Replace with clean markdown card
Browse files
README.md
CHANGED
|
@@ -10,18 +10,16 @@ tags:
|
|
| 10 |
- braindecode
|
| 11 |
- foundation-model
|
| 12 |
- convolutional
|
| 13 |
-
- transformer
|
| 14 |
---
|
| 15 |
|
| 16 |
# MEDFormer
|
| 17 |
|
| 18 |
-
Medformer from Wang et al (2024) .
|
| 19 |
|
| 20 |
-
> **Architecture-only repository.**
|
| 21 |
> `braindecode.models.MEDFormer` class. **No pretrained weights are
|
| 22 |
-
> distributed here**
|
| 23 |
-
> data
|
| 24 |
-
> separately.
|
| 25 |
|
| 26 |
## Quick start
|
| 27 |
|
|
@@ -40,841 +38,46 @@ model = MEDFormer(
|
|
| 40 |
)
|
| 41 |
```
|
| 42 |
|
| 43 |
-
The signal-shape arguments above are
|
| 44 |
-
|
| 45 |
|
| 46 |
## Documentation
|
| 47 |
-
|
| 48 |
-
-
|
| 49 |
-
<https://braindecode.org/stable/generated/braindecode.models.MEDFormer.html>
|
| 50 |
-
- Interactive browser with live instantiation:
|
| 51 |
<https://huggingface.co/spaces/braindecode/model-explorer>
|
| 52 |
- Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/medformer.py#L20>
|
| 53 |
|
| 54 |
-
## Architecture description
|
| 55 |
|
| 56 |
-
|
| 57 |
-
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 58 |
|
| 59 |
-
<div class='bd-doc'><main>
|
| 60 |
-
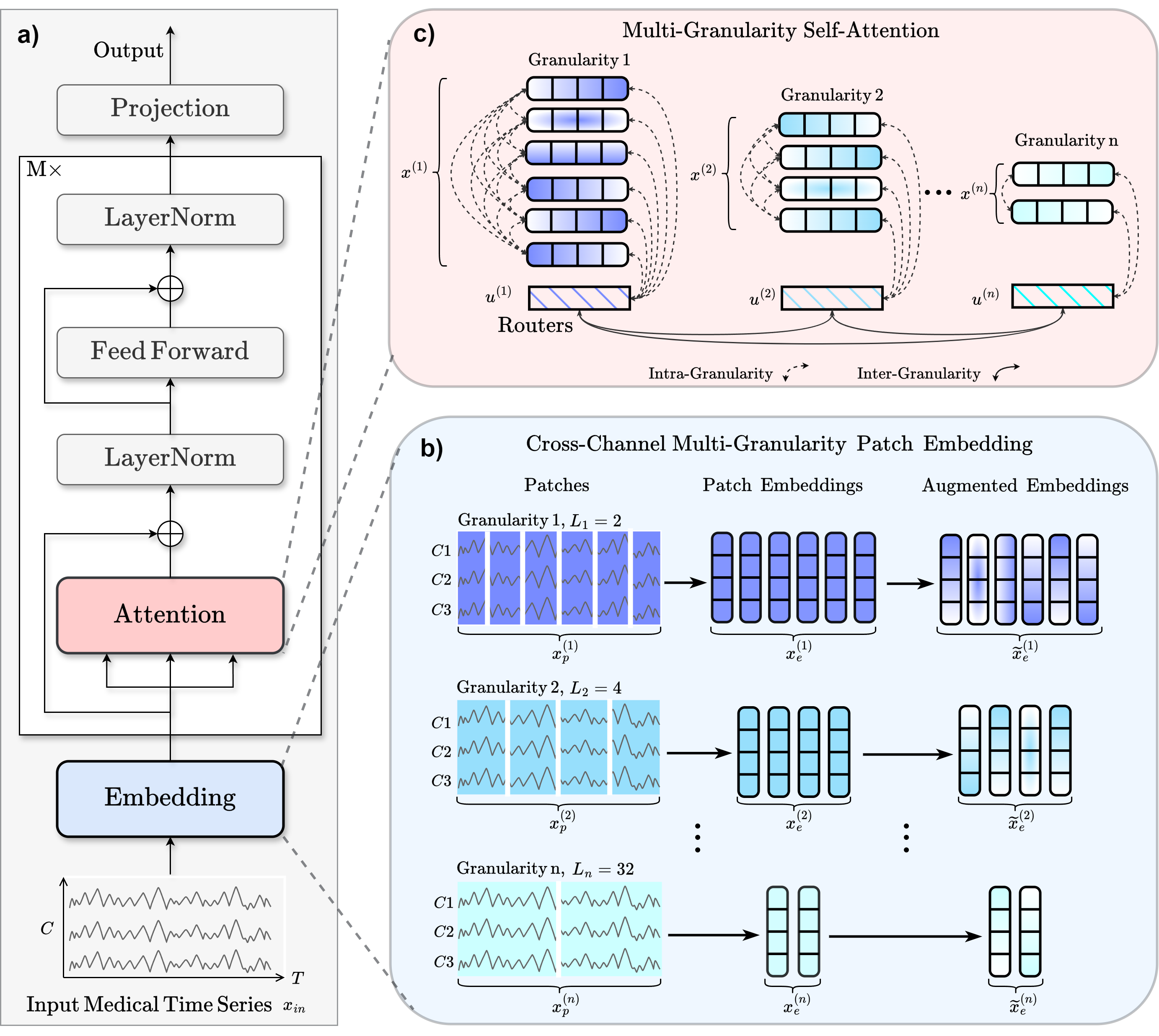
<p>Medformer from Wang et al (2024) <a class="citation-reference" href="#medformer2024" id="citation-reference-1" role="doc-biblioref">[Medformer2024]</a>.</p>
|
| 61 |
-
<span style="display:inline-block;padding:2px 8px;border-radius:4px;background:#5cb85c;color:white;font-size:11px;font-weight:600;margin-right:4px;">Convolution</span><span style="display:inline-block;padding:2px 8px;border-radius:4px;background:#d9534f;color:white;font-size:11px;font-weight:600;margin-right:4px;">Foundation Model</span><figure class="align-center">
|
| 62 |
-
<img alt="MEDFormer Architecture." src="https://raw.githubusercontent.com/DL4mHealth/Medformer/refs/heads/main/figs/medformer_architecture.png" />
|
| 63 |
-
<figcaption>
|
| 64 |
-
<p>a) Workflow. b) For the input sample <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 65 |
-
<msub>
|
| 66 |
-
<mi>x</mi>
|
| 67 |
-
<mtext>in</mtext>
|
| 68 |
-
</msub>
|
| 69 |
-
</math>, the authors apply <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 70 |
-
<mi>n</mi>
|
| 71 |
-
</math>
|
| 72 |
-
different patch lengths in parallel to create patched features <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 73 |
-
<msubsup>
|
| 74 |
-
<mi>x</mi>
|
| 75 |
-
<mi>p</mi>
|
| 76 |
-
<mrow>
|
| 77 |
-
<mo stretchy="false">(</mo>
|
| 78 |
-
<mi>i</mi>
|
| 79 |
-
<mo stretchy="false">)</mo>
|
| 80 |
-
</mrow>
|
| 81 |
-
</msubsup>
|
| 82 |
-
</math>, where <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 83 |
-
<mi>i</mi>
|
| 84 |
-
</math>
|
| 85 |
-
ranges from 1 to <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 86 |
-
<mi>n</mi>
|
| 87 |
-
</math>. Each patch length represents a different granularity. These patched
|
| 88 |
-
features are linearly transformed into <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 89 |
-
<msubsup>
|
| 90 |
-
<mi>x</mi>
|
| 91 |
-
<mi>e</mi>
|
| 92 |
-
<mrow>
|
| 93 |
-
<mo stretchy="false">(</mo>
|
| 94 |
-
<mi>i</mi>
|
| 95 |
-
<mo stretchy="false">)</mo>
|
| 96 |
-
</mrow>
|
| 97 |
-
</msubsup>
|
| 98 |
-
</math> and augmented into <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 99 |
-
<msup>
|
| 100 |
-
<munderover>
|
| 101 |
-
<mi>x</mi>
|
| 102 |
-
<mi>e</mi>
|
| 103 |
-
<mo accent="true">~</mo>
|
| 104 |
-
</munderover>
|
| 105 |
-
<mrow>
|
| 106 |
-
<mo stretchy="false">(</mo>
|
| 107 |
-
<mi>i</mi>
|
| 108 |
-
<mo stretchy="false">)</mo>
|
| 109 |
-
</mrow>
|
| 110 |
-
</msup>
|
| 111 |
-
</math>.
