mhg-gnn
This repository provides PyTorch source code assosiated with our publication, "MHG-GNN: Combination of Molecular Hypergraph Grammar with Graph Neural Network"
Paper: Arxiv Link
Introduction
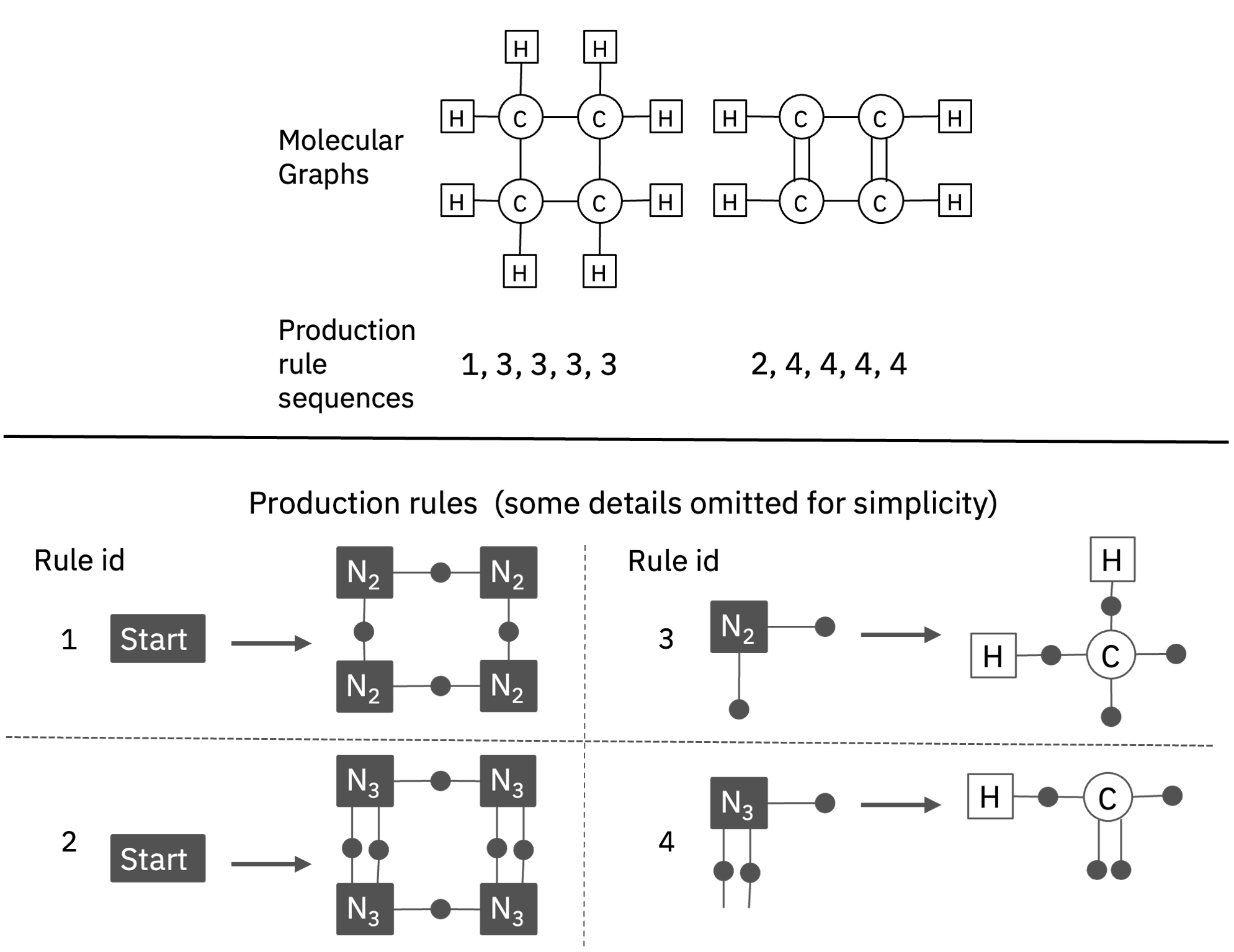
We present MHG-GNN, an autoencoder architecture
that has an encoder based on GNN and a decoder based on a sequential model with MHG.
Since the encoder is a GNN variant, MHG-GNN can accept any molecule as input, and
demonstrate high predictive performance on molecular graph data.
In addition, the decoder inherits the theoretical guarantee of MHG on always generating a structurally valid molecule as output.
Table of Contents
Getting Started
This code and environment have been tested on Intel E5-2667 CPUs at 3.30GHz and NVIDIA A100 Tensor Core GPUs.
Pretrained Models and Training Logs
We provide checkpoints of the MHG-GNN model pre-trained on a dataset of ~1.34M molecules curated from PubChem. (later) For model weights: HuggingFace Link
Add the MHG-GNN pre-trained weights.pt to the models/ directory according to your needs.
Installation
We recommend to create a virtual environment. For example:
python3 -m venv .venv
. .venv/bin/activate
Type the following command once the virtual environment is activated:
git clone git@github.ibm.com:CMD-TRL/mhg-gnn.git
cd ./mhg-gnn
pip install .
Feature Extraction
The example notebook mhg-gnn_encoder_decoder_example.ipynb contains code to load checkpoint files and use the pre-trained model for encoder and decoder tasks.
To load mhg-gnn, you can simply use:
import torch
import load
model = load.load()
To encode SMILES into embeddings, you can use:
with torch.no_grad():
repr = model.encode(["CCO", "O=C=O", "OC(=O)c1ccccc1C(=O)O"])
For decoder, you can use the function, so you can return from embeddings to SMILES strings:
orig = model.decode(repr)
- Downloads last month
- 332