Replace with clean markdown card
Browse files
README.md
CHANGED
|
@@ -14,13 +14,12 @@ tags:
|
|
| 14 |
|
| 15 |
# CBraMod
|
| 16 |
|
| 17 |
-
**C**\ riss-\ **C**\ ross **Bra**\ in **Mod**\ el for EEG Decoding from Wang et al. (2025) .
|
| 18 |
|
| 19 |
-
> **Architecture-only repository.**
|
| 20 |
> `braindecode.models.CBraMod` class. **No pretrained weights are
|
| 21 |
-
> distributed here**
|
| 22 |
-
> data
|
| 23 |
-
> separately.
|
| 24 |
|
| 25 |
## Quick start
|
| 26 |
|
|
@@ -39,161 +38,43 @@ model = CBraMod(
|
|
| 39 |
)
|
| 40 |
```
|
| 41 |
|
| 42 |
-
The signal-shape arguments above are
|
| 43 |
-
|
| 44 |
|
| 45 |
## Documentation
|
| 46 |
-
|
| 47 |
-
-
|
| 48 |
-
<https://braindecode.org/stable/generated/braindecode.models.CBraMod.html>
|
| 49 |
-
- Interactive browser with live instantiation:
|
| 50 |
<https://huggingface.co/spaces/braindecode/model-explorer>
|
| 51 |
- Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/cbramod.py#L23>
|
| 52 |
|
| 53 |
-
|
| 54 |
-
|
| 55 |
-
|
| 56 |
-
|
| 57 |
-
|
| 58 |
-
|
| 59 |
-
|
| 60 |
-
|
| 61 |
-
|
| 62 |
-
|
| 63 |
-
|
| 64 |
-
|
| 65 |
-
|
| 66 |
-
|
| 67 |
-
|
| 68 |
-
|
| 69 |
-
|
| 70 |
-
|
| 71 |
-
|
| 72 |
-
|
| 73 |
-
|
| 74 |
-
|
| 75 |
-
|
| 76 |
-
|
| 77 |
-
leverages the inherent structural characteristics of EEG signals. The criss-cross approach
|
| 78 |
-
reduces computational complexity (FLOPs reduced by ~32% compared to full attention) while
|
| 79 |
-
improving performance and enabling faster convergence.</p>
|
| 80 |
-
<p><strong>Asymmetric Conditional Positional Encoding (ACPE)</strong></p>
|
| 81 |
-
<p>Rather than using fixed positional embeddings, CBraMod employs <strong>Asymmetric Conditional
|
| 82 |
-
Positional Encoding</strong> that dynamically generates positional embeddings using a convolutional
|
| 83 |
-
network. This enables the model to:</p>
|
| 84 |
-
<ul class="simple">
|
| 85 |
-
<li><p>Capture relative positional information adaptively</p></li>
|
| 86 |
-
<li><p>Handle diverse EEG channel formats (different channel counts and reference schemes)</p></li>
|
| 87 |
-
<li><p>Generalize to arbitrary downstream EEG formats without retraining</p></li>
|
| 88 |
-
<li><p>Support various reference schemes (earlobe, average, REST, bipolar)</p></li>
|
| 89 |
-
</ul>
|
| 90 |
-
<p><strong>Pretraining Highlights</strong></p>
|
| 91 |
-
<ul class="simple">
|
| 92 |
-
<li><p><strong>Pretraining Dataset</strong>: Temple University Hospital EEG Corpus (TUEG), the largest public EEG corpus</p></li>
|
| 93 |
-
<li><p><strong>Pretraining Task</strong>: Self-supervised masked EEG patch reconstruction from both time-domain
|
| 94 |
-
and frequency-domain EEG signals</p></li>
|
| 95 |
-
<li><p><strong>Model Parameters</strong>: ~4.0M parameters (very compact compared to other foundation models)</p></li>
|
| 96 |
-
<li><p><strong>Fast Convergence</strong>: Achieves decent results in first epoch on downstream tasks,
|
| 97 |
-
full convergence within ~10 epochs (vs. ~30 for supervised models like EEGConformer)</p></li>
|
| 98 |
-
</ul>
|
| 99 |
-
<p><strong>Macro Components</strong></p>
|
| 100 |
-
<ul class="simple">
|
| 101 |
-
<li><p><strong>Patch Encoding Network</strong>: Converts raw EEG patches into embeddings</p></li>
|
| 102 |
-
<li><p><strong>Asymmetric Conditional Positional Encoding (ACPE)</strong>: Generates spatial-temporal positional
|
| 103 |
-
embeddings adaptively from input EEG format</p></li>
|
| 104 |
-
<li><p><strong>Criss-Cross Transformer Blocks</strong> (12 layers): Alternates spatial and temporal attention
|
| 105 |
-
to learn EEG representations</p></li>