|
| 112 |
-
c) The final patch embedding <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 113 |
-
<msup>
|
| 114 |
-
<mi>x</mi>
|
| 115 |
-
<mrow>
|
| 116 |
-
<mo stretchy="false">(</mo>
|
| 117 |
-
<mi>i</mi>
|
| 118 |
-
<mo stretchy="false">)</mo>
|
| 119 |
-
</mrow>
|
| 120 |
-
</msup>
|
| 121 |
-
</math> fuses augmented <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 122 |
-
<msup>
|
| 123 |
-
<munderover>
|
| 124 |
-
<mi>x</mi>
|
| 125 |
-
<mi>e</mi>
|
| 126 |
-
<mo accent="true">~</mo>
|
| 127 |
-
</munderover>
|
| 128 |
-
<mrow>
|
| 129 |
-
<mo stretchy="false">(</mo>
|
| 130 |
-
<mi>i</mi>
|
| 131 |
-
<mo stretchy="false">)</mo>
|
| 132 |
-
</mrow>
|
| 133 |
-
</msup>
|
| 134 |
-
</math> with the
|
| 135 |
-
positional embedding <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 136 |
-
<msub>
|
| 137 |
-
<mi>W</mi>
|
| 138 |
-
<mtext>pos</mtext>
|
| 139 |
-
</msub>
|
| 140 |
-
</math> and the granularity embedding <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 141 |
-
<msubsup>
|
| 142 |
-
<mi>W</mi>
|
| 143 |
-
<mtext>gr</mtext>
|
| 144 |
-
<mrow>
|
| 145 |
-
<mo stretchy="false">(</mo>
|
| 146 |
-
<mi>i</mi>
|
| 147 |
-
<mo stretchy="false">)</mo>
|
| 148 |
-
</mrow>
|
| 149 |
-
</msubsup>
|
| 150 |
-
</math>.
|
| 151 |
-
Each granularity employs a router <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 152 |
-
<msup>
|
| 153 |
-
<mi>u</mi>
|
| 154 |
-
<mrow>
|
| 155 |
-
<mo stretchy="false">(</mo>
|
| 156 |
-
<mi>i</mi>
|
| 157 |
-
<mo stretchy="false">)</mo>
|
| 158 |
-
</mrow>
|
| 159 |
-
</msup>
|
| 160 |
-
</math> to capture aggregated information.
|
| 161 |
-
Intra-granularity attention focuses within individual granularities, and inter-granularity attention
|
| 162 |
-
leverages the routers to integrate information across granularities.</p>
|
| 163 |
-
</figcaption>
|
| 164 |
-
</figure>
|
| 165 |
-
<p>The <strong>MedFormer</strong> is a multi-granularity patching transformer tailored to medical
|
| 166 |
-
time-series (MedTS) classification, with an emphasis on EEG and ECG signals. It captures
|
| 167 |
-
local temporal dynamics, inter-channel correlations, and multi-scale temporal structure
|
| 168 |
-
through cross-channel patching, multi-granularity embeddings, and two-stage attention
|
| 169 |
-
<a class="citation-reference" href="#medformer2024" id="citation-reference-2" role="doc-biblioref">[Medformer2024]</a>.</p>
|
| 170 |
-
<p><strong>Architecture Overview</strong></p>
|
| 171 |
-
<p>MedFormer integrates three mechanisms to enhance representation learning <a class="citation-reference" href="#medformer2024" id="citation-reference-3" role="doc-biblioref">[Medformer2024]</a>:</p>
|
| 172 |
-
<ol class="arabic simple">
|
| 173 |
-
<li><p><strong>Cross-channel patching.</strong> Leverages inter-channel correlations by forming patches
|
| 174 |
-
across multiple channels and timestamps, capturing multi-timestamp and cross-channel
|
| 175 |
-
patterns.</p></li>
|
| 176 |
-
<li><p><strong>Multi-granularity embedding.</strong> Extracts features at different temporal scales from
|
| 177 |
-
:attr:`patch_len_list`, emulating frequency-band behavior without hand-crafted filters.</p></li>
|
| 178 |
-
<li><p><strong>Two-stage multi-granularity self-attention.</strong> Learns intra- and inter-granularity
|
| 179 |
-
correlations to fuse information across temporal scales.</p></li>
|
| 180 |
-
</ol>
|
| 181 |
-
<p><strong>Macro Components</strong></p>
|
| 182 |
-
<dl>
|
| 183 |
-
<dt><span class="docutils literal">MEDFormer.enc_embedding</span> (Embedding Layer)</dt>
|
| 184 |
-
<dd><p><strong>Operations.</strong> :class:`~braindecode.models.medformer._ListPatchEmbedding` implements
|
| 185 |
-
cross-channel multi-granularity patching. For each patch length <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 186 |
-
<msub>
|
| 187 |
-
<mi>L</mi>
|
| 188 |
-
<mi>i</mi>
|
| 189 |
-
</msub>
|
| 190 |
-
</math>, the input
|
| 191 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 192 |
-
<msub>
|
| 193 |
-
<mi>𝐱</mi>
|
| 194 |
-
<mtext>in</mtext>
|
| 195 |
-
</msub>
|
| 196 |
-
<mo>∈</mo>
|
| 197 |
-
<msup>
|
| 198 |
-
<mi>ℝ</mi>
|
| 199 |
-
<mrow>
|
| 200 |
-
<mi>T</mi>
|
| 201 |
-
<mo>×</mo>
|
| 202 |
-
<mi>C</mi>
|
| 203 |
-
</mrow>
|
| 204 |
-
</msup>
|
| 205 |
-
</math> is segmented into
|
| 206 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 207 |
-
<msub>
|
| 208 |
-
<mi>N</mi>
|
| 209 |
