Spaces:
Sleeping
A newer version of the Gradio SDK is available:
4.36.1
title: Diffdock
emoji: 🐠
colorFrom: indigo
colorTo: pink
sdk: gradio
app_file: app.py
pinned: false
license: mit
duplicated_from: simonduerr/diffdock
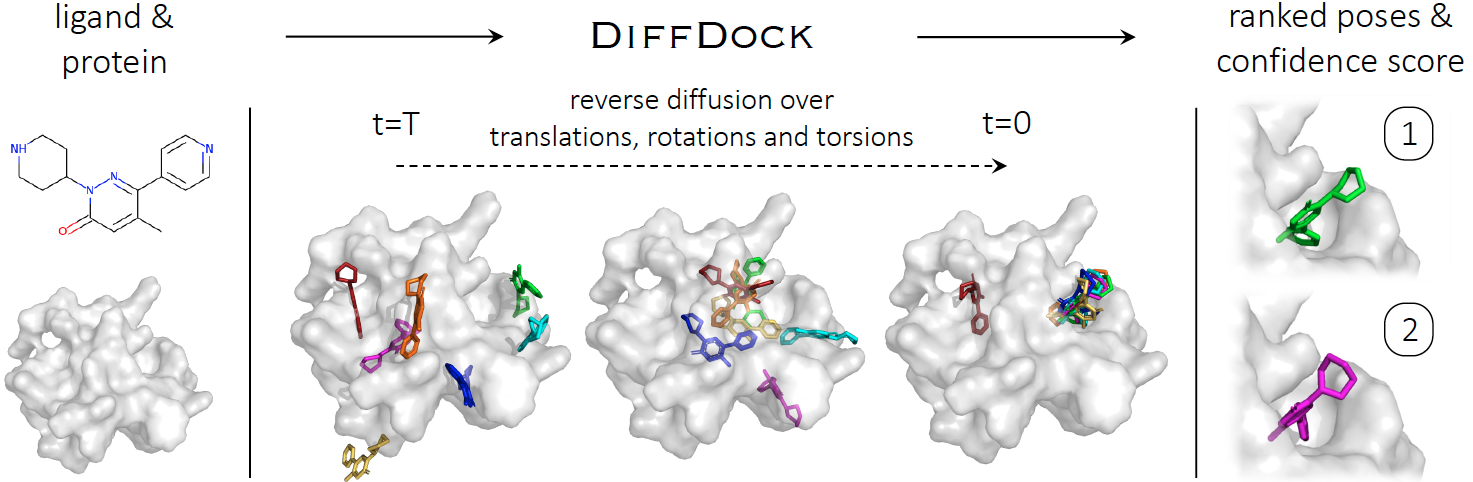
DiffDock: Diffusion Steps, Twists, and Turns for Molecular Docking
Paper on arXiv
Implementation of DiffDock, state-of-the-art method for molecular docking, by Gabriele Corso*, Hannes Stark*, Bowen Jing*, Regina Barzilay and Tommi Jaakkola. This repository contains all code, instructions and model weights necessary to run the method or to retrain a model. If you have any question, feel free to open an issue or reach out to us: gcorso@mit.edu, hstark@mit.edu, bjing@mit.edu.
The repository also contains all the scripts to run the baselines and generate the figures.
Additionally, there are visualization videos in visualizations.
Dataset
The files in data contain the names for the time-based data split.
If you want to train one of our models with the data then:
- download it from zenodo
- unzip the directory and place it into
datasuch that you have the pathdata/PDBBind_processed
Setup Environment
We will set up the environment using Anaconda. Clone the current repo
git clone https://github.com/gcorso/DiffDock.git
Create a new environment with all required packages using environment.yml. While in the project directory run:
conda env create
Activate the environment
conda activate diffdock
If you want to install the packages yourself in case something does not work, these are the required ones:
pytorch
pyg
pyyaml
scipy
networkx
biopython
rdkit-pypi
e3nn
spyrmsd
pandas
biopandas
Running DiffDock on your own complexes
We support multiple input formats depending on whether you only want to make predictions for a single complex or for many at once.
The protein inputs need to be .pdb files. The ligand input can either be a SMILES string or a filetype that RDKit can read like .sdf or .mol2.
For a single complex: specify the protein with, e.g., --protein_path protein.pdb and the ligand with --ligand_path ligand.sdf or --ligand_smiles COc(cc1)ccc1C#N
For many complexes: create a csv file with paths to proteins and ligand files or SMILES. The first column of the .csv has to be called protein_path and the second one ligand.
An example .csv is at data/protein_ligand_example_csv.csv and you would use it with --protein_ligand_csv protein_ligand_example_csv.csv.
Generate the ESM2 embeddings for the proteins
We will soon also provide weights of a trained model without ESM2 embeddings such that this step is not necessary. Luckily, it is rather easy. First prepare a fasta for ESM2 (for a single protein use --protein_path protein.pdb instead):
python datasets/esm_embedding_preparation.py --protein_ligand_csv data/protein_ligand_example_csv.csv --out_file data/prepared_for_esm.fasta
Generate the embeddings with ESM2 (assuming that you are in the DiffDock directory):
git clone https://github.com/facebookresearch/esm
cd esm
pip install -e .
cd ..
HOME=esm/model_weights python esm/scripts/extract.py esm2_t33_650M_UR50D data/prepared_for_esm.fasta data/esm2_output --repr_layers 33 --include per_tok
And done, that is it!
