Replace with clean markdown card
Browse files
README.md
CHANGED
|
@@ -9,19 +9,17 @@ tags:
|
|
| 9 |
- neuroscience
|
| 10 |
- braindecode
|
| 11 |
- convolutional
|
| 12 |
-
- transformer
|
| 13 |
- sleep-staging
|
| 14 |
---
|
| 15 |
|
| 16 |
# USleep
|
| 17 |
|
| 18 |
-
Sleep staging architecture from Perslev et al (2021) .
|
| 19 |
|
| 20 |
-
> **Architecture-only repository.**
|
| 21 |
> `braindecode.models.USleep` class. **No pretrained weights are
|
| 22 |
-
> distributed here**
|
| 23 |
-
> data
|
| 24 |
-
> separately.
|
| 25 |
|
| 26 |
## Quick start
|
| 27 |
|
|
@@ -40,228 +38,46 @@ model = USleep(
|
|
| 40 |
)
|
| 41 |
```
|
| 42 |
|
| 43 |
-
The signal-shape arguments above are
|
| 44 |
-
|
| 45 |
|
| 46 |
## Documentation
|
| 47 |
-
|
| 48 |
-
-
|
| 49 |
-
<https://braindecode.org/stable/generated/braindecode.models.USleep.html>
|
| 50 |
-
- Interactive browser with live instantiation:
|
| 51 |
<https://huggingface.co/spaces/braindecode/model-explorer>
|
| 52 |
- Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/usleep.py#L14>
|
| 53 |
|
| 54 |
-
## Architecture description
|
| 55 |
|
| 56 |
-
|
| 57 |
-
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 58 |
|
| 59 |
-
<div class='bd-doc'><main>
|
| 60 |
-
<p>Sleep staging architecture from Perslev et al (2021) <a class="brackets" href="#footnote-1" id="footnote-reference-1" role="doc-noteref"><span class="fn-bracket">[</span>1<span class="fn-bracket">]</span></a>.</p>
|
| 61 |
-
<span style="display:inline-block;padding:2px 8px;border-radius:4px;background:#5cb85c;color:white;font-size:11px;font-weight:600;margin-right:4px;">Convolution</span><figure class="align-center">
|
| 62 |
-
<img alt="USleep Architecture" src="https://media.springernature.com/full/springer-static/image/art%3A10.1038%2Fs41746-021-00440-5/MediaObjects/41746_2021_440_Fig2_HTML.png" />
|
| 63 |
-
<figcaption>
|
| 64 |
-
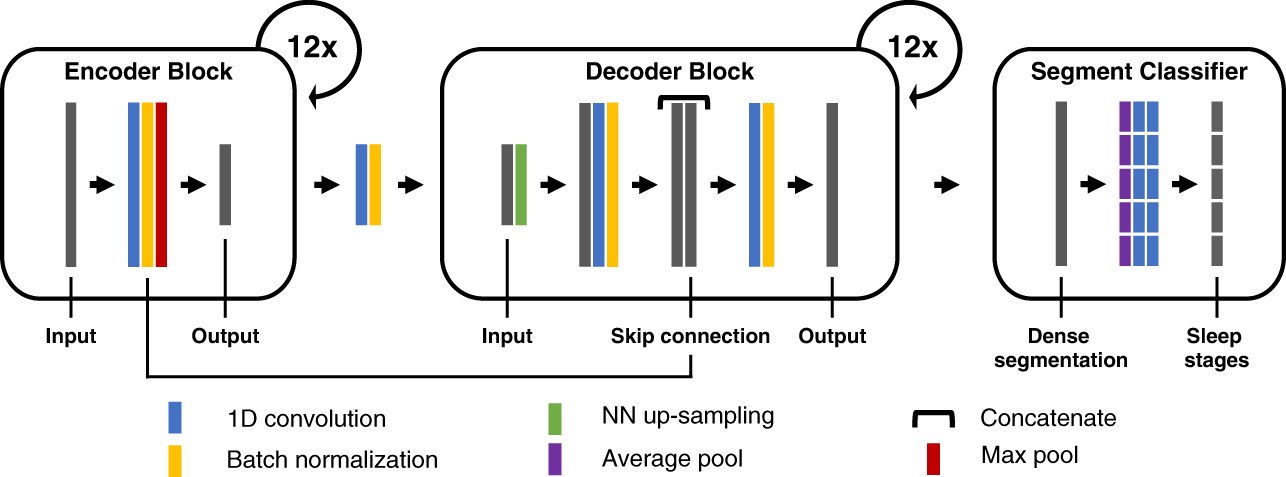
<p>Figure: U-Sleep consists of an encoder (left) which encodes the input signals into dense feature representations, a decoder (middle) which projects
|
| 65 |
-
the learned features into the input space to generate a dense sleep stage representation, and finally a specially designed segment
|
| 66 |
-
classifier (right) which generates sleep stages at a chosen temporal resolution.</p>