-
<mi>i</mi>
|
| 210 |
-
</msub>
|
| 211 |
-
</math> cross-channel non-overlapping patches
|
| 212 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 213 |
-
<msubsup>
|
| 214 |
-
<mi>𝐱</mi>
|
| 215 |
-
<mi>p</mi>
|
| 216 |
-
<mrow>
|
| 217 |
-
<mo stretchy="false">(</mo>
|
| 218 |
-
<mi>i</mi>
|
| 219 |
-
<mo stretchy="false">)</mo>
|
| 220 |
-
</mrow>
|
| 221 |
-
</msubsup>
|
| 222 |
-
<mo>∈</mo>
|
| 223 |
-
<msup>
|
| 224 |
-
<mi>ℝ</mi>
|
| 225 |
-
<mrow>
|
| 226 |
-
<msub>
|
| 227 |
-
<mi>N</mi>
|
| 228 |
-
<mi>i</mi>
|
| 229 |
-
</msub>
|
| 230 |
-
<mo>×</mo>
|
| 231 |
-
<mo stretchy="false">(</mo>
|
| 232 |
-
<msub>
|
| 233 |
-
<mi>L</mi>
|
| 234 |
-
<mi>i</mi>
|
| 235 |
-
</msub>
|
| 236 |
-
<mo>⋅</mo>
|
| 237 |
-
<mi>C</mi>
|
| 238 |
-
<mo stretchy="false">)</mo>
|
| 239 |
-
</mrow>
|
| 240 |
-
</msup>
|
| 241 |
-
</math>, where
|
| 242 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 243 |
-
<msub>
|
| 244 |
-
<mi>N</mi>
|
| 245 |
-
<mi>i</mi>
|
| 246 |
-
</msub>
|
| 247 |
-
<mo>=</mo>
|
| 248 |
-
<mo>⌈</mo>
|
| 249 |
-
<mi>T</mi>
|
| 250 |
-
<mo stretchy="false">/</mo>
|
| 251 |
-
<msub>
|
| 252 |
-
<mi>L</mi>
|
| 253 |
-
<mi>i</mi>
|
| 254 |
-
</msub>
|
| 255 |
-
<mo>⌉</mo>
|
| 256 |
-
</math>. Each patch is linearly projected via
|
| 257 |
-
:class:`~braindecode.models.medformer._CrossChannelTokenEmbedding` to obtain
|
| 258 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 259 |
-
<msubsup>
|
| 260 |
-
<mi>𝐱</mi>
|
| 261 |
-
<mi>e</mi>
|
| 262 |
-
<mrow>
|
| 263 |
-
<mo stretchy="false">(</mo>
|
| 264 |
-
<mi>i</mi>
|
| 265 |
-
<mo stretchy="false">)</mo>
|
| 266 |
-
</mrow>
|
| 267 |
-
</msubsup>
|
| 268 |
-
<mo>∈</mo>
|
| 269 |
-
<msup>
|
| 270 |
-
<mi>ℝ</mi>
|
| 271 |
-
<mrow>
|
| 272 |
-
<msub>
|
| 273 |
-
<mi>N</mi>
|
| 274 |
-
<mi>i</mi>
|
| 275 |
-
</msub>
|
| 276 |
-
<mo>×</mo>
|
| 277 |
-
<mi>D</mi>
|
| 278 |
-
</mrow>
|
| 279 |
-
</msup>
|
| 280 |
-
</math>. Data augmentations
|
| 281 |
-
(masking, jittering) produce augmented embeddings <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 282 |
-
<msup>
|
| 283 |
-
<munderover>
|
| 284 |
-
<mi>𝐱</mi>
|
| 285 |
-
<mi>e</mi>
|
| 286 |
-
<mo stretchy="false">~</mo>
|
| 287 |
-
</munderover>
|
| 288 |
-
<mrow>
|
| 289 |
-
<mo stretchy="false">(</mo>
|
| 290 |
-
<mi>i</mi>
|
| 291 |
-
<mo stretchy="false">)</mo>
|
| 292 |
-
</mrow>
|
| 293 |
-
</msup>
|
| 294 |
-
</math>.
|
| 295 |
-
The final embedding combines augmented patches, fixed positional embeddings
|
| 296 |
-
(:class:`~braindecode.models.medformer._PositionalEmbedding`), and learnable
|
| 297 |
-
granularity embeddings <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 298 |
-
<msubsup>
|
| 299 |
-
<mi>𝐖</mi>
|
| 300 |
-
<mtext>gr</mtext>
|
| 301 |
-
<mrow>
|
| 302 |
-
<mo stretchy="false">(</mo>
|
| 303 |
-
<mi>i</mi>
|
| 304 |
-
<mo stretchy="false">)</mo>
|
| 305 |
-
</mrow>
|
| 306 |
-
</msubsup>
|
| 307 |
-
</math>:</p>
|
| 308 |
-
<div>
|
| 309 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML" display="block">
|
| 310 |
-
<msup>
|
| 311 |
-
<mi>𝐱</mi>
|
| 312 |
-
<mrow>
|
| 313 |
-
<mo stretchy="false">(</mo>
|
| 314 |
-
<mi>i</mi>
|
| 315 |
-
<mo stretchy="false">)</mo>
|
| 316 |
-
</mrow>
|
| 317 |
-
</msup>
|
| 318 |
-
<mo>=</mo>
|
| 319 |
-
<msup>
|
| 320 |
-
<munderover>
|
| 321 |
-
<mi>𝐱</mi>
|
| 322 |
-
<mi>e</mi>
|
| 323 |
-
<mo stretchy="false">~</mo>
|
| 324 |
-
</munderover>
|
| 325 |
-
<mrow>
|
| 326 |
-
<mo stretchy="false">(</mo>
|
| 327 |
-
<mi>i</mi>
|
| 328 |
-
<mo stretchy="false">)</mo>
|
| 329 |
-
</mrow>
|
| 330 |
-
</msup>
|
| 331 |
-
<mo>+</mo>
|
| 332 |
-
<msub>
|
| 333 |
-
<mi>𝐖</mi>
|
| 334 |
-
<mtext>pos</mtext>
|
| 335 |
-
</msub>
|
| 336 |
-
<mo stretchy="false">[</mo>
|
| 337 |
-
<mn>1</mn>
|
| 338 |
-
<mo>∶</mo>
|
| 339 |
-
<msub>
|
| 340 |
-
<mi>N</mi>
|
| 341 |
-
<mi>i</mi>
|
| 342 |
-
</msub>
|
| 343 |
-
<mo stretchy="false">]</mo>
|
| 344 |
-
<mo>+</mo>
|
| 345 |
-
<msubsup>
|
| 346 |
-
<mi>𝐖</mi>
|
| 347 |
-
<mtext>gr</mtext>
|
| 348 |
-
<mrow>
|
| 349 |
-
<mo stretchy="false">(</mo>
|
| 350 |
-
<mi>i</mi>
|
| 351 |
-
<mo stretchy="false">)</mo>
|
| 352 |
-
</mrow>
|
| 353 |
-
</msubsup>
|
| 354 |
-
</math>
|
| 355 |
-
</div>
|
| 356 |
-
<p>Additionally, a router token is initialized for each granularity:</p>
|
| 357 |
-
<div>
|
| 358 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML" display="block">
|
| 359 |
-
<msup>
|
| 360 |
-
<mi>𝐮</mi>
