Replace with clean markdown card
Browse files
README.md
CHANGED
|
@@ -13,13 +13,12 @@ tags:
|
|
| 13 |
|
| 14 |
# EEGNeX
|
| 15 |
|
| 16 |
-
EEGNeX model from Chen et al (2024) .
|
| 17 |
|
| 18 |
-
> **Architecture-only repository.**
|
| 19 |
> `braindecode.models.EEGNeX` class. **No pretrained weights are
|
| 20 |
-
> distributed here**
|
| 21 |
-
> data
|
| 22 |
-
> separately.
|
| 23 |
|
| 24 |
## Quick start
|
| 25 |
|
|
@@ -38,246 +37,47 @@ model = EEGNeX(
|
|
| 38 |
)
|
| 39 |
```
|
| 40 |
|
| 41 |
-
The signal-shape arguments above are
|
| 42 |
-
|
| 43 |
|
| 44 |
## Documentation
|
| 45 |
-
|
| 46 |
-
-
|
| 47 |
-
<https://braindecode.org/stable/generated/braindecode.models.EEGNeX.html>
|
| 48 |
-
- Interactive browser with live instantiation:
|
| 49 |
<https://huggingface.co/spaces/braindecode/model-explorer>
|
| 50 |
- Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/eegnex.py#L16>
|
| 51 |
|
| 52 |
-
## Architecture description
|
| 53 |
-
|
| 54 |
-
The block below is the rendered class docstring (parameters,
|
| 55 |
-
references, architecture figure where available).
|
| 56 |
-
|
| 57 |
-
<div class='bd-doc'><main>
|
| 58 |
-
<p>EEGNeX model from Chen et al (2024) [eegnex]_.</p>
|
| 59 |
-
<span style="display:inline-block;padding:2px 8px;border-radius:4px;background:#5cb85c;color:white;font-size:11px;font-weight:600;margin-right:4px;">Convolution</span>
|
| 60 |
-
|
| 61 |
-
|
| 62 |
-
|
| 63 |
-
.. figure:: https://braindecode.org/dev/_static/model/eegnex.jpg
|
| 64 |
-
:align: center
|
| 65 |
-
:alt: EEGNeX Architecture
|
| 66 |
-
:width: 620px
|
| 67 |
-
|
| 68 |
-
.. rubric:: Architectural Overview
|
| 69 |
-
|
| 70 |
-
EEGNeX is a **purely convolutional** architecture that refines the EEGNet-style stem
|
| 71 |
-
and deepens the temporal stack with **dilated temporal convolutions**. The end-to-end
|
| 72 |
-
flow is:
|
| 73 |
-
|
| 74 |
-
- (i) **Block-1/2**: two temporal convolutions ``(1 x L)`` with BN refine a
|
| 75 |
-
learned FIR-like *temporal filter bank* (no pooling yet);
|
| 76 |
-
- (ii) **Block-3**: depthwise **spatial** convolution across electrodes
|
| 77 |
-
``(n_chans x 1)`` with max-norm constraint, followed by ELU → AvgPool (time) → Dropout;
|
| 78 |
-
- (iii) **Block-4/5**: two additional **temporal** convolutions with increasing **dilation**
|
| 79 |
-
to expand the receptive field; the last block applies ELU → AvgPool → Dropout → Flatten;
|
| 80 |
-
- (iv) **Classifier**: a max-norm–constrained linear layer.
|
| 81 |
-
|
| 82 |
-
The published work positions EEGNeX as a compact, conv-only alternative that consistently
|
| 83 |
-
outperforms prior baselines across MOABB-style benchmarks, with the popular
|
| 84 |
-
“EEGNeX-8,32” shorthand denoting *8 temporal filters* and *kernel length 32*.
|
| 85 |
-
|
| 86 |
-
|
| 87 |
-
.. rubric:: Macro Components
|
| 88 |
-
|
| 89 |
-
- **Block-1 / Block-2 — Temporal filter (learned).**
|
| 90 |
-
|
| 91 |
-
- *Operations.*
|
| 92 |
-
- :class:`torch.nn.Conv2d` with kernels ``(1, L)``
|
| 93 |
-
- :class:`torch.nn.BatchNorm2d` (no nonlinearity until Block-3, mirroring a linear FIR analysis stage).
|
| 94 |
-
These layers set up frequency-selective detectors before spatial mixing.