|
| 106 |
-
<li><p><strong>Reconstruction Head</strong>: Reconstructs masked EEG patches during pretraining</p></li>
|
| 107 |
-
<li><dl class="simple">
|
| 108 |
-
<dt><strong>Task head</strong> (<span class="docutils literal">final_layer</span>): flatten summary tokens across patches and map to</dt>
|
| 109 |
-
<dd><p><span class="docutils literal">n_outputs</span>; if <span class="docutils literal">return_encoder_output=True</span>, return the encoder features instead.</p>
|
| 110 |
-
</dd>
|
| 111 |
-
</dl>
|
| 112 |
-
</li>
|
| 113 |
-
</ul>
|
| 114 |
-
<p>The model is highly efficient, requiring only ~318.9M FLOPs on a typical 16-channel, 10-second
|
| 115 |
-
EEG recording (significantly lower than full attention baselines).</p>
|
| 116 |
-
<p><strong>Known Limitations</strong></p>
|
| 117 |
-
<ul class="simple">
|
| 118 |
-
<li><p><strong>Data Quality</strong>: TUEG corpus contains "dirty data"; pretraining used crude filtering,
|
| 119 |
-
reducing available pre-training data</p></li>
|
| 120 |
-
<li><p><strong>Channel Dependency</strong>: Performance degrades with very sparse electrode setups (e.g., <4 channels)</p></li>
|
| 121 |
-
<li><p><strong>Computational Resources</strong>: While efficient, foundation models have higher deployment
|
| 122 |
-
requirements than lightweight models</p></li>
|
| 123 |
-
<li><p><strong>Limited Scaling Exploration</strong>: Future work should explore scaling laws at billion-parameter levels
|
| 124 |
-
and integration with large pre-trained vision/language models</p></li>
|
| 125 |
-
</ul>
|
| 126 |
-
<aside class="admonition important">
|
| 127 |
-
<p class="admonition-title">Important</p>
|
| 128 |
-
<p><strong>Pre-trained Weights Available</strong></p>
|
| 129 |
-
<p>This model has pre-trained weights available on the Hugging Face Hub.
|
| 130 |
-
You can load them using:</p>
|
| 131 |
-
<p>To push your own trained model to the Hub:</p>
|
| 132 |
-
<p>Requires installing <span class="docutils literal">braindecode[hug]</span> for Hub integration.</p>
|
| 133 |
-
</aside>
|
| 134 |
-
<section id="parameters">
|
| 135 |
-
<h2>Parameters</h2>
|
| 136 |
-
<dl class="simple">
|
| 137 |
-
<dt>patch_size<span class="classifier">int, default=200</span></dt>
|
| 138 |
-
<dd><p>Temporal patch size in samples (200 samples = 1 second at 200 Hz).</p>
|
| 139 |
-
</dd>
|
| 140 |
-
<dt>dim_feedforward<span class="classifier">int, default=800</span></dt>
|
| 141 |
-
<dd><p>Dimension of the feedforward network in Transformer layers.</p>
|
| 142 |
-
</dd>
|
| 143 |
-
<dt>n_layer<span class="classifier">int, default=12</span></dt>
|
| 144 |
-
<dd><p>Number of Transformer layers.</p>
|
| 145 |
-
</dd>
|
| 146 |
-
<dt>nhead<span class="classifier">int, default=8</span></dt>
|
| 147 |
-
<dd><p>Number of attention heads.</p>
|
| 148 |
-
</dd>
|
| 149 |
-
<dt>activation<span class="classifier">type[nn.Module], default=nn.GELU</span></dt>
|
| 150 |
-
<dd><p>Activation function used in Transformer feedforward layers.</p>
|
| 151 |
-
</dd>
|
| 152 |
-
<dt>emb_dim<span class="classifier">int, default=200</span></dt>
|
| 153 |
-
<dd><p>Output embedding dimension.</p>
|
| 154 |
-
</dd>
|
| 155 |
-
<dt>drop_prob<span class="classifier">float, default=0.1</span></dt>
|
| 156 |
-
<dd><p>Dropout probability.</p>
|
| 157 |
-
</dd>
|
| 158 |
-
<dt>return_encoder_output<span class="classifier">bool, default=False</span></dt>
|
| 159 |
-
<dd><p>If false (default), the features are flattened and passed through a final linear layer
|
| 160 |
-
to produce class logits of size <span class="docutils literal">n_outputs</span>.