|
| 361 |
-
<mrow>
|
| 362 |
-
<mo stretchy="false">(</mo>
|
| 363 |
-
<mi>i</mi>
|
| 364 |
-
<mo stretchy="false">)</mo>
|
| 365 |
-
</mrow>
|
| 366 |
-
</msup>
|
| 367 |
-
<mo>=</mo>
|
| 368 |
-
<msub>
|
| 369 |
-
<mi>𝐖</mi>
|
| 370 |
-
<mtext>pos</mtext>
|
| 371 |
-
</msub>
|
| 372 |
-
<mo stretchy="false">[</mo>
|
| 373 |
-
<msub>
|
| 374 |
-
<mi>N</mi>
|
| 375 |
-
<mi>i</mi>
|
| 376 |
-
</msub>
|
| 377 |
-
<mo>+</mo>
|
| 378 |
-
<mn>1</mn>
|
| 379 |
-
<mo stretchy="false">]</mo>
|
| 380 |
-
<mo>+</mo>
|
| 381 |
-
<msubsup>
|
| 382 |
-
<mi>𝐖</mi>
|
| 383 |
-
<mtext>gr</mtext>
|
| 384 |
-
<mrow>
|
| 385 |
-
<mo stretchy="false">(</mo>
|
| 386 |
-
<mi>i</mi>
|
| 387 |
-
<mo stretchy="false">)</mo>
|
| 388 |
-
</mrow>
|
| 389 |
-
</msubsup>
|
| 390 |
-
</math>
|
| 391 |
-
</div>
|
| 392 |
-
<p><strong>Role.</strong> Converts raw input into granularity-specific patch embeddings
|
| 393 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 394 |
-
<mo>{</mo>
|
| 395 |
-
<msup>
|
| 396 |
-
<mi>𝐱</mi>
|
| 397 |
-
<mrow>
|
| 398 |
-
<mo stretchy="false">(</mo>
|
| 399 |
-
<mn>1</mn>
|
| 400 |
-
<mo stretchy="false">)</mo>
|
| 401 |
-
</mrow>
|
| 402 |
-
</msup>
|
| 403 |
-
<mo>,</mo>
|
| 404 |
-
<mi>…</mi>
|
| 405 |
-
<mo>,</mo>
|
| 406 |
-
<msup>
|
| 407 |
-
<mi>𝐱</mi>
|
| 408 |
-
<mrow>
|
| 409 |
-
<mo stretchy="false">(</mo>
|
| 410 |
-
<mi>n</mi>
|
| 411 |
-
<mo stretchy="false">)</mo>
|
| 412 |
-
</mrow>
|
| 413 |
-
</msup>
|
| 414 |
-
<mo>}</mo>
|
| 415 |
-
</math> and router embeddings
|
| 416 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 417 |
-
<mo>{</mo>
|
| 418 |
-
<msup>
|
| 419 |
-
<mi>𝐮</mi>
|
| 420 |
-
<mrow>
|
| 421 |
-
<mo stretchy="false">(</mo>
|
| 422 |
-
<mn>1</mn>
|
| 423 |
-
<mo stretchy="false">)</mo>
|
| 424 |
-
</mrow>
|
| 425 |
-
</msup>
|
| 426 |
-
<mo>,</mo>
|
| 427 |
-
<mi>…</mi>
|
| 428 |
-
<mo>,</mo>
|
| 429 |
-
<msup>
|
| 430 |
-
<mi>𝐮</mi>
|
| 431 |
-
<mrow>
|
| 432 |
-
<mo stretchy="false">(</mo>
|
| 433 |
-
<mi>n</mi>
|
| 434 |
-
<mo stretchy="false">)</mo>
|
| 435 |
-
</mrow>
|
| 436 |
-
</msup>
|
| 437 |
-
<mo>}</mo>
|
| 438 |
-
</math> for multi-scale processing.</p>
|
| 439 |
-
</dd>
|
| 440 |
-
<dt><span class="docutils literal">MEDFormer.encoder</span> (Transformer Encoder Stack)</dt>
|
| 441 |
-
<dd><p><strong>Operations.</strong> A stack of :class:`~braindecode.models.medformer._EncoderLayer` modules,
|
| 442 |
-
each containing a :class:`~braindecode.models.medformer._MedformerLayer` that implements
|
| 443 |
-
two-stage self-attention. The two-stage mechanism splits self-attention into:</p>
|
| 444 |
-
<p><strong>(a) Intra-Granularity Self-Attention.</strong> For granularity <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 445 |
-
<mi>i</mi>
|
| 446 |
-
</math>, the patch embedding
|
| 447 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 448 |
-
<msup>
|
| 449 |
-
<mi>𝐱</mi>
|
| 450 |
-
<mrow>
|
| 451 |
-
<mo stretchy="false">(</mo>
|
| 452 |
-
<mi>i</mi>
|
| 453 |
-
<mo stretchy="false">)</mo>
|
| 454 |
-
</mrow>
|
| 455 |
-
</msup>
|
| 456 |
-
<mo>∈</mo>
|
| 457 |
-
<msup>
|
| 458 |
-
<mi>ℝ</mi>
|
| 459 |
-
<mrow>
|
| 460 |
-
<msub>
|
| 461 |
-
<mi>N</mi>
|
| 462 |
-
<mi>i</mi>
|
| 463 |
-
</msub>
|
| 464 |
-
<mo>×</mo>
|
| 465 |
-
<mi>D</mi>
|
| 466 |
-
</mrow>
|
| 467 |
-
</msup>
|
| 468 |
-
</math> and router embedding
|
| 469 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 470 |
-
<msup>
|
| 471 |
-
<mi>𝐮</mi>
|
| 472 |
-
<mrow>
|
| 473 |
-
<mo stretchy="false">(</mo>
|
| 474 |
-
<mi>i</mi>
|
| 475 |
-
<mo stretchy="false">)</mo>
|
| 476 |
-
</mrow>
|
| 477 |
-
</msup>
|
| 478 |
-
<mo>∈</mo>
|
| 479 |
-
<msup>
|
| 480 |
-
<mi>ℝ</mi>
|
| 481 |
-
<mrow>
|
| 482 |
-
<mn>1</mn>
|
| 483 |
-
<mo>×</mo>
|
| 484 |
-
<mi>D</mi>
|
| 485 |
-
</mrow>
|
| 486 |
-
</msup>
|
| 487 |
-
</math> are concatenated:</p>
|
| 488 |
-
<div>
|
| 489 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML" display="block">
|
| 490 |
-
<msup>
|
| 491 |
-
<mi>𝐳</mi>
|
| 492 |
-
<mrow>
|
| 493 |
-
<mo stretchy="false">(</mo>
|
| 494 |
-
<mi>i</mi>
|
| 495 |
-
<mo stretchy="false">)</mo>
|
| 496 |
-
</mrow>
|
| 497 |
-
</msup>
|
| 498 |
-
<mo>=</mo>
|
| 499 |
-
<mo stretchy="false">[</mo>
|
| 500 |
-
<msup>
|
| 501 |
-
<mi>𝐱</mi>
|
| 502 |
-
<mrow>
|
| 503 |
-
<mo stretchy="false">(</mo>
|
| 504 |
-
<mi>i</mi>
|
| 505 |
-
<mo stretchy="false">)</mo>
|
| 506 |
-
</mrow>
|
| 507 |
-
</msup>
|
| 508 |