Run inference
python -m inference --protein_ligand_csv data/protein_ligand_example_csv.csv --out_dir results/user_predictions_small --inference_steps 20 --samples_per_complex 40 --batch_size 10
Running DiffDock to reproduce paper numbers
Download the data and place it as described in the "Dataset" section above.
Generate the ESM2 embeddings for the proteins
First run:
python datasets/pdbbind_lm_embedding_preparation.py
Use the generated file data/pdbbind_sequences.fasta to generate the ESM2 language model embeddings using the library https://github.com/facebookresearch/esm by installing their repository and executing the following in their repository:
python scripts/extract.py esm2_t33_650M_UR50D pdbbind_sequences.fasta embeddings_output --repr_layers 33 --include per_tok
This generates the embeddings_output directory which you have to copy into the data folder of our repository to have data/embeddings_output.
Then run the command:
python datasets/esm_embeddings_to_pt.py
Using the provided model weights for evaluation
To predict binding structures using the provided model weights run:
python -m evaluate --model_dir workdir/paper_score_model --ckpt best_ema_inference_epoch_model.pt --confidence_ckpt best_model_epoch75.pt --confidence_model_dir workdir/paper_confidence_model --run_name DiffDockInference --inference_steps 20 --split_path data/splits/timesplit_test --samples_per_complex 40 --batch_size 10
To additionally save the .sdf files of the generated molecules, add the flag --save_visualisation
Training a model yourself and using those weights
Train the large score model:
python -m train --run_name big_score_model --test_sigma_intervals --esm_embeddings_path data/esm2_3billion_embeddings.pt --log_dir workdir --lr 1e-3 --tr_sigma_min 0.1 --tr_sigma_max 19 --rot_sigma_min 0.03 --rot_sigma_max 1.55 --batch_size 16 --ns 48 --nv 10 --num_conv_layers 6 --dynamic_max_cross --scheduler plateau --scale_by_sigma --dropout 0.1 --sampling_alpha 1 --sampling_beta 1 --remove_hs --c_alpha_max_neighbors 24 --receptor_radius 15 --num_dataloader_workers 1 --cudnn_benchmark --rot_alpha 1 --rot_beta 1 --tor_alpha 1 --tor_beta 1 --val_inference_freq 5 --num_inference_complexes 500 --use_ema --distance_embed_dim 64 --cross_distance_embed_dim 64 --sigma_embed_dim 64 --scheduler_patience 30 --n_epochs 850
The model weights are saved in the workdir directory.
Train a small score model with higher maximum translation sigma that will be used to generate the samples for training the confidence model:
python -m train --run_name small_score_model --test_sigma_intervals --esm_embeddings_path data/esm2_3billion_embeddings.pt --log_dir workdir --lr 1e-3 --tr_sigma_min 0.1 --tr_sigma_max 34 --rot_sigma_min 0.03 --rot_sigma_max 1.55 --batch_size 16 --ns 24 --nv 6 --num_conv_layers 5 --dynamic_max_cross --scheduler plateau --scale_by_sigma --dropout 0.1 --sampling_alpha 1 --sampling_beta 1 --remove_hs --c_alpha_max_neighbors 24 --receptor_radius 15 --num_dataloader_workers 1 --cudnn_benchmark --rot_alpha 1 --rot_beta 1 --tor_alpha 1 --tor_beta 1 --val_inference_freq 5 --num_inference_complexes 500 --use_ema --scheduler_patience 30 --n_epochs 300
In practice, you could also likely achieve the same or better results by using the first score model for creating the samples to train the confidence model, but this is what we did in the paper.
The score model used to generate the samples to train the confidence model does not have to be the same as the score model that is used with that confidence model during inference.
Train the confidence model by running the following:
python -m confidence.confidence_train --original_model_dir workdir/small_score_model --run_name confidence_model --inference_steps 20 --samples_per_complex 7 --inf_sched_alpha 1 --inf_sched_beta 1 --batch_size 16 --n_epochs 100 --lr 3e-4 --scheduler_patience 50 --tr_sigma_min 0.1 --tr_sigma_max 34 --rot_sigma_min 0.03 --rot_sigma_max 1.55 --ns 24 --nv 6 --num_conv_layers 5 --dynamic_max_cross --scale_by_sigma --dropout 0.1 --all_atoms --remove_hs --c_alpha_max_neighbors 24 --receptor_radius 15 --esm_embeddings_path data/esm2_3billion_embeddings.pt --main_metric loss --main_metric_goal min --best_model_save_frequency 5 --rmsd_classification_cutoff 2 --cache_creation_id 1 --cache_ids_to_combine 1 2 3 4
first with --cache_creation_id 1 then --cache_creation_id 2 etc. up to 4
Now everything is trained and you can run inference with:
python -m evaluate --model_dir workdir/big_score_model --ckpt best_ema_inference_epoch_model.pt --confidence_ckpt best_model_epoch75.pt --confidence_model_dir workdir/confidence_model --run_name DiffDockInference --inference_steps 20 --split_path data/splits/timesplit_test --samples_per_complex 40 --batch_size 10
Citation
@article{corso2022diffdock,
title={DiffDock: Diffusion Steps, Twists, and Turns for Molecular Docking},
author = {Corso, Gabriele and Stärk, Hannes and Jing, Bowen and Barzilay, Regina and Jaakkola, Tommi},
journal={arXiv preprint arXiv:2210.01776},
year={2022}
}
License
MIT