|
| 67 |
-
</figcaption>
|
| 68 |
-
</figure>
|
| 69 |
-
<p><strong>Architectural Overview</strong></p>
|
| 70 |
-
<p>U-Sleep is a <strong>fully convolutional</strong>, feed-forward encoder-decoder with a <em>segment classifier</em> head for
|
| 71 |
-
time-series <strong>segmentation</strong> (sleep staging). It maps multi-channel PSG (EEG+EOG) to a <em>dense, high-frequency</em>
|
| 72 |
-
per-sample representation, then aggregates it into fixed-length stage labels (e.g., 30 s). The network
|
| 73 |
-
processes arbitrarily long inputs in <strong>one forward pass</strong> (resampling to 128 Hz), allowing whole-night
|
| 74 |
-
hypnograms in seconds.</p>
|
| 75 |
-
<ul class="simple">
|
| 76 |
-
<li><p>(i). :class:`_EncoderBlock` extracts progressively deeper temporal features at lower resolution;</p></li>
|
| 77 |
-
<li><p>(ii). :class:`_Decoder` upsamples and fuses encoder features via U-Net-style skips to recover a per-sample stage map;</p></li>
|
| 78 |
-
<li><p>(iii). Segment Classifier mean-pools over the target epoch length and applies two pointwise convs to yield
|
| 79 |
-
per-epoch probabilities. Integrates into the USleep class.</p></li>
|
| 80 |
-
</ul>
|
| 81 |
-
<p><strong>Macro Components</strong></p>
|
| 82 |
-
<ul>
|
| 83 |
-
<li><p>Encoder :class:`_EncoderBlock` <strong>(multi-scale temporal feature extractor; downsampling x2 per block)</strong></p>
|
| 84 |
-
<blockquote>
|
| 85 |
-
<ul class="simple">
|
| 86 |
-
<li><p><em>Operations.</em></p></li>
|
| 87 |
-
<li><p><strong>Conv1d</strong> (:class:`torch.nn.Conv1d`) with kernel <span class="docutils literal">9</span> (stride <span class="docutils literal">1</span>, no dilation)</p></li>
|
| 88 |
-
<li><p><strong>ELU</strong> (:class:`torch.nn.ELU`)</p></li>
|
| 89 |
-
<li><p><strong>Batch Norm</strong> (:class:`torch.nn.BatchNorm1d`)</p></li>
|
| 90 |
-
<li><p><strong>Max Pool 1d</strong>, :class:`torch.nn.MaxPool1d` (<span class="docutils literal">kernel=2, stride=2</span>).</p></li>
|
| 91 |
-
</ul>
|
| 92 |
-
<p>Filters grow with depth by a factor of <span class="docutils literal">sqrt(2)</span> (start <span class="docutils literal">c_1=5</span>); each block exposes a <strong>skip</strong>
|
| 93 |
-
(pre-pooling activation) to the matching decoder block.
