|
--- |
|
|
|
tags: |
|
- feature-extraction |
|
- image-classification |
|
- timm |
|
- biology |
|
- cancer |
|
- owkin |
|
- histology |
|
library_name: timm |
|
model-index: |
|
- name: owkin_pancancer |
|
results: |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: Camelyon16[Meta] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 94.5 ± 4.4 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-BRCA[Hist] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 96.2 ± 3.3 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-BRCA[HRD] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 79.3 ± 2.4 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-BRCA[Mol] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 81.7 ± 1.6 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-BRCA[OS] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 64.7 ± 5.7 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-CRC[MSI] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 91.0 ± 2.2 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-COAD[OS] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 63.4 ± 7.4 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-NSCLC[CType] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 97.7 ± 1.3 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-LUAD[OS] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 53.8 ± 4.5 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-LUSC[OS] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 62.2 ± 2.9 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-OV[HRD] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 74.2 ± 8.6 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-RCC[CType] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 99.5 ± 0.2 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-STAD[MSI] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 89.9 ± 3.9 |
|
name: ROC AUC |
|
verified: false |
|
- task: |
|
type: image-classification |
|
name: Image Classification |
|
dataset: |
|
name: TCGA-PAAD[OS] |
|
type: image-classification |
|
metrics: |
|
- type: accuracy |
|
value: 59.2 ± 4.1 |
|
name: ROC AUC |
|
verified: false |
|
widget: |
|
- src: https://github.com/owkin/HistoSSLscaling/raw/main/assets/example.tif |
|
example_title: pancancer tile |
|
co2_eq_emissions: |
|
emissions: 14590 |
|
source: https://www.medrxiv.org/content/10.1101/2023.07.21.23292757v2 |
|
training_type: pre-training |
|
geographical_location: Jean Zay cluster, France (~40 gCO₂eq/kWh) |
|
hardware_used: 32 V100 32Gb GPUs, 1216 GPU hours |
|
license: other |
|
license_name: owkin-non-commercial |
|
license_link: https://github.com/owkin/HistoSSLscaling/blob/main/LICENSE.txt |
|
pipeline_tag: feature-extraction |
|
inference: false |
|
datasets: |
|
- owkin/camelyon16-features |
|
- owkin/nct-crc-he |
|
metrics: |
|
- roc_auc |
|
--- |
|
|
|
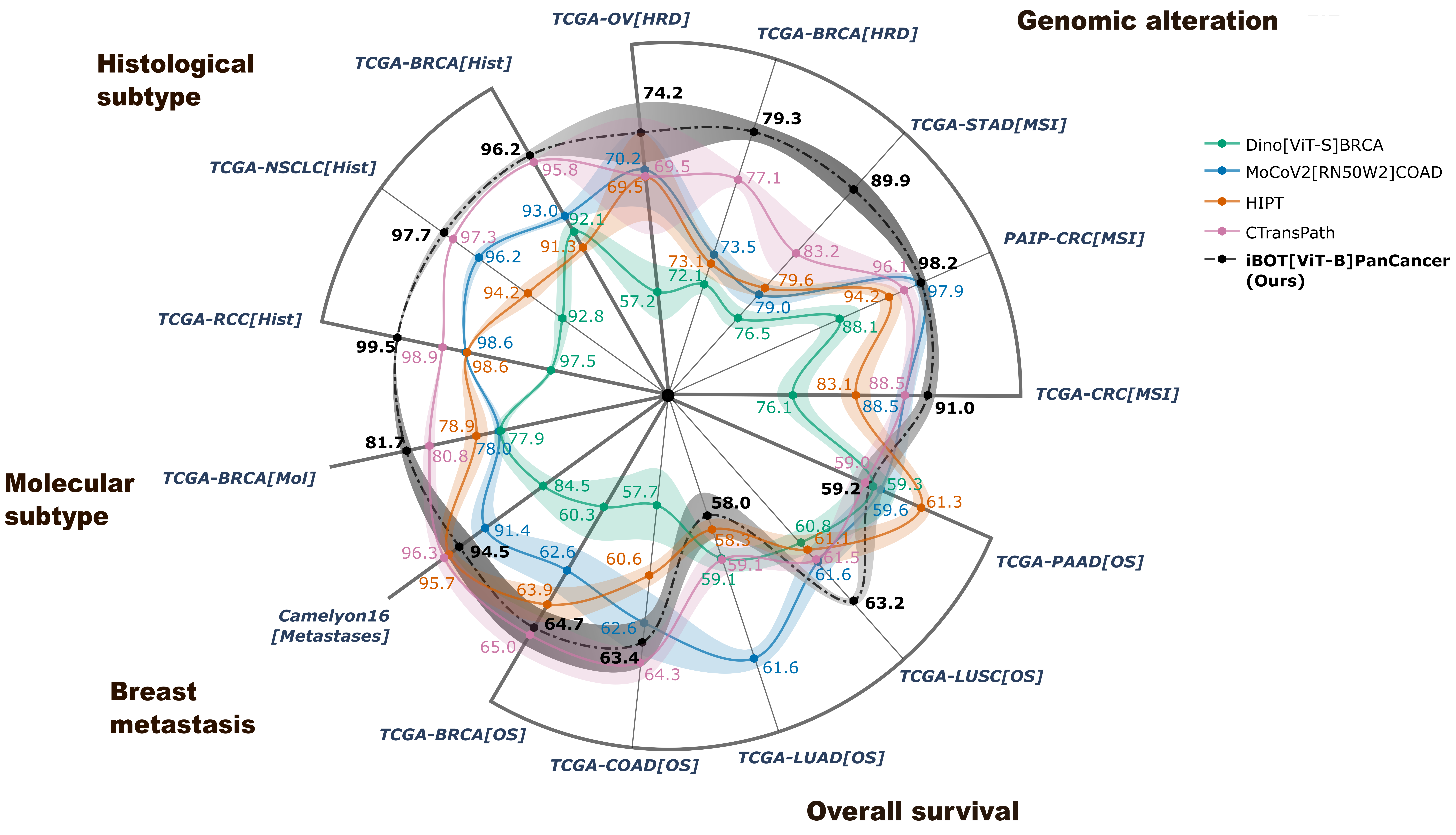
# Model card for vit_base_patch16_224.owkin_pancancer |
|
|
|
A Vision Transformer (ViT) image classification model. \ |
