genes. Acts as a transcription activator/factor. Could maintain the noradrenergic phenotype. |
What is the functional description of the protein sequence? | MDTGPDQSYFSGNHWFVFSVYLLTFLVGLPLNLLALVVFVGKLQRRPVAVDVLLLNLTASDLLLLLFLPFRMVEAANGMHWPLPFILCPLSGFIFFTTIYLTALFLAAVSIERFLSVAHPLWYKTRPRLGQAGLVSVACWLLASAHCSVVYVIEFSGDISHSQGTNGTCYLEFRKDQLAILLPVRLEMAVVLFVVPLIITSYCYSRLVWILGRGGSHRRQRRVAGLLAATLLNFLVCFGPYNVSHVVGYICGESPAWRIYVTLLSTLNSCVDPFVYYFSSSGFQADFHELLRRLCGLWGQWQQESSMELKEQKGGEEQRADRPAERKTSEHSQGCGTGGQVACAES | G protein-coupled receptor that is activated by a major product of dietary fiber digestion, the short chain fatty acids (SCFAs), and that plays a role in the regulation of whole-body energy homeostasis and in intestinal immunity. In omnivorous mammals, the short chain fatty acids acetate, propionate and butyrate are produced primarily by the gut microbiome that metabolizes dietary fibers. SCFAs serve as a source of energy but also act as signaling molecules. That G protein-coupled receptor is probably coupled to the pertussis toxin-sensitive, G(i/o)-alpha family of G proteins. Its activation results in the formation of inositol 1,4,5-trisphosphate, the mobilization of intracellular calcium, the phosphorylation of the MAPK3/ERK1 and MAPK1/ERK2 kinases and the inhibition of intracellular cAMP accumulation. Activated by SCFAs and by beta-hydroxybutyrate, a ketone body produced by the liver upon starvation, it inhibits N-type calcium channels and modulates the activity of sympathetic neurons through a signaling cascade involving the beta and gamma subunits of its coupled G protein, phospholipase C and MAP kinases. Thereby, it may regulate energy expenditure through the control of the sympathetic nervous system that controls for instance heart rate. Upon activation by SCFAs accumulating in the intestine, it may also signal to the brain via neural circuits which in turn would regulate intestinal gluconeogenesis. May also control the production of hormones involved in whole-body energy homeostasis. May for instance, regulate blood pressure through renin secretion. May also regulate secretion of the PYY peptide by enteroendocrine cells and control gut motility, intestinal transit rate, and the harvesting of energy from SCFAs produced by gut microbiota. May also indirectly regulate the production of LEP/Leptin, a hormone acting on the CNS to inhibit food intake, in response to the presence of short-chain fatty acids in the intestine. Finally, may also play a role in glucose homeostasis. Besides its role in energy homeostasis, may play a role in intestinal immunity. May mediate the activation of the inflammatory and immune response by SCFAs in the gut, regulating the rapid production of chemokines and cytokines by intestinal epithelial cells. Among SCFAs, the fatty acids containing less than 6 carbons, the most potent activators are probably propionate, butyrate and pentanoate while acetate is a poor activator. |