-
<mo>‖</mo>
|
| 509 |
-
<msup>
|
| 510 |
-
<mi>𝐮</mi>
|
| 511 |
-
<mrow>
|
| 512 |
-
<mo stretchy="false">(</mo>
|
| 513 |
-
<mi>i</mi>
|
| 514 |
-
<mo stretchy="false">)</mo>
|
| 515 |
-
</mrow>
|
| 516 |
-
</msup>
|
| 517 |
-
<mo stretchy="false">]</mo>
|
| 518 |
-
<mo>∈</mo>
|
| 519 |
-
<msup>
|
| 520 |
-
<mi>ℝ</mi>
|
| 521 |
-
<mrow>
|
| 522 |
-
<mo stretchy="false">(</mo>
|
| 523 |
-
<msub>
|
| 524 |
-
<mi>N</mi>
|
| 525 |
-
<mi>i</mi>
|
| 526 |
-
</msub>
|
| 527 |
-
<mo>+</mo>
|
| 528 |
-
<mn>1</mn>
|
| 529 |
-
<mo stretchy="false">)</mo>
|
| 530 |
-
<mo>×</mo>
|
| 531 |
-
<mi>D</mi>
|
| 532 |
-
</mrow>
|
| 533 |
-
</msup>
|
| 534 |
-
</math>
|
| 535 |
-
</div>
|
| 536 |
-
<p>Self-attention is applied to update both embeddings:</p>
|
| 537 |
-
<div>
|
| 538 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML" display="block">
|
| 539 |
-
<mtable class="ams-align" displaystyle="true">
|
| 540 |
-
<mtr>
|
| 541 |
-
<mtd>
|
| 542 |
-
<msup>
|
| 543 |
-
<mi>𝐱</mi>
|
| 544 |
-
<mrow>
|
| 545 |
-
<mo stretchy="false">(</mo>
|
| 546 |
-
<mi>i</mi>
|
| 547 |
-
<mo stretchy="false">)</mo>
|
| 548 |
-
</mrow>
|
| 549 |
-
</msup>
|
| 550 |
-
</mtd>
|
| 551 |
-
<mtd>
|
| 552 |
-
<mo>←</mo>
|
| 553 |
-
<msub>
|
| 554 |
-
<mtext>Attn</mtext>
|
| 555 |
-
<mtext>intra</mtext>
|
| 556 |
-
</msub>
|
| 557 |
-
<mo stretchy="false">(</mo>
|
| 558 |
-
<msup>
|
| 559 |
-
<mi>𝐱</mi>
|
| 560 |
-
<mrow>
|
| 561 |
-
<mo stretchy="false">(</mo>
|
| 562 |
-
<mi>i</mi>
|
| 563 |
-
<mo stretchy="false">)</mo>
|
| 564 |
-
</mrow>
|
| 565 |
-
</msup>
|
| 566 |
-
<mo>,</mo>
|
| 567 |
-
<msup>
|
| 568 |
-
<mi>𝐳</mi>
|
| 569 |
-
<mrow>
|
| 570 |
-
<mo stretchy="false">(</mo>
|
| 571 |
-
<mi>i</mi>
|
| 572 |
-
<mo stretchy="false">)</mo>
|
| 573 |
-
</mrow>
|
| 574 |
-
</msup>
|
| 575 |
-
<mo>,</mo>
|
| 576 |
-
<msup>
|
| 577 |
-
<mi>𝐳</mi>
|
| 578 |
-
<mrow>
|
| 579 |
-
<mo stretchy="false">(</mo>
|
| 580 |
-
<mi>i</mi>
|
| 581 |
-
<mo stretchy="false">)</mo>
|
| 582 |
-
</mrow>
|
| 583 |
-
</msup>
|
| 584 |
-
<mo stretchy="false">)</mo>
|
| 585 |
-
</mtd>
|
| 586 |
-
</mtr>
|
| 587 |
-
<mtr>
|
| 588 |
-
<mtd>
|
| 589 |
-
<msup>
|
| 590 |
-
<mi>𝐮</mi>
|
| 591 |
-
<mrow>
|
| 592 |
-
<mo stretchy="false">(</mo>
|
| 593 |
-
<mi>i</mi>
|
| 594 |
-
<mo stretchy="false">)</mo>
|
| 595 |
-
</mrow>
|
| 596 |
-
</msup>
|
| 597 |
-
</mtd>
|
| 598 |
-
<mtd>
|
| 599 |
-
<mo>←</mo>
|
| 600 |
-
<msub>
|
| 601 |
-
<mtext>Attn</mtext>
|
| 602 |
-
<mtext>intra</mtext>
|
| 603 |
-
</msub>
|
| 604 |
-
<mo stretchy="false">(</mo>
|
| 605 |
-
<msup>
|
| 606 |
-
<mi>𝐮</mi>
|
| 607 |
-
<mrow>
|
| 608 |
-
<mo stretchy="false">(</mo>
|
| 609 |
-
<mi>i</mi>
|
| 610 |
-
<mo stretchy="false">)</mo>
|
| 611 |
-
</mrow>
|
| 612 |
-
</msup>
|
| 613 |
-
<mo>,</mo>
|
| 614 |
-
<msup>
|
| 615 |
-
<mi>𝐳</mi>
|
| 616 |
-
<mrow>
|
| 617 |
-
<mo stretchy="false">(</mo>
|
| 618 |
-
<mi>i</mi>
|
| 619 |
-
<mo stretchy="false">)</mo>
|
| 620 |
-
</mrow>
|
| 621 |
-
</msup>
|
| 622 |
-
<mo>,</mo>
|
| 623 |
-
<msup>
|
| 624 |
-
<mi>𝐳</mi>
|
| 625 |
-
<mrow>
|
| 626 |
-
<mo stretchy="false">(</mo>
|
| 627 |
-
<mi>i</mi>
|
| 628 |
-
<mo stretchy="false">)</mo>
|
| 629 |
-
</mrow>
|
| 630 |
-
</msup>
|
| 631 |
-
<mo stretchy="false">)</mo>
|
| 632 |
-
</mtd>
|
| 633 |
-
</mtr>
|
| 634 |
-
</mtable>
|
| 635 |
-
</math>
|
| 636 |
-
</div>
|
| 637 |
-
<p>This captures temporal features within each granularity independently.</p>
|
| 638 |
-
<p><strong>(b) Inter-Granularity Self-Attention.</strong> All router embeddings are concatenated:</p>
|
| 639 |
-
<div>
|
| 640 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML" display="block">
|
| 641 |
-
<mi>𝐔</mi>
|
| 642 |
-
<mo>=</mo>
|
| 643 |
-
<mo stretchy="false">[</mo>
|
| 644 |
-
<msup>
|
| 645 |
-
<mi>𝐮</mi>
|
| 646 |
-
<mrow>
|
| 647 |
-
<mo stretchy="false">(</mo>
|
| 648 |
-
<mn>1</mn>
|
| 649 |
-
<mo stretchy="false">)</mo>
|
| 650 |
-
</mrow>
|
| 651 |
-
</msup>
|
| 652 |
-
<mo>‖</mo>
|
| 653 |
-
<msup>
|
| 654 |
-
<mi>𝐮</mi>
|
| 655 |
-
<mrow>
|
| 656 |
-
<mo stretchy="false">(</mo>
|
| 657 |
-
<mn>2</mn>
|
| 658 |
-
<mo stretchy="false">)</mo>
|
| 659 |
-
</mrow>
|
| 660 |
-
</msup>
|
| 661 |
-
<mo>‖</mo>
|
| 662 |
-
<mi>⋯</mi>
|
| 663 |
-
<mo>‖</mo>
|
| 664 |
-
<msup>
|
| 665 |
-
<mi>𝐮</mi>
|
| 666 |
-
<mrow>
|
| 667 |
-
<mo stretchy="false">(</mo>
|
| 668 |
-
<mi>n</mi>
|
| 669 |
-
<mo stretchy="false">)</mo>
|
| 670 |
-
</mrow>
|
| 671 |
-
</msup>
|