|
| 94 |
-
<em>Role.</em> Slow, uniform downsampling preserves early information while expanding the effective temporal
|
| 95 |
-
context over minutes—foundational for robust cross-cohort staging.</p>
|
| 96 |
-
</blockquote>
|
| 97 |
-
</li>
|
| 98 |
-
</ul>
|
| 99 |
-
<p>The number of filters grows with depth (capacity scaling); each block also exposes a <strong>skip</strong> (pre-pool)
|
| 100 |
-
to the matching decoder block.</p>
|
| 101 |
-
<dl class="simple">
|
| 102 |
-
<dt><strong>Rationale.</strong></dt>
|
| 103 |
-
<dd><ul class="simple">
|
| 104 |
-
<li><p>Slow, uniform downsampling (x2 each level) preserves information in early layers while expanding the temporal receptive field over the minutes.</p></li>
|
| 105 |
-
</ul>
|
| 106 |
-
</dd>
|
| 107 |
-
</dl>
|
| 108 |
-
<ul>
|
| 109 |
-
<li><p>Decoder :class:`_DecoderBlock` <strong>(progressive upsampling + skip fusion to high-frequency map, 12 blocks; upsampling x2 per block)</strong></p>
|
| 110 |
-
<blockquote>
|
| 111 |
-
<ul>
|
| 112 |
-
<li><p><em>Operations.</em></p>
|
| 113 |
-
<blockquote>
|
| 114 |
-
<ul class="simple">
|
| 115 |
-
<li><p><strong>Nearest-neighbor upsample</strong>, :class:`nn.Upsample` (x2)</p></li>
|
| 116 |
-
<li><p><strong>Convolution2d</strong> (k=2), :class:`torch.nn.Conv2d`</p></li>
|
| 117 |
-
<li><p>ELU, :class:`torch.nn.ELU`</p></li>
|
| 118 |
-
<li><p>Batch Norm, :class:`torch.nn.BatchNorm2d`</p></li>
|
| 119 |
-
<li><p><strong>Concatenate</strong> with the encoder skip at the same temporal scale, <span class="docutils literal">torch.cat</span></p></li>
|
| 120 |
-
<li><p><strong>Convolution</strong>, :class:`torch.nn.Conv2d`</p></li>
|
| 121 |
-
<li><p>ELU, :class:`torch.nn.ELU`</p></li>
|
| 122 |
-
<li><p>Batch Norm, :class:`torch.nn.BatchNorm2d`.</p></li>
|
| 123 |
-
</ul>
|
| 124 |
-
</blockquote>
|
| 125 |
-
</li>
|
| 126 |
-
</ul>
|
| 127 |
-
</blockquote>
|
| 128 |
-
</li>
|
| 129 |
-
</ul>
|
| 130 |
-
<p><strong>Output</strong>: A multi-class, <strong>high-frequency</strong> per-sample representation aligned to the input rate (128 Hz).</p>
|
| 131 |
-
<ul>
|
| 132 |
-
<li><p><strong>Segment Classifier incorporate into :class:`braindecode.models.USleep` (aggregation to fixed epochs)</strong></p>
|
| 133 |
-
<blockquote>
|
| 134 |
-
<ul>
|
| 135 |
-
<li><p><em>Operations.</em></p>
|
| 136 |
-
<blockquote>
|
| 137 |
-
<ul class="simple">
|
| 138 |
-
<li><p><strong>Mean-pool</strong>, :class:`torch.nn.AvgPool2d` per class with kernel = epoch length <em>i</em> and stride <em>i</em></p></li>
|
| 139 |
-
<li><p><strong>1x1 conv</strong>, :class:`torch.nn.Conv2d`</p></li>
|
| 140 |
-
<li><p>ELU, :class:`torch.nn.ELU`</p></li>
|
| 141 |
-
<li><p><strong>1x1 conv</strong>, :class:`torch.nn.Conv2d` with <span class="docutils literal">(T, K)</span> (epochs x stages).</p></li>
|
| 142 |
-
</ul>
|
| 143 |
-
</blockquote>
|
| 144 |
-
</li>
|
| 145 |
-
</ul>
|
| 146 |
-
</blockquote>
|
| 147 |
-
</li>
|
| 148 |
-
</ul>
|
| 149 |
-
<p><strong>Role</strong>: Learns a <strong>non-linear</strong> weighted combination over each 30-s window (unlike U-Time's linear combiner).</p>
|
| 150 |
-
<p><strong>Convolutional Details</strong></p>
|
| 151 |
-
<ul>
|
| 152 |
-
<li><p><strong>Temporal (where time-domain patterns are learned).</strong></p>
|
| 153 |
-
<p>All convolutions are <strong>1-D along time</strong>; depth (12 levels) plus pooling yields an extensive receptive field
|
| 154 |
-
(reported sensitivity to ±6.75 min around each epoch; theoretical field ≈ 9.6 min at the deepest layer).