Predict the functional description of the protein sequence below. | MSYIPGQPVTAVVQRVEIHKLRQGENLILGFSIGGGIDQDPSQNPFSEDKTDKGIYVTRVSEGGPAEIAGLQIGDKIMQVNGWDMTMVTHDQARKRLTKRSEEVVRLLVTRQSLQKAVQQSMLS | May regulate a number of protein-protein interactions by competing for PDZ domain binding sites. Binds CTNNB1 and may thereby act as an inhibitor of the Wnt signaling pathway. Competes with LIN7A for KCNJ4 binding, and thereby promotes KCNJ4 internalization. May play a role in the Rho signaling pathway. May play a role in activation of CDC42 by the viral protein HPV16 E6. |
Can you provide the functional description of the following protein sequence? | MPLGLGRRKKAPPLVENEEAEPGRGGLGVGEPGPLGGGGSGGPQMGLPPPPPALRPRLVFHTQLAHGSPTGRIEGFTNVKELYGKIAEAFRLPTAEVMFCTLNTHKVDMDKLLGGQIGLEDFIFAHVKGQRKEVEVFKSEDALGLTITDNGAGYAFIKRIKEGSVIDHIHLISVGDMIEAINGQSLLGCRHYEVARLLKELPRGRTFTLKLTEPRKAFDMISQRSAGGRPGSGPQLGTGRGTLRLRSRGPATVEDLPSAFEEKAIEKVDDLLESYMGIRDTELAATMVELGKDKRNPDELAEALDERLGDFAFPDEFVFDVWGAIGDAKVGRY | May be involved in G protein-linked signaling. |
Generate the functional description in free-text form based on the protein sequence. | MLKPSVTSAPTADMATLTVVQPLTLDRDVARAIELLEKLQESGEVPVHKLQSLKKVLQSEFCTAIREVYQYMHETITVNGCPEFRARATAKATVAAFAASEGHSHPRVVELPKTDEGLGFNVMGGKEQNSPIYISRIIPGGVAERHGGLKRGDQLLSVNGVSVEGEHHEKAVELLKAAKDSVKLVVRYTPKVLEEMEARFEKLRTARRRQQQQLLIQQQQQQQQQQTQQNHMS | Plays a role in establishing and maintaining the asymmetric distribution of channels and receptors at the plasma membrane of polarized cells. Forms membrane-associated multiprotein complexes that may regulate delivery and recycling of proteins to the correct membrane domains. The tripartite complex composed of LIN7 (LIN7A, LIN7B or LIN7C), CASK and APBA1 associates with the motor protein KIF17 to transport vesicles containing N-methyl-D-aspartate (NMDA) receptor subunit NR2B along microtubules (By similarity). This complex may have the potential to couple synaptic vesicle exocytosis to cell adhesion in brain. Ensures the proper localization of GRIN2B (subunit 2B of the NMDA receptor) to neuronal postsynaptic density and may function in localizing synaptic vesicles at synapses where it is recruited by beta-catenin and cadherin. Required to localize Kir2 channels, GABA transporter (SLC6A12) and EGFR/ERBB1, ERBB2, ERBB3 and ERBB4 to the basolateral membrane of epithelial cells. |
Predict the functional description of the protein sequence below. | MEGGGGSGNKTTGGLAGFFGAGGAGYSHADLAGVPLTGMNPLSPYLNVDPRYLVQDTDEFILPTGANKTRGRFELAFFTIGGCCMTGAAFGAMNGLRLGLKETQNMAWSKPRNVQILNMVTRQGALWANTLGSLALLYSAFGVIIEKTRGAEDDLNTVAAGTMTGMLYKCTGGLRGIARGGLTGLTLTSLYALYNNWEHMKGSLLQQSL | Essential component of the TIM23 complex, a complex that mediates the translocation of transit peptide-containing proteins across the mitochondrial inner membrane. |