|
| 95 |
-
|
| 96 |
-
- *Interpretability.* Kernels can be inspected as FIR filters; two stacked temporal
|
| 97 |
-
convs allow longer effective kernels without parameter blow-up.
|
| 98 |
-
|
| 99 |
-
- **Block-3 — Spatial projection + condensation.**
|
| 100 |
-
|
| 101 |
-
- *Operations.*
|
| 102 |
-
- :class:`braindecode.modules.Conv2dWithConstraint` with kernel``(n_chans, 1)``
|
| 103 |
-
and ``groups = filter_2`` (depthwise across filters)
|
| 104 |
-
- :class:`torch.nn.BatchNorm2d`
|
| 105 |
-
- :class:`torch.nn.ELU`
|
| 106 |
-
- :class:`torch.nn.AvgPool2d` (time)
|
| 107 |
-
- :class:`torch.nn.Dropout`.
|
| 108 |
-
|
| 109 |
-
**Role**: Learns per-filter spatial patterns over the **full montage** while temporal
|
| 110 |
-
pooling stabilizes and compresses features; max-norm encourages well-behaved spatial
|
| 111 |
-
weights similar to EEGNet practice.
|
| 112 |
-
|
| 113 |
-
- **Block-4 / Block-5 — Dilated temporal integration.**
|
| 114 |
-
|
| 115 |
-
- *Operations.*
|
| 116 |
-
- :class:`torch.nn.Conv2d` with kernels ``(1, k)`` and **dilations**
|
| 117 |
-
(e.g., 2 then 4);
|
| 118 |
-
- :class:`torch.nn.BatchNorm2d`
|
| 119 |
-
- :class:`torch.nn.ELU`
|
| 120 |
-
- :class:`torch.nn.AvgPool2d` (time)
|
| 121 |
-
- :class:`torch.nn.Dropout`
|
| 122 |
-
- :class:`torch.nn.Flatten`.
|
| 123 |
-
|
| 124 |
-
**Role**: Expands the temporal receptive field efficiently to capture rhythms and
|
| 125 |
-
long-range context after condensation.
|
| 126 |
-
|
| 127 |
-
- **Final Classifier — Max-norm linear.**
|
| 128 |
-
|
| 129 |
-
- *Operations.*
|
| 130 |
-
- :class:`braindecode.modules.LinearWithConstraint` maps the flattened
|
| 131 |
-
vector to the target classes; the max-norm constraint regularizes the readout.
|
| 132 |
-
|
| 133 |
-
|
| 134 |
-
.. rubric:: Convolutional Details
|
| 135 |
-
|
| 136 |
-
- **Temporal (where time-domain patterns are learned).**
|
| 137 |
-
Blocks 1-2 learn the primary filter bank (oscillations/transients), while Blocks 4-5
|
| 138 |
-
use **dilation** to integrate over longer horizons without extra pooling. The final
|
| 139 |
-
AvgPool in Block-5 sets the output token rate and helps noise suppression.
|
| 140 |
-
|
| 141 |
-
- **Spatial (how electrodes are processed).**
|
| 142 |
-
A *single* depthwise spatial conv (Block-3) spans the entire electrode set
|
| 143 |
-
(kernel ``(n_chans, 1)``), producing per-temporal-filter topographies; no cross-filter
|
| 144 |
-
mixing occurs at this stage, aiding interpretability.
|
| 145 |
-
|
| 146 |
-
- **Spectral (how frequency content is captured).**
|
| 147 |
-
Frequency selectivity emerges from the learned temporal kernels; dilation broadens effective
|
| 148 |
-
bandwidth coverage by composing multiple scales.
|
| 149 |
-
|
| 150 |
-
.. rubric:: Additional Mechanisms
|
| 151 |
-
|
| 152 |
-
- **EEGNeX-8,32 naming.** “8,32” indicates *8 temporal filters* and *kernel length 32*,
|
| 153 |
-
reflecting the paper's ablation path from EEGNet-8,2 toward thicker temporal kernels
|
| 154 |
-
and a deeper conv stack.
|
| 155 |
-
- **Max-norm constraints.** Spatial (Block-3) and final linear layers use max-norm
|
| 156 |
-
regularization—standard in EEG CNNs—to reduce overfitting and encourage stable spatial
|
| 157 |
-
patterns.