|
Trained by Owkin on 40 million pan-cancer histology tiles from TCGA-COAD. |
|
|
|
A version using the transformers library is also available here: https://huggingface.co/owkin/phikon |
|
|
|
 |
|
|
|
## Model Details |
|
|
|
- **Model Type:** Feature backbone |
|
- **Developed by**: Owkin |
|
- **Funded by**: Owkin and IDRIS |
|
- **Model Stats:** |
|
- Params: 85.8M (base) |
|
- Image size: 224 x 224 x 3 |
|
- Patch size: 16 x 16 x 3 |
|
- **Pre-training:** |
|
- Dataset: Pancancer40M, created from [TCGA-COAD](https://portal.gdc.cancer.gov/repository?facetTab=cases&filters=%7B%22content%22%3A%5B%7B%22content%22%3A%7B%22field%22%3A%22cases.project.project_id%22%2C%22value%22%3A%5B%22TCGA-COAD%22%5D%7D%2C%22op%22%3A%22in%22%7D%2C%7B%22content%22%3A%7B%22field%22%3A%22files.experimental_strategy%22%2C%22value%22%3A%5B%22Diagnostic%20Slide%22%5D%7D%2C%22op%22%3A%22in%22%7D%5D%2C%22op%22%3A%22and%22%7D&searchTableTab=cases) |
|
- Framework: [iBOT](https://github.com/bytedance/ibot), self-supervised, masked image modeling, self-distillation |
|
- **Papers:** |
|
- [Scaling Self-Supervised Learning for Histopathology with Masked Image Modeling](https://www.medrxiv.org/content/10.1101/2023.07.21.23292757v2) |
|
- **Original:** https://github.com/owkin/HistoSSLscaling |
|
- **License:** [Owkin non-commercial license](https://github.com/owkin/HistoSSLscaling/blob/main/LICENSE.txt) |
|
|
|
## Model Usage |
|
|
|
### Image Embeddings |
|
```python |
|
from urllib.request import urlopen |
|
from PIL import Image |
|
import timm |
|
|
|
# get example histology image |
|
img = Image.open( |
|
urlopen( |
|
"https://github.com/owkin/HistoSSLscaling/raw/main/assets/example.tif" |
|
) |
|
) |
|
|
|
# load model from the hub |
|
model = timm.create_model( |
|
model_name="hf-hub:1aurent/vit_base_patch16_224.owkin_pancancer", |
|
pretrained=True, |
|
).eval() |
|
|
|
# get model specific transforms (normalization, resize) |
|
data_config = timm.data.resolve_model_data_config(model) |
|
transforms = timm.data.create_transform(**data_config, is_training=False) |
|
|
|
data = transforms(img).unsqueeze(0) # input is a (batch_size, num_channels, img_size, img_size) shaped tensor |
|
output = model(data) # output is a (batch_size, num_features) shaped tensor |
|
``` |
|
|
|
## Citation |
|
```bibtex |
|
@article{Filiot2023.07.21.23292757, |
|
author = {Alexandre Filiot and Ridouane Ghermi and Antoine Olivier and Paul Jacob and Lucas Fidon and Alice Mac Kain and Charlie Saillard and Jean-Baptiste Schiratti}, |
|
title = {Scaling Self-Supervised Learning for Histopathology with Masked Image Modeling}, |
|
elocation-id = {2023.07.21.23292757}, |
|
year = {2023}, |
|
doi = {10.1101/2023.07.21.23292757}, |
|
publisher = {Cold Spring Harbor Laboratory Press}, |
|
url = {https://www.medrxiv.org/content/early/2023/09/14/2023.07.21.23292757}, |
|
eprint = {https://www.medrxiv.org/content/early/2023/09/14/2023.07.21.23292757.full.pdf}, |
|
journal = {medRxiv} |
|
} |
|
``` |