Can you provide the functional description of the following protein sequence? | MPTNGLHQVLKIQFGLVNDTDRYLTAESFGFKVNASAPSLKRKQTWVLEPDPGQGTAVLLRSSHLGRYLSAEEDGRVACEAEQPGRDCRFLVLPQPDGRWVLRSEPHGRFFGGTEDQLSCFATAVSPAELWTVHLAIHPQAHLLSVSRRRYVHLCPREDEMAADGDKPWGVDALLTLIFRSRRYCLKSCDSRYLRSDGRLVWEPEPRACYTLEFKAGKLAFKDCDGHYLAPVGPAGTLKAGRNTRPGKDELFDLEESHPQVVLVAANHRYVSVRQGVNVSANQDDELDHETFLMQIDQETKKCTFYSSTGGYWTLVTHGGIHATATQVSANTMFEMEWRGRRVALKASNGRYVCMKKNGQLAAISDFVGKDEEFTLKLINRPILVLRGLDGFVCHHRGSNQLDTNRSVYDVFHLSFSDGAYRIRGRDGGFWYTGSHGSVCSDGERAEDFVFEFRERGRLAIRARSGKYLRGGASGLLRADADAPAGTALWEY | Acts as an actin bundling protein. May play a pivotal role in photoreceptor cell-specific events, such as disk morphogenesis. |
What is the functional description of the protein sequence? | MMASMRVVKELEDLQKKPPPYLRNLSSDDANVLVWHALLLPDQPPYHLKAFNLRISFPPEYPFKPPMIKFTTKIYHPNVDENGQICLPIISSENWKPCTKTCQVLEALNVLVNRPNIREPLRMDLADLLTQNPELFRKNAEEFTLRFGVDRPS | Catalyzes the covalent attachment of ubiquitin or ISG15 to other proteins. Functions in the E6/E6-AP-induced ubiquitination of p53/TP53. Promotes ubiquitination and subsequent proteasomal degradation of FLT3. |
Github:
https://github.com/baaihealth/opi
Paper:
OPI: An Open Instruction Dataset for Adapting Large Language Models to Protein-Related Tasks has been accepted by NeurIPS 2024 Workshop: Foundation Models for Science: Progress, Opportunities, and Challenges.
Dataset Overview
Dataset size:
- Thera are 1.64M samples, including training (1,615,661) and testing (26,607) sets, in OPI dataset, covering 9 protein-related tasks.
We are excited to announce the release of the Open Protein Instructions (OPI) dataset, a curated collection of instructions covering 9 tasks for adapting LLMs to protein biology. The dataset is designed to advance LLM-driven research in the field of protein biology. We welcome contributions and enhancements to this dataset from the community.
OPI is the initial part of Open Biology Instructions(OBI) project, together with the subsequent Open Molecule Instructions(OMI), Open DNA Instructions(ODI), Open RNA Instructions(ORI) and Open Single-cell Instructions (OSCI). OBI is a project which aims to fully leverage the potential ability of Large Language Models(LLMs), especially the scientific LLMs like Galactica, to facilitate research in AI for Life Science community. While OBI is still in an early stage, we hope to provide a starting point for the community to bridge LLMs and biological domain knowledge.
Dataset Update
The previous version of OPI dataset is based on the release 2022_01 of UniProtKB/Swiss-Prot protein knowledgebase. At current, OPI is updated to contain the latest release 2023_05, which can be accessed via the dataset file OPI_updated_160k.json.
Reference:
- https://ftp.uniprot.org/pub/databases/uniprot/previous_releases/release-2022_01/knowledgebase/UniProtKB_SwissProt-relstat.html
- https://ftp.uniprot.org/pub/databases/uniprot/previous_releases/release-2023_05/knowledgebase/UniProtKB_SwissProt-relstat.html
OPI Dataset Construction Pipeline
The OPI dataset is curated on our own by extracting key information from Swiss-Prot database. The following figure shows the general construction process.