|
| 155 |
-
The decoder restores sample-level resolution before epoch aggregation.</p>
|
| 156 |
-
</li>
|
| 157 |
-
<li><p><strong>Spatial (how channels are processed).</strong></p>
|
| 158 |
-
<p>Convolutions mix across the <em>channel</em> dimension jointly with time (no separate spatial operator). The system
|
| 159 |
-
is <strong>montage-agnostic</strong> (any reasonable EEG/EOG pair) and was trained across diverse cohorts/protocols,
|
| 160 |
-
supporting robustness to channel placement and hardware differences.</p>
|
| 161 |
-
</li>
|
| 162 |
-
<li><p><strong>Spectral (how frequency content is captured).</strong></p>
|
| 163 |
-
<p>No explicit Fourier/wavelet transform is used; the <strong>stack of temporal convolutions</strong> acts as a learned
|
| 164 |
-
filter bank whose effective bandwidth grows with depth. The high-frequency decoder output (128 Hz)
|
| 165 |
-
retains fine temporal detail for the segment classifier.</p>
|
| 166 |
-
</li>
|
| 167 |
-
</ul>
|
| 168 |
-
<p><strong>Attention / Sequential Modules</strong></p>
|
| 169 |
-
<p>U-Sleep contains <strong>no attention or recurrent units</strong>; it is a <em>pure</em> feed-forward, fully convolutional
|
| 170 |
-
segmentation network inspired by U-Net/U-Time, favoring training stability and cross-dataset portability.</p>
|
| 171 |
-
<p><strong>Additional Mechanisms</strong></p>
|
| 172 |
-
<ul class="simple">
|
| 173 |
-
<li><p><strong>U-Net lineage with task-specific head.</strong> U-Sleep extends U-Time by being <strong>deeper</strong> (12 vs. 4 levels),
|
| 174 |
-
switching ReLU→**ELU**, using uniform pooling (2) at all depths, and replacing the linear combiner with a
|
| 175 |
-
<strong>two-layer</strong> pointwise head—improving capacity and resilience across datasets.</p></li>
|
| 176 |
-
<li><p><strong>Arbitrary-length inference.</strong> Thanks to full convolutionality and tiling-free design, entire nights can be
|
| 177 |
-
staged in a single pass on commodity hardware. Inputs shorter than ≈ 17.5 min may reduce performance by
|
| 178 |
-
limiting long-range context.</p></li>
|
| 179 |
-
<li><p><strong>Complexity scaling (alpha).</strong> Filter counts can be adjusted by a global <strong>complexity factor</strong> to trade accuracy
|
| 180 |
-
and memory (as described in the paper's topology table).</p></li>
|
| 181 |
-
</ul>
|
| 182 |
-
<p><strong>Usage and Configuration</strong></p>
|
| 183 |
-
<ul class="simple">
|
| 184 |
-
<li><p><strong>Practice.</strong> Resample PSG to <strong>128 Hz</strong> and provide at least two channels (one EEG, one EOG). Choose epoch
|
| 185 |
-
length <em>i</em> (often 30 s); ensure windows long enough to exploit the model's receptive field (e.g., training on
|
| 186 |
-
≥ 17.5 min chunks).</p></li>
|
| 187 |
-
</ul>
|
| 188 |
-
<section id="parameters">
|
| 189 |
-
<h2>Parameters</h2>
|
| 190 |
-
<dl class="simple">
|
| 191 |
-
<dt>n_chans<span class="classifier">int</span></dt>
|
| 192 |