|
| 161 |
-
If True, the model returns the encoder output features.</p>
|
| 162 |
-
</dd>
|
| 163 |
-
</dl>
|
| 164 |
-
</section>
|
| 165 |
-
<section id="references">
|
| 166 |
-
<h2>References</h2>
|
| 167 |
-
<div role="list" class="citation-list">
|
| 168 |
-
<div class="citation" id="cbramod" role="doc-biblioentry">
|
| 169 |
-
<span class="label"><span class="fn-bracket">[</span><a role="doc-backlink" href="#citation-reference-1">cbramod</a><span class="fn-bracket">]</span></span>
|
| 170 |
-
<p>Wang, J., Zhao, S., Luo, Z., Zhou, Y., Jiang, H., Li, S., Li, T., & Pan, G. (2025).
|
| 171 |
-
CBraMod: A Criss-Cross Brain Foundation Model for EEG Decoding.
|
| 172 |
-
In The Thirteenth International Conference on Learning Representations (ICLR 2025).
|
| 173 |
-
<a class="reference external" href="https://arxiv.org/abs/2412.07236">https://arxiv.org/abs/2412.07236</a></p>
|
| 174 |
-
</div>
|
| 175 |
-
</div>
|
| 176 |
-
<p><strong>Hugging Face Hub integration</strong></p>
|
| 177 |
-
<p>When the optional <span class="docutils literal">huggingface_hub</span> package is installed, all models
|
| 178 |
-
automatically gain the ability to be pushed to and loaded from the
|
| 179 |
-
Hugging Face Hub. Install with:</p>
|
| 180 |
-
<pre class="literal-block">pip install braindecode[hub]</pre>
|
| 181 |
-
<p><strong>Pushing a model to the Hub:</strong></p>
|
| 182 |
-
<p><strong>Loading a model from the Hub:</strong></p>
|
| 183 |
-
<p><strong>Extracting features and replacing the head:</strong></p>
|
| 184 |
-
<p><strong>Saving and restoring full configuration:</strong></p>
|
| 185 |
-
<p>All model parameters (both EEG-specific and model-specific such as
|
| 186 |
-
dropout rates, activation functions, number of filters) are automatically
|
| 187 |
-
saved to the Hub and restored when loading.</p>
|
| 188 |
-
<p>See :ref:`load-pretrained-models` for a complete tutorial.</p>
|
| 189 |
-
</section>
|
| 190 |
-
</main>
|
| 191 |
-
</div>
|
| 192 |
|
| 193 |
## Citation
|
| 194 |
|
| 195 |
-
|
| 196 |
-
*References* section above) and braindecode:
|
| 197 |
|
| 198 |
```bibtex
|
| 199 |
@article{aristimunha2025braindecode,
|
|
|
|
| 14 |
|
| 15 |
# CBraMod
|
| 16 |
|
| 17 |
+
**C**\ riss-\ **C**\ ross **Bra**\ in **Mod**\ el for EEG Decoding from Wang et al. (2025) [cbramod].
|
| 18 |
|
| 19 |
+
> **Architecture-only repository.** Documents the
|
| 20 |
> `braindecode.models.CBraMod` class. **No pretrained weights are
|