|
| 158 |
-
|
| 159 |
-
.. rubric:: Usage and Configuration
|
| 160 |
-
|
| 161 |
-
- **Kernel schedule.** Start with the canonical **EEGNeX-8,32** (``filter_1=8``,
|
| 162 |
-
``kernel_block_1_2=32``) and keep **Block-3** depth multiplier modest (e.g., 2) to match
|
| 163 |
-
the paper's “pure conv” profile.
|
| 164 |
-
- **Pooling vs. dilation.** Use pooling in Blocks 3 and 5 to control compute and variance;
|
| 165 |
-
increase dilations (Blocks 4-5) to widen temporal context when windows are short.
|
| 166 |
-
- **Regularization.** Combine dropout (Blocks 3 & 5) with max-norm on spatial and
|
| 167 |
-
classifier layers; prefer ELU activations for stable training on small EEG datasets.
|
| 168 |
-
|
| 169 |
-
|
| 170 |
-
- The braindecode implementation follows the paper's conv-only design with five blocks
|
| 171 |
-
and reproduces the depthwise spatial step and dilated temporal stack. See the class
|
| 172 |
-
reference for exact kernel sizes, dilations, and pooling defaults. You can check the
|
| 173 |
-
original implementation at [EEGNexCode]_.
|
| 174 |
-
|
| 175 |
-
.. versionadded:: 1.1
|
| 176 |
-
|
| 177 |
-
|
| 178 |
-
Parameters
|
| 179 |
-
----------
|
| 180 |
-
activation : nn.Module, optional
|
| 181 |
-
Activation function to use. Default is `nn.ELU`.
|
| 182 |
-
depth_multiplier : int, optional
|
| 183 |
-
Depth multiplier for the depthwise convolution. Default is 2.
|
| 184 |
-
filter_1 : int, optional
|
| 185 |
-
Number of filters in the first convolutional layer. Default is 8.
|
| 186 |
-
filter_2 : int, optional
|
| 187 |
-
Number of filters in the second convolutional layer. Default is 32.
|
| 188 |
-
drop_prob: float, optional
|
| 189 |
-
Dropout rate. Default is 0.5.
|
| 190 |
-
kernel_block_4 : tuple[int, int], optional
|
| 191 |
-
Kernel size for block 4. Default is (1, 16).
|
| 192 |
-
dilation_block_4 : tuple[int, int], optional
|
| 193 |
-
Dilation rate for block 4. Default is (1, 2).
|
| 194 |
-
avg_pool_block4 : tuple[int, int], optional
|
| 195 |
-
Pooling size for block 4. Default is (1, 4).
|
| 196 |
-
kernel_block_5 : tuple[int, int], optional
|
| 197 |
-
Kernel size for block 5. Default is (1, 16).
|
| 198 |
-
dilation_block_5 : tuple[int, int], optional
|
| 199 |
-
Dilation rate for block 5. Default is (1, 4).
|
| 200 |
-
avg_pool_block5 : tuple[int, int], optional
|
| 201 |
-
Pooling size for block 5. Default is (1, 8).
|
| 202 |
-
|
| 203 |
-
References
|
| 204 |
-
----------
|
| 205 |
-
.. [eegnex] Chen, X., Teng, X., Chen, H., Pan, Y., & Geyer, P. (2024).
|
| 206 |
-
Toward reliable signals decoding for electroencephalogram: A benchmark
|
| 207 |
-
study to EEGNeX. Biomedical Signal Processing and Control, 87, 105475.
|
| 208 |
-
.. [EEGNexCode] Chen, X., Teng, X., Chen, H., Pan, Y., & Geyer, P. (2024).
|
| 209 |
-
Toward reliable signals decoding for electroencephalogram: A benchmark
|
| 210 |
-
study to EEGNeX. https://github.com/chenxiachan/EEGNeX
|
| 211 |
-
|
| 212 |
-
.. rubric:: Hugging Face Hub integration
|
| 213 |
-
|
| 214 |
-
When the optional ``huggingface_hub`` package is installed, all models
|
| 215 |
-
automatically gain the ability to be pushed to and loaded from the
|
| 216 |
-
Hugging Face Hub. Install with::
|
| 217 |
-
|
| 218 |
-
pip install braindecode[hub]
|
| 219 |
-
|
| 220 |
-
**Pushing a model to the Hub:**
|
| 221 |
-
|
| 222 |
-
.. code::
|
| 223 |
-
from braindecode.models import EEGNeX
|
| 224 |
-
|
| 225 |
-
# Train your model
|
| 226 |
-
model = EEGNeX(n_chans=22, n_outputs=4, n_times=1000)