| 672 |
-
<mo stretchy="false">]</mo>
|
| 673 |
-
<mo>∈</mo>
|
| 674 |
-
<msup>
|
| 675 |
-
<mi>ℝ</mi>
|
| 676 |
-
<mrow>
|
| 677 |
-
<mi>n</mi>
|
| 678 |
-
<mo>×</mo>
|
| 679 |
-
<mi>D</mi>
|
| 680 |
-
</mrow>
|
| 681 |
-
</msup>
|
| 682 |
-
</math>
|
| 683 |
-
</div>
|
| 684 |
-
<p>Self-attention among routers exchanges information across granularities:</p>
|
| 685 |
-
<div>
|
| 686 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML" display="block">
|
| 687 |
-
<msup>
|
| 688 |
-
<mi>𝐮</mi>
|
| 689 |
-
<mrow>
|
| 690 |
-
<mo stretchy="false">(</mo>
|
| 691 |
-
<mi>i</mi>
|
| 692 |
-
<mo stretchy="false">)</mo>
|
| 693 |
-
</mrow>
|
| 694 |
-
</msup>
|
| 695 |
-
<mo>←</mo>
|
| 696 |
-
<msub>
|
| 697 |
-
<mtext>Attn</mtext>
|
| 698 |
-
<mtext>inter</mtext>
|
| 699 |
-
</msub>
|
| 700 |
-
<mo stretchy="false">(</mo>
|
| 701 |
-
<msup>
|
| 702 |
-
<mi>𝐮</mi>
|
| 703 |
-
<mrow>
|
| 704 |
-
<mo stretchy="false">(</mo>
|
| 705 |
-
<mi>i</mi>
|
| 706 |
-
<mo stretchy="false">)</mo>
|
| 707 |
-
</mrow>
|
| 708 |
-
</msup>
|
| 709 |
-
<mo>,</mo>
|
| 710 |
-
<mi>𝐔</mi>
|
| 711 |
-
<mo>,</mo>
|
| 712 |
-
<mi>𝐔</mi>
|
| 713 |
-
<mo stretchy="false">)</mo>
|
| 714 |
-
</math>
|
| 715 |
-
</div>
|
| 716 |
-
<p><strong>Role.</strong> Learns representations and correlations within and across temporal scales while
|
| 717 |
-
reducing complexity from <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 718 |
-
<mi>O</mi>
|
| 719 |
-
<mo stretchy="false">(</mo>
|
| 720 |
-
<mo stretchy="false">(</mo>
|
| 721 |
-
<munder>
|
| 722 |
-
<mo movablelimits="true">∑</mo>
|
| 723 |
-
<mi>i</mi>
|
| 724 |
-
</munder>
|
| 725 |
-
<msub>
|
| 726 |
-
<mi>N</mi>
|
| 727 |
-
<mi>i</mi>
|
| 728 |
-
</msub>
|
| 729 |
-
<msup>
|
| 730 |
-
<mo stretchy="false">)</mo>
|
| 731 |
-
<mn>2</mn>
|
| 732 |
-
</msup>
|
| 733 |
-
<mo stretchy="false">)</mo>
|
| 734 |
-
</math> to
|
| 735 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 736 |
-
<mi>O</mi>
|
| 737 |
-
<mo stretchy="false">(</mo>
|
| 738 |
-
<munder>
|
| 739 |
-
<mo movablelimits="true">∑</mo>
|
| 740 |
-
<mi>i</mi>
|
| 741 |
-
</munder>
|
| 742 |
-
<msubsup>
|
| 743 |
-
<mi>N</mi>
|
| 744 |
-
<mi>i</mi>
|
| 745 |
-
<mn>2</mn>
|
| 746 |
-
</msubsup>
|
| 747 |
-
<mo>+</mo>
|
| 748 |
-
<msup>
|
| 749 |
-
<mi>n</mi>
|
| 750 |
-
<mn>2</mn>
|
| 751 |
-
</msup>
|
| 752 |
-
<mo stretchy="false">)</mo>
|
| 753 |
-
</math> through the router mechanism.</p>
|
| 754 |
-
</dd>
|
| 755 |
-
</dl>
|
| 756 |
-
<p><strong>Temporal, Spatial, and Spectral Encoding</strong></p>
|
| 757 |
-
<ul class="simple">
|
| 758 |
-
<li><p><strong>Temporal:</strong> Multiple patch lengths in :attr:`patch_len_list` capture features at several
|
| 759 |
-
temporal granularities, while intra-granularity attention supports long-range temporal
|
| 760 |
-
dependencies.</p></li>
|
| 761 |
-
<li><p><strong>Spatial:</strong> Cross-channel patching embeds inter-channel dependencies by applying kernels
|
| 762 |
-
that span every input channel.</p></li>
|
| 763 |
-
<li><p><strong>Spectral:</strong> Differing patch lengths simulate multiple sampling frequencies analogous to
|
| 764 |
-
clinically relevant bands (e.g., alpha, beta, gamma).</p></li>
|
| 765 |
-
</ul>
|
| 766 |
-
<p><strong>Additional Mechanisms</strong></p>
|
| 767 |
-
<ul class="simple">
|
| 768 |
-
<li><p><strong>Granularity router:</strong> Each granularity <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 769 |
-
<mi>i</mi>
|
| 770 |
-
</math> receives a dedicated router token
|
| 771 |
-
<math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 772 |
-
<msup>
|
| 773 |
-
<mi>𝐮</mi>
|
| 774 |
-
<mrow>
|
| 775 |
-
<mo stretchy="false">(</mo>
|
| 776 |
-
<mi>i</mi>
|
| 777 |
-
<mo stretchy="false">)</mo>
|
| 778 |
-
</mrow>
|
| 779 |
-
</msup>
|
| 780 |
-
</math>. Intra-attention updates the token, and inter-attention exchanges
|
| 781 |
-
aggregated information across scales.</p></li>
|
| 782 |
-
<li><p><strong>Complexity:</strong> Router-mediated two-stage attention maintains <math xmlns="http://www.w3.org/1998/Math/MathML">
|
| 783 |
-
<mi>O</mi>
|
| 784 |
-
<mo stretchy="false">(</mo>
|
| 785 |
-
<msup>
|
| 786 |
-
<mi>T</mi>
|
| 787 |
-
<mn>2</mn>
|
| 788 |
-
</msup>
|
| 789 |
-
<mo stretchy="false">)</mo>
|
| 790 |
-
</math> complexity for
|
| 791 |
-
suitable patch lengths (e.g., power series), preserving transformer-like efficiency while
|
| 792 |
-
modeling multiple granularities.</p></li>