| 21 |
+
> distributed here.** Instantiate the model and train it on your own
|
| 22 |
+
> data.
|
|
|
|
| 23 |
|
| 24 |
## Quick start
|
| 25 |
|
|
|
|
| 38 |
)
|
| 39 |
```
|
| 40 |
|
| 41 |
+
The signal-shape arguments above are illustrative defaults — adjust to
|
| 42 |
+
match your recording.
|
| 43 |
|
| 44 |
## Documentation
|
| 45 |
+
- Full API reference: <https://braindecode.org/stable/generated/braindecode.models.CBraMod.html>
|
| 46 |
+
- Interactive browser (live instantiation, parameter counts):
|
|
|
|
|
|
|
| 47 |
<https://huggingface.co/spaces/braindecode/model-explorer>
|
| 48 |
- Source on GitHub: <https://github.com/braindecode/braindecode/blob/master/braindecode/models/cbramod.py#L23>
|
| 49 |
|
| 50 |
+
|
| 51 |
+
## Architecture
|
| 52 |
+
|
| 53 |
+
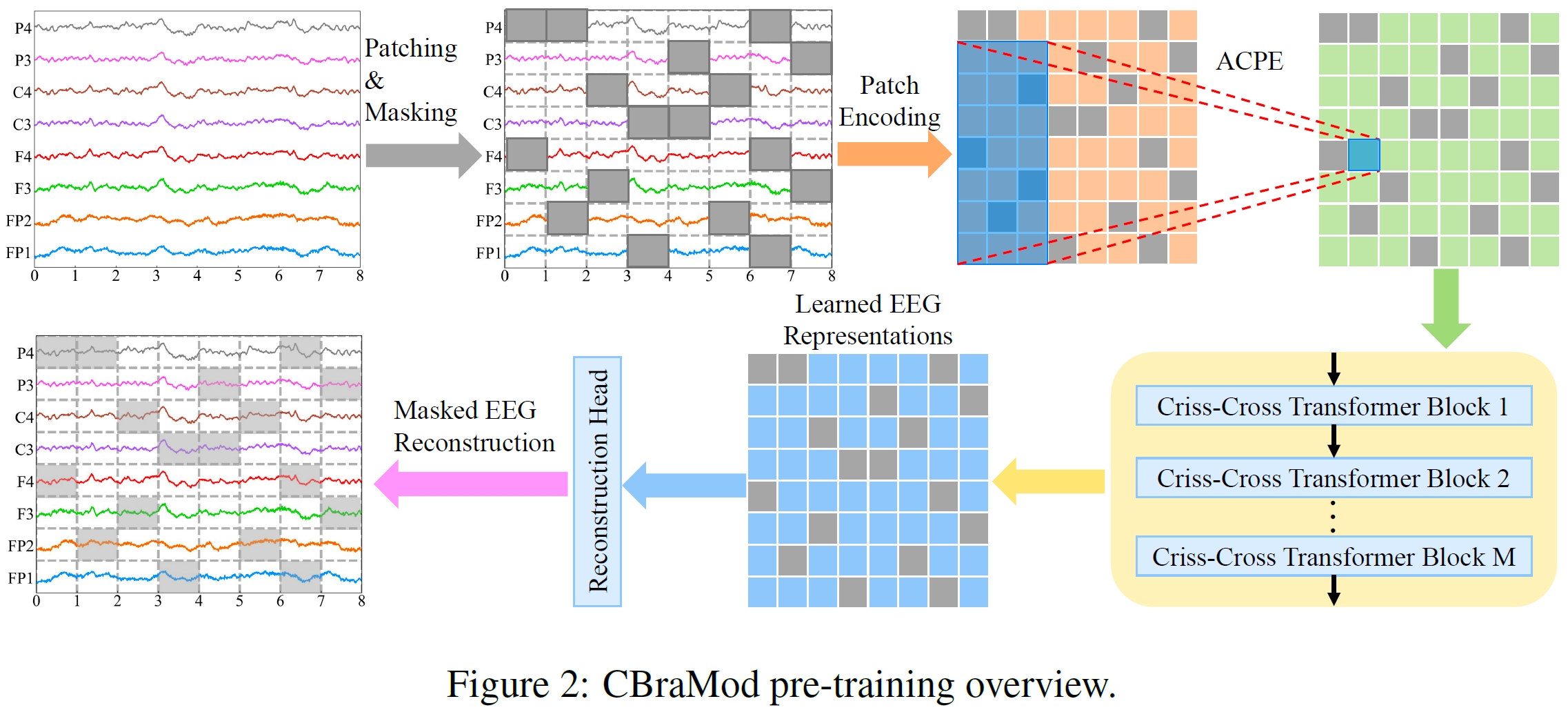
|
| 54 |
+
|
| 55 |
+
|
| 56 |
+
## Parameters
|
| 57 |
+
|
| 58 |
+
| Parameter | Type | Description |
|
| 59 |
+
|---|---|---|
|
| 60 |
+
| `patch_size` | int, default=200 | Temporal patch size in samples (200 samples = 1 second at 200 Hz). |
|
| 61 |
+
| `dim_feedforward` | int, default=800 | Dimension of the feedforward network in Transformer layers. |
|
| 62 |
+
| `n_layer` | int, default=12 | Number of Transformer layers. |
|
| 63 |
+
| `nhead` | int, default=8 | Number of attention heads. |
|
| 64 |
+
| `activation` | type[nn.Module], default=nn.GELU | Activation function used in Transformer feedforward layers. |
|
| 65 |
+
| `emb_dim` | int, default=200 | Output embedding dimension. |
|
| 66 |
+
| `drop_prob` | float, default=0.1 | Dropout probability. |
|
| 67 |
+
| `return_encoder_output` | bool, default=False | If false (default), the features are flattened and passed through a final linear layer to produce class logits of size `n_outputs`. If True, the model returns the encoder output features. |
|
| 68 |
+
|
| 69 |
+
|
| 70 |
+
## References
|
| 71 |
+
|
| 72 |
+
1. Wang, J., Zhao, S., Luo, Z., Zhou, Y., Jiang, H., Li, S., Li, T., & Pan, G. (2025). CBraMod: A Criss-Cross Brain Foundation Model for EEG Decoding. In The Thirteenth International Conference on Learning Representations (ICLR 2025). https://arxiv.org/abs/2412.07236
|
| 73 |
+
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 74 |
|
| 75 |
## Citation
|
| 76 |
|
| 77 |
+
Cite the original architecture paper (see *References* above) and braindecode:
|
|
|
|
| 78 |
|
| 79 |
```bibtex
|
| 80 |
@article{aristimunha2025braindecode,
|