|
| 227 |
-
# ... training code ...
|
| 228 |
-
|
| 229 |
-
# Push to the Hub
|
| 230 |
-
model.push_to_hub(
|
| 231 |
-
repo_id="username/my-eegnex-model",
|
| 232 |
-
commit_message="Initial model upload",
|
| 233 |
-
)
|
| 234 |
-
|
| 235 |
-
**Loading a model from the Hub:**
|
| 236 |
-
|
| 237 |
-
.. code::
|
| 238 |
-
from braindecode.models import EEGNeX
|
| 239 |
-
|
| 240 |
-
# Load pretrained model
|
| 241 |
-
model = EEGNeX.from_pretrained("username/my-eegnex-model")
|
| 242 |
-
|
| 243 |
-
# Load with a different number of outputs (head is rebuilt automatically)
|
| 244 |
-
model = EEGNeX.from_pretrained("username/my-eegnex-model", n_outputs=4)
|
| 245 |
-
|
| 246 |
-
**Extracting features and replacing the head:**
|
| 247 |
|
| 248 |
-
|
| 249 |
-
import torch
|
| 250 |
|
| 251 |
-
|
| 252 |
-
# Extract encoder features (consistent dict across all models)
|
| 253 |
-
out = model(x, return_features=True)
|
| 254 |
-
features = out["features"]
|
| 255 |
|
| 256 |
-
# Replace the classification head
|
| 257 |
-
model.reset_head(n_outputs=10)
|
| 258 |
|
| 259 |
-
|
| 260 |
|
| 261 |
-
|
| 262 |
-
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 263 |
|
| 264 |
-
config = model.get_config() # all __init__ params
|
| 265 |
-
with open("config.json", "w") as f:
|
| 266 |
-
json.dump(config, f)
|
| 267 |
|
| 268 |
-
|
| 269 |
|
| 270 |
-
|
| 271 |
-
|
| 272 |
-
saved to the Hub and restored when loading.
|
| 273 |
|
| 274 |
-
See :ref:`load-pretrained-models` for a complete tutorial.</main>
|
| 275 |
-
</div>
|
| 276 |
|
| 277 |
## Citation
|
| 278 |
|
| 279 |
-
|
| 280 |
-
*References* section above) and braindecode:
|
| 281 |
|
| 282 |
```bibtex
|
| 283 |
@article{aristimunha2025braindecode,
|
|
|
|
| 13 |
|
| 14 |
# EEGNeX
|
| 15 |
|
| 16 |
+
EEGNeX model from Chen et al (2024) [eegnex].
|
| 17 |
|
| 18 |
+
> **Architecture-only repository.** Documents the
|
| 19 |
> `braindecode.models.EEGNeX` class. **No pretrained weights are
|
| 20 |
+
> distributed here.** Instantiate the model and train it on your own
|
| 21 |
+
> data.
|
|
|
|
| 22 |
|
| 23 |
## Quick start
|
| 24 |
|
|
|
|
| 37 |
)
|
| 38 |
```
|
| 39 |
|
| 40 |
+
The signal-shape arguments above are illustrative defaults — adjust to
|
| 41 |
+
match your recording.