|
| 793 |
-
</ul>
|
| 794 |
-
<section id="parameters">
|
| 795 |
-
<h2>Parameters</h2>
|
| 796 |
-
<dl class="simple">
|
| 797 |
-
<dt>patch_len_list<span class="classifier">list of int, optional</span></dt>
|
| 798 |
-
<dd><p>Patch lengths for multi-granularity patching; each entry selects a temporal scale.
|
| 799 |
-
The default is <span class="docutils literal">[14, 44, 45]</span>.</p>
|
| 800 |
-
</dd>
|
| 801 |
-
<dt>embed_dim<span class="classifier">int, optional</span></dt>
|
| 802 |
-
<dd><p>Embedding dimensionality. The default is <span class="docutils literal">128</span>.</p>
|
| 803 |
-
</dd>
|
| 804 |
-
<dt>num_heads<span class="classifier">int, optional</span></dt>
|
| 805 |
-
<dd><p>Number of attention heads, which must divide :attr:`d_model`. The default is <span class="docutils literal">8</span>.</p>
|
| 806 |
-
</dd>
|
| 807 |
-
<dt>drop_prob<span class="classifier">float, optional</span></dt>
|
| 808 |
-
<dd><p>Dropout probability. The default is <span class="docutils literal">0.1</span>.</p>
|
| 809 |
-
</dd>
|
| 810 |
-
<dt>no_inter_attn<span class="classifier">bool, optional</span></dt>
|
| 811 |
-
<dd><p>If <span class="docutils literal">True</span>, disables inter-granularity attention. The default is <span class="docutils literal">False</span>.</p>
|
| 812 |
-
</dd>
|
| 813 |
-
<dt>num_layers<span class="classifier">int, optional</span></dt>
|
| 814 |
-
<dd><p>Number of encoder layers. The default is <span class="docutils literal">6</span>.</p>
|
| 815 |
-
</dd>
|
| 816 |
-
<dt>dim_feedforward<span class="classifier">int, optional</span></dt>
|
| 817 |
-
<dd><p>Feedforward dimensionality. The default is <span class="docutils literal">256</span>.</p>
|
| 818 |
-
</dd>
|
| 819 |
-
<dt>activation_trans<span class="classifier">nn.Module, optional</span></dt>
|
| 820 |
-
<dd><p>Activation module used in transformer encoder layers. The default is :class:`nn.ReLU`.</p>
|
| 821 |
-
</dd>
|
| 822 |
-
<dt>single_channel<span class="classifier">bool, optional</span></dt>
|
| 823 |
-
<dd><p>If <span class="docutils literal">True</span>, processes each channel independently, increasing capacity and cost. The default is <span class="docutils literal">False</span>.</p>
|
| 824 |
-
</dd>
|
| 825 |
-
<dt>output_attention<span class="classifier">bool, optional</span></dt>
|
| 826 |
-
<dd><p>If <span class="docutils literal">True</span>, returns attention weights for interpretability. The default is <span class="docutils literal">True</span>.</p>
|
| 827 |
-
</dd>
|
| 828 |
-
<dt>activation_class<span class="classifier">nn.Module, optional</span></dt>
|
| 829 |
-
<dd><p>Activation used in the final classification layer. The default is :class:`nn.GELU`.</p>
|
| 830 |
-
</dd>
|
| 831 |
-
</dl>
|
| 832 |
-
</section>
|
| 833 |
-
<section id="notes">
|
| 834 |
-
<h2>Notes</h2>
|
| 835 |
-
<ul class="simple">
|
| 836 |
-
<li><p>MedFormer outperforms strong baselines across six metrics on five MedTS datasets in a
|
| 837 |
-
subject-independent evaluation <a class="citation-reference" href="#medformer2024" id="citation-reference-4" role="doc-biblioref">[Medformer2024]</a>.</p></li>
|
| 838 |
-
<li><p>Cross-channel patching provides the largest F1 improvement in ablation studies (average
|
| 839 |
-
+6.10%), highlighting its importance for MedTS tasks <a class="citation-reference" href="#medformer2024" id="citation-reference-5" role="doc-biblioref">[Medformer2024]</a>.</p></li>
|
| 840 |
-
<li><p>Setting :attr:`no_inter_attn` to <span class="docutils literal">True</span> disables inter-granularity attention while retaining
|
| 841 |
-
intra-granularity attention.</p></li>
|
| 842 |
-
</ul>
|
| 843 |
-
</section>
|
| 844 |
-
<section id="references">
|
| 845 |
-
<h2>References</h2>
|
| 846 |
-
<div role="list" class="citation-list">
|
| 847 |
-
<div class="citation" id="medformer2024" role="doc-biblioentry">
|
| 848 |
-
<span class="label"><span class="fn-bracket">[</span>Medformer2024<span class="fn-bracket">]</span></span>
|
| 849 |
-
<span class="backrefs">(<a role="doc-backlink" href="#citation-reference-1">1</a>,<a role="doc-backlink" href="#citation-reference-2">2</a>,<a role="doc-backlink" href="#citation-reference-3">3</a>,<a role="doc-backlink" href="#citation-reference-4">4</a>,<a role="doc-backlink" href="#citation-reference-5">5</a>)</span>
|
| 850 |
-
<p>Wang, Y., Huang, N., Li, T., Yan, Y., & Zhang, X. (2024).