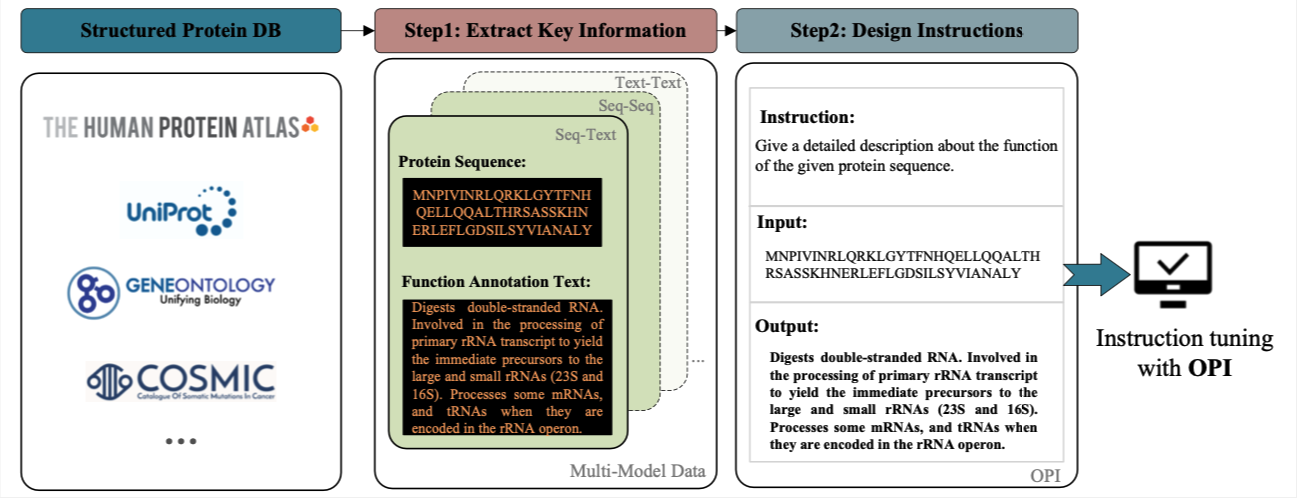
OPI Dataset Folder Structure
The OPI dataset is organized into the three subfolders—AP, KM, and SU—by in the OPI_DATA directory within this repository, where you can find a subset for each specific task as well as the full dataset file: OPI_full_1.61M_train.json.
./OPI_DATA/
└── SU
│ ├── EC_number
│ │ ├── test
│ │ │ ├── CLEAN_EC_number_new_test.jsonl
│ │ │ └── CLEAN_EC_number_price_test.jsonl
│ │ └── train
│ │ ├── CLEAN_EC_number_train.json
│ ├── Fold_type
│ │ ├── test
│ │ │ └── fold_type_test.jsonl
│ │ └── train
│ │ └── fold_type_train.json
│ └── Subcellular_localization
│ ├── test
│ │ ├── subcell_loc_test.jsonl
│ └── train
└── subcell_loc_train.json
├── AP
│ └── Keywords
│ │ ├── test
│ │ │ ├── CASPSimilarSeq_keywords_test.jsonl
│ │ │ ├── IDFilterSeq_keywords_test.jsonl
│ │ │ └── UniProtSeq_keywords_test.jsonl
│ │ └── train
│ │ ├── keywords_train.json
│ ├── GO
│ │ ├── test
│ │ │ ├── CASPSimilarSeq_go_terms_test.jsonl
│ │ │ ├── IDFilterSeq_go_terms_test.jsonl
│ │ │ └── UniProtSeq_go_terms_test.jsonl
│ │ └── train
│ │ ├── go_terms_train.json
│ ├── Function
│ ├── test
│ │ ├── CASPSimilarSeq_function_test.jsonl
│ │ ├── IDFilterSeq_function_test.jsonl
│ │ └── UniProtSeq_function_test.jsonl
│ └── train
│ ├── function_train.json
├── KM
└── gSymbol2Tissue
│ ├── test
│ │ └── gene_symbol_to_tissue_test.jsonl
│ └── train
│ └── gene_symbol_to_tissue_train.json
├── gSymbol2Cancer
│ ├── test
│ │ └── gene_symbol_to_cancer_test.jsonl
│ └── train
│ └── gene_symbol_to_cancer_train.json
├── gName2Cancer
├── test
│ └── gene_name_to_cancer_test.jsonl
└── train
└── gene_name_to_cancer_train.json
Dataset Examples
An example of OPI training data:
instruction:
What is the EC classification of the input protein sequence based on its biological function?