-
<dd><p>Number of EEG or EOG channels. Set to 2 in <a class="brackets" href="#footnote-1" id="footnote-reference-2" role="doc-noteref"><span class="fn-bracket">[</span>1<span class="fn-bracket">]</span></a> (1 EEG, 1 EOG).</p>
|
| 193 |
-
</dd>
|
| 194 |
-
<dt>sfreq<span class="classifier">float</span></dt>
|
| 195 |
-
<dd><p>EEG sampling frequency. Set to 128 in <a class="brackets" href="#footnote-1" id="footnote-reference-3" role="doc-noteref"><span class="fn-bracket">[</span>1<span class="fn-bracket">]</span></a>.</p>
|
| 196 |
-
</dd>
|
| 197 |
-
<dt>depth<span class="classifier">int</span></dt>
|
| 198 |
-
<dd><p>Number of conv blocks in encoding layer (number of 2x2 max pools).
|
| 199 |
-
Note: each block halves the spatial dimensions of the features.</p>
|
| 200 |
-
</dd>
|
| 201 |
-
<dt>n_time_filters<span class="classifier">int</span></dt>
|
| 202 |
-
<dd><p>Initial number of convolutional filters. Set to 5 in <a class="brackets" href="#footnote-1" id="footnote-reference-4" role="doc-noteref"><span class="fn-bracket">[</span>1<span class="fn-bracket">]</span></a>.</p>
|
| 203 |
-
</dd>
|
| 204 |
-
<dt>complexity_factor<span class="classifier">float</span></dt>
|
| 205 |
-
<dd><p>Multiplicative factor for the number of channels at each layer of the U-Net.
|
| 206 |
-
Set to 2 in <a class="brackets" href="#footnote-1" id="footnote-reference-5" role="doc-noteref"><span class="fn-bracket">[</span>1<span class="fn-bracket">]</span></a>.</p>
|
| 207 |
-
</dd>
|
| 208 |
-
<dt>with_skip_connection<span class="classifier">bool</span></dt>
|
| 209 |
-
<dd><p>If True, use skip connections in decoder blocks.</p>
|
| 210 |
-
</dd>
|
| 211 |
-
<dt>n_outputs<span class="classifier">int</span></dt>
|
| 212 |
-
<dd><p>Number of outputs/classes. Set to 5.</p>
|
| 213 |
-
</dd>
|
| 214 |
-
<dt>input_window_seconds<span class="classifier">float</span></dt>
|
| 215 |
-
<dd><p>Size of the input, in seconds. Set to 30 in <a class="brackets" href="#footnote-1" id="footnote-reference-6" role="doc-noteref"><span class="fn-bracket">[</span>1<span class="fn-bracket">]</span></a>.</p>
|
| 216 |
-
</dd>
|
| 217 |
-
<dt>time_conv_size_s<span class="classifier">float</span></dt>
|
| 218 |
-
<dd><p>Size of the temporal convolution kernel, in seconds. Set to 9 / 128 in
|
| 219 |
-
<a class="brackets" href="#footnote-1" id="footnote-reference-7" role="doc-noteref"><span class="fn-bracket">[</span>1<span class="fn-bracket">]</span></a>.</p>
|
| 220 |
-
</dd>
|
| 221 |
-
<dt>ensure_odd_conv_size<span class="classifier">bool</span></dt>
|
| 222 |
-
<dd><p>If True and the size of the convolutional kernel is an even number, one
|
| 223 |
-
will be added to it to ensure it is odd, so that the decoder blocks can
|
| 224 |
-
work. This can be useful when using different sampling rates from 128
|
| 225 |
-
or 100 Hz.</p>
|
| 226 |
-
</dd>
|
| 227 |
-
<dt>activation<span class="classifier">nn.Module, default=nn.ELU</span></dt>
|
| 228 |