|
| 851 |
-
Medformer: A Multi-Granularity Patching Transformer for Medical Time-Series Classification.
|
| 852 |
-
In A. Globerson, L. Mackey, D. Belgrave, A. Fan, U. Paquet, J. Tomczak, & C. Zhang (Eds.),
|
| 853 |
-
Advances in Neural Information Processing Systems (Vol. 37, pp. 36314-36341).
|
| 854 |
-
doi:10.52202/079017-1145.</p>
|
| 855 |
-
</div>
|
| 856 |
-
</div>
|
| 857 |
-
<p><strong>Hugging Face Hub integration</strong></p>
|
| 858 |
-
<p>When the optional <span class="docutils literal">huggingface_hub</span> package is installed, all models
|
| 859 |
-
automatically gain the ability to be pushed to and loaded from the
|
| 860 |
-
Hugging Face Hub. Install with:</p>
|
| 861 |
-
<pre class="literal-block">pip install braindecode[hub]</pre>
|
| 862 |
-
<p><strong>Pushing a model to the Hub:</strong></p>
|
| 863 |
-
<p><strong>Loading a model from the Hub:</strong></p>
|
| 864 |
-
<p><strong>Extracting features and replacing the head:</strong></p>
|
| 865 |
-
<p><strong>Saving and restoring full configuration:</strong></p>
|
| 866 |
-
<p>All model parameters (both EEG-specific and model-specific such as
|
| 867 |
-
dropout rates, activation functions, number of filters) are automatically
|
| 868 |
-
saved to the Hub and restored when loading.</p>
|
| 869 |
-
<p>See :ref:`load-pretrained-models` for a complete tutorial.</p>
|
| 870 |
-
</section>
|
| 871 |
-
</main>
|
| 872 |
-
</div>
|
| 873 |
|
| 874 |
## Citation
|
| 875 |
|
| 876 |
-
|
| 877 |
-
*References* section above) and braindecode:
|
| 878 |
|
| 879 |
```bibtex
|
| 880 |
@article{aristimunha2025braindecode,
|
|
|
|
| 10 |
- braindecode
|
| 11 |
- foundation-model
|
| 12 |
- convolutional
|
|
|
|
| 13 |
---
|
| 14 |
|
| 15 |
# MEDFormer
|
| 16 |
|
| 17 |
+
Medformer from Wang et al (2024) [Medformer2024].
|
| 18 |
|
| 19 |
+
> **Architecture-only repository.** Documents the
|
| 20 |
> `braindecode.models.MEDFormer` class. **No pretrained weights are
|
| 21 |
+
> distributed here.** Instantiate the model and train it on your own
|
| 22 |
+
> data.
|
|
|
|
| 23 |
|
| 24 |
## Quick start
|
| 25 |
|
|
|
|
| 38 |
)
|
| 39 |
```
|
| 40 |
|
| 41 |
+
The signal-shape arguments above are illustrative defaults — adjust to
|
| 42 |
+
match your recording.
|
| 43 |
|
| 44 |
## Documentation
|
| 45 |
+
- Full API reference: <https://braindecode.org/stable/generated/braindecode.models.MEDFormer.html>
|
| 46 |
+
- Interactive browser (live instantiation, parameter counts):
|
|
|
|
|
|
|
| 47 |
<https://huggingface.co/spaces/braindecode/model-explorer>
|
| 48 |
- Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/medformer.py#L20>
|
| 49 |
|
|
|
|
| 50 |
|
| 51 |
+
## Architecture
|
| 52 |
+
|
| 53 |
+
|
| 54 |
+
|
| 55 |
+
|
| 56 |
+
## Parameters
|
| 57 |
+
|
| 58 |
+
| Parameter | Type | Description |
|
| 59 |
+
|---|---|---|
|
| 60 |
+
| `patch_len_list` | list of int, optional | Patch lengths for multi-granularity patching; each entry selects a temporal scale. The default is `[14, 44, 45]`. |
|
| 61 |
+
| `embed_dim` | int, optional | Embedding dimensionality. The default is `128`. |
|
| 62 |
+
| `num_heads` | int, optional | Number of attention heads, which must divide :attr:`d_model`. The default is `8`. |
|
| 63 |
+
| `drop_prob` | float, optional | Dropout probability. The default is `0.1`. |
|
| 64 |
+
| `no_inter_attn` | bool, optional | If `True`, disables inter-granularity attention. The default is `False`. |
|
| 65 |
+
| `num_layers` | int, optional | Number of encoder layers. The default is `6`. |
|
| 66 |
+
| `dim_feedforward` | int, optional | Feedforward dimensionality. The default is `256`. |
|
| 67 |
+
| `activation_trans` | nn.Module, optional | Activation module used in transformer encoder layers. The default is :class:`nn.ReLU`. |
|
| 68 |
+
| `single_channel` | bool, optional | If `True`, processes each channel independently, increasing capacity and cost. The default is `False`. |
|
| 69 |
+
| `output_attention` | bool, optional | If `True`, returns attention weights for interpretability. The default is `True`. |
|
| 70 |
+
| `activation_class` | nn.Module, optional | Activation used in the final classification layer. The default is :class:`nn.GELU`. |
|
| 71 |
+
|
| 72 |
+
|
| 73 |
+
## References
|
| 74 |
+
|
| 75 |
+
1. Wang, Y., Huang, N., Li, T., Yan, Y., & Zhang, X. (2024). Medformer: A Multi-Granularity Patching Transformer for Medical Time-Series Classification. In A. Globerson, L. Mackey, D. Belgrave, A. Fan, U. Paquet, J. Tomczak, & C. Zhang (Eds.), Advances in Neural Information Processing Systems (Vol. 37, pp. 36314-36341). doi:10.52202/079017-1145.
|
| 76 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 77 |
|
| 78 |
## Citation
|
| 79 |
|
| 80 |
+
Cite the original architecture paper (see *References* above) and braindecode:
|
|
|
|
| 81 |
|
| 82 |
```bibtex
|
| 83 |
@article{aristimunha2025braindecode,
|