input:
MGLVSSKKPDKEKPIKEKDKGQWSPLKVSAQDKDAPPLPPLVVFNHLTPPPPDEHLDEDKHFVVALYDYTAMNDRDLQMLKGEKLQVLKGTGDWWLARS
LVTGREGYVPSNFVARVESLEMERWFFRSQGRKEAERQLLAPINKAGSFLIRESETNKGAFSLSVKDVTTQGELIKHYKIRCLDEGGYYISPRITFPSL
QALVQHYSKKGDGLCQRLTLPCVRPAPQNPWAQDEWEIPRQSLRLVRKLGSGQFGEVWMGYYKNNMKVAIKTLKEGTMSPEAFLGEANVMKALQHERLV
RLYAVVTKEPIYIVTEYMARGCLLDFLKTDEGSRLSLPRLIDMSAQIAEGMAYIERMNSIHRDLRAANILVSEALCCKIADFGLARIIDSEYTAQEGAK
FPIKWTAPEAIHFGVFTIKADVWSFGVLLMEVVTYGRVPYPGMSNPEVIRNLERGYRMPRPDTCPPELYRGVIAECWRSRPEERPTFEFLQSVLEDFYT
ATERQYELQP
output:
2.7.10.2
An example of OPI testing data:
{"id": "seed_task_0", "name": "EC number of price dataset from CLEAN", "instruction":
"Return the EC number of the protein sequence.", "instances": [{"input":
"MAIPPYPDFRSAAFLRQHLRATMAFYDPVATDASGGQFHFFLDDGTVYNTHTRHLVSATRFVVTHAMLYRTTGEARYQVGMRHALEFLRTAFLDPATGGY
AWLIDWQDGRATVQDTTRHCYGMAFVMLAYARAYEAGVPEARVWLAEAFDTAEQHFWQPAAGLYADEASPDWQLTSYRGQNANMHACEAMISAFRATGERR
YIERAEQLAQGICQRQAALSDRTHAPAAEGWVWEHFHADWSVDWDYNRHDRSNIFRPWGYQVGHQTEWAKLLLQLDALLPADWHLPCAQRLFDTAVERGWD
AEHGGLYYGMAPDGSICDDGKYHWVQAESMAAAAVLAVRTGDARYWQWYDRIWAYCWAHFVDHEHGAWFRILHRDNRNTTREKSNAGKVDYHNMGACYDVL
LWALDAPGFSKESRSAALGRP", "output": "5.3.1.7"}], "is_classification": false}
OPEval: Nine evaluation tasks using the OPI dataset
To assess the effectiveness of instruction tuning with the OPI dataset, we developed OPEval, which comprises three categories of evaluation tasks. Each category includes three specific tasks. The table below outlines the task types, names, and the corresponding sizes of the training and testing sets.
| Task Type | Type Abbr. | Task Name | Task Abbr. | Training set size | Testing set size |
|---|---|---|---|---|---|
| Sequence Understanding | SU | EC Number Prediction | EC_number | 227,362 | 392 (NEW-392), 149 (Price-149) |
| Fold Type Prediction | Fold_type | 12,312 | 718 (Fold), 1254 (Superfamily), 1272 (Family) | ||
| Subcellular Localization Prediction | Subcellular_localization | 11,230 | 2,772 | ||
| Annotation Prediction | AP | Function Keywords Prediction | Keywords | 451,618 | 184 (CASPSimilarSeq), 1,112 (IDFilterSeq), 4562 (UniprotSeq) |
| Gene Ontology(GO) Terms Prediction | GO | 451,618 | 184 (CASPSimilarSeq), 1,112 (IDFilterSeq), 4562 (UniprotSeq) | ||
| Function Description Prediction | Function | 451,618 | 184 (CASPSimilarSeq), 1,112 (IDFilterSeq), 4562 (UniprotSeq) | ||
| Knowledge Mining | KM | Tissue Location Prediction from Gene Symbol | gSymbol2Tissue | 8,723 | 2,181 |
| Cancer Prediction from Gene Symbol | gSymbol2Cancer | 590 | 148 | ||
| Cancer Prediction from Gene Name | gName2Cancer | 590 | 148 |
License
The dataset is licensed under a Creative Commons Attribution Non Commercial 4.0 License. The use of this dataset should also abide by the original License & Disclaimer and Privacy Notice of UniProt.
- Downloads last month
- 479