-
<dd><p>Activation function class to apply. Should be a PyTorch activation
|
| 229 |
-
module class like <span class="docutils literal">nn.ReLU</span> or <span class="docutils literal">nn.ELU</span>. Default is <span class="docutils literal">nn.ELU</span>.</p>
|
| 230 |
-
</dd>
|
| 231 |
-
</dl>
|
| 232 |
-
</section>
|
| 233 |
-
<section id="references">
|
| 234 |
-
<h2>References</h2>
|
| 235 |
-
<aside class="footnote-list brackets">
|
| 236 |
-
<aside class="footnote brackets" id="footnote-1" role="doc-footnote">
|
| 237 |
-
<span class="label"><span class="fn-bracket">[</span>1<span class="fn-bracket">]</span></span>
|
| 238 |
-
<span class="backrefs">(<a role="doc-backlink" href="#footnote-reference-1">1</a>,<a role="doc-backlink" href="#footnote-reference-2">2</a>,<a role="doc-backlink" href="#footnote-reference-3">3</a>,<a role="doc-backlink" href="#footnote-reference-4">4</a>,<a role="doc-backlink" href="#footnote-reference-5">5</a>,<a role="doc-backlink" href="#footnote-reference-6">6</a>,<a role="doc-backlink" href="#footnote-reference-7">7</a>)</span>
|
| 239 |
-
<p>Perslev M, Darkner S, Kempfner L, Nikolic M, Jennum PJ, Igel C.
|
| 240 |
-
U-Sleep: resilient high-frequency sleep staging. <em>npj Digit. Med.</em> 4, 72 (2021).
|
| 241 |
-
<a class="reference external" href="https://github.com/perslev/U-Time/blob/master/utime/models/usleep.py">https://github.com/perslev/U-Time/blob/master/utime/models/usleep.py</a></p>
|
| 242 |
-
</aside>
|
| 243 |
-
</aside>
|
| 244 |
-
<p><strong>Hugging Face Hub integration</strong></p>
|
| 245 |
-
<p>When the optional <span class="docutils literal">huggingface_hub</span> package is installed, all models
|
| 246 |
-
automatically gain the ability to be pushed to and loaded from the
|
| 247 |
-
Hugging Face Hub. Install with:</p>
|
| 248 |
-
<pre class="literal-block">pip install braindecode[hub]</pre>
|
| 249 |
-
<p><strong>Pushing a model to the Hub:</strong></p>
|
| 250 |
-
<p><strong>Loading a model from the Hub:</strong></p>
|
| 251 |
-
<p><strong>Extracting features and replacing the head:</strong></p>
|
| 252 |
-
<p><strong>Saving and restoring full configuration:</strong></p>
|
| 253 |
-
<p>All model parameters (both EEG-specific and model-specific such as
|
| 254 |
-
dropout rates, activation functions, number of filters) are automatically
|
| 255 |
-
saved to the Hub and restored when loading.</p>
|
| 256 |
-
<p>See :ref:`load-pretrained-models` for a complete tutorial.</p>
|
| 257 |
-
</section>
|
| 258 |
-
</main>
|
| 259 |
-
</div>
|
| 260 |
|
| 261 |
## Citation
|
| 262 |
|
| 263 |
-
|
| 264 |
-
*References* section above) and braindecode:
|
| 265 |
|
| 266 |
```bibtex
|
| 267 |
@article{aristimunha2025braindecode,
|
|
|
|
| 9 |
- neuroscience
|
| 10 |
- braindecode
|
| 11 |
- convolutional
|
|
|
|
| 12 |
- sleep-staging
|
| 13 |
---
|
| 14 |
|
| 15 |
# USleep
|
| 16 |
|
| 17 |
+
Sleep staging architecture from Perslev et al (2021) [1].