|
| 42 |
|
| 43 |
## Documentation
|
| 44 |
+
- Full API reference: <https://braindecode.org/stable/generated/braindecode.models.EEGNeX.html>
|
| 45 |
+
- Interactive browser (live instantiation, parameter counts):
|
|
|
|
|
|
|
| 46 |
<https://huggingface.co/spaces/braindecode/model-explorer>
|
| 47 |
- Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/eegnex.py#L16>
|
| 48 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 49 |
|
| 50 |
+
## Architecture
|
|
|
|
| 51 |
|
| 52 |
+
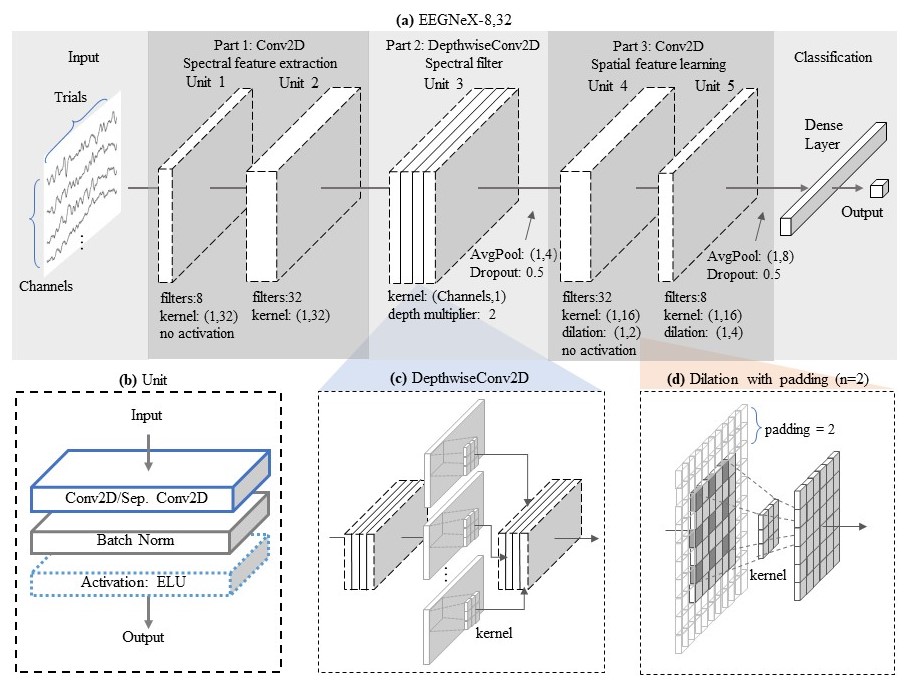
|
|
|
|
|
|
|
|
|
|
| 53 |
|
|
|
|
|
|
|
| 54 |
|
| 55 |
+
## Parameters
|
| 56 |
|
| 57 |
+
| Parameter | Type | Description |
|
| 58 |
+
|---|---|---|
|
| 59 |
+
| `activation` | nn.Module, optional | Activation function to use. Default is `nn.ELU`. |
|
| 60 |
+
| `depth_multiplier` | int, optional | Depth multiplier for the depthwise convolution. Default is 2. |
|
| 61 |
+
| `filter_1` | int, optional | Number of filters in the first convolutional layer. Default is 8. |
|
| 62 |
+
| `filter_2` | int, optional | Number of filters in the second convolutional layer. Default is 32. |
|
| 63 |
+
| `drop_prob: float, optional` | — | Dropout rate. Default is 0.5. |
|
| 64 |
+
| `kernel_block_4` | tuple[int, int], optional | Kernel size for block 4. Default is (1, 16). |
|
| 65 |
+
| `dilation_block_4` | tuple[int, int], optional | Dilation rate for block 4. Default is (1, 2). |
|
| 66 |
+
| `avg_pool_block4` | tuple[int, int], optional | Pooling size for block 4. Default is (1, 4). |
|
| 67 |
+
| `kernel_block_5` | tuple[int, int], optional | Kernel size for block 5. Default is (1, 16). |
|
| 68 |
+
| `dilation_block_5` | tuple[int, int], optional | Dilation rate for block 5. Default is (1, 4). |
|
| 69 |
+
| `avg_pool_block5` | tuple[int, int], optional | Pooling size for block 5. Default is (1, 8). |
|
| 70 |
|
|
|
|
|
|
|
|
|
|
| 71 |
|
| 72 |
+
## References
|
| 73 |
|
| 74 |
+
1. Chen, X., Teng, X., Chen, H., Pan, Y., & Geyer, P. (2024). Toward reliable signals decoding for electroencephalogram: A benchmark study to EEGNeX. Biomedical Signal Processing and Control, 87, 105475.
|
| 75 |
+
2. Chen, X., Teng, X., Chen, H., Pan, Y., & Geyer, P. (2024). Toward reliable signals decoding for electroencephalogram: A benchmark study to EEGNeX. https://github.com/chenxiachan/EEGNeX
|
|
|
|
| 76 |
|
|
|
|
|
|
|
| 77 |
|
| 78 |
## Citation
|
| 79 |
|
| 80 |
+
Cite the original architecture paper (see *References* above) and braindecode:
|
|
|
|
| 81 |
|
| 82 |
```bibtex
|
| 83 |
@article{aristimunha2025braindecode,
|