|
| 18 |
|
| 19 |
+
> **Architecture-only repository.** Documents the
|
| 20 |
> `braindecode.models.USleep` class. **No pretrained weights are
|
| 21 |
+
> distributed here.** Instantiate the model and train it on your own
|
| 22 |
+
> data.
|
|
|
|
| 23 |
|
| 24 |
## Quick start
|
| 25 |
|
|
|
|
| 38 |
)
|
| 39 |
```
|
| 40 |
|
| 41 |
+
The signal-shape arguments above are illustrative defaults — adjust to
|
| 42 |
+
match your recording.
|
| 43 |
|
| 44 |
## Documentation
|
| 45 |
+
- Full API reference: <https://braindecode.org/stable/generated/braindecode.models.USleep.html>
|
| 46 |
+
- Interactive browser (live instantiation, parameter counts):
|
|
|
|
|
|
|
| 47 |
<https://huggingface.co/spaces/braindecode/model-explorer>
|
| 48 |
- Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/usleep.py#L14>
|
| 49 |
|
|
|
|
| 50 |
|
| 51 |
+
## Architecture
|
| 52 |
+
|
| 53 |
+
|
| 54 |
+
|
| 55 |
+
|
| 56 |
+
## Parameters
|
| 57 |
+
|
| 58 |
+
| Parameter | Type | Description |
|
| 59 |
+
|---|---|---|
|
| 60 |
+
| `n_chans` | int | Number of EEG or EOG channels. Set to 2 in [1] (1 EEG, 1 EOG). |
|
| 61 |
+
| `sfreq` | float | EEG sampling frequency. Set to 128 in [1]. |
|
| 62 |
+
| `depth` | int | Number of conv blocks in encoding layer (number of 2x2 max pools). Note: each block halves the spatial dimensions of the features. |
|
| 63 |
+
| `n_time_filters` | int | Initial number of convolutional filters. Set to 5 in [1]. |
|
| 64 |
+
| `complexity_factor` | float | Multiplicative factor for the number of channels at each layer of the U-Net. Set to 2 in [1]. |
|
| 65 |
+
| `with_skip_connection` | bool | If True, use skip connections in decoder blocks. |
|
| 66 |
+
| `n_outputs` | int | Number of outputs/classes. Set to 5. |
|
| 67 |
+
| `input_window_seconds` | float | Size of the input, in seconds. Set to 30 in [1]. |
|
| 68 |
+
| `time_conv_size_s` | float | Size of the temporal convolution kernel, in seconds. Set to 9 / 128 in [1]. |
|
| 69 |
+
| `ensure_odd_conv_size` | bool | If True and the size of the convolutional kernel is an even number, one will be added to it to ensure it is odd, so that the decoder blocks can work. This can be useful when using different sampling rates from 128 or 100 Hz. |
|
| 70 |
+
| `activation` | nn.Module, default=nn.ELU | Activation function class to apply. Should be a PyTorch activation module class like `nn.ReLU` or `nn.ELU`. Default is `nn.ELU`. |
|
| 71 |
+
|
| 72 |
+
|
| 73 |
+
## References
|
| 74 |
+
|
| 75 |
+
1. Perslev M, Darkner S, Kempfner L, Nikolic M, Jennum PJ, Igel C. U-Sleep: resilient high-frequency sleep staging. *npj Digit. Med.* 4, 72 (2021). https://github.com/perslev/U-Time/blob/master/utime/models/usleep.py
|
| 76 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 77 |
|
| 78 |
## Citation
|
| 79 |
|
| 80 |
+
Cite the original architecture paper (see *References* above) and braindecode:
|
|
|
|
| 81 |
|
| 82 |
```bibtex
|
| 83 |
@article{aristimunha2025braindecode,
|