markdown stringlengths 0 1.02M | code stringlengths 0 832k | output stringlengths 0 1.02M | license stringlengths 3 36 | path stringlengths 6 265 | repo_name stringlengths 6 127 |
|---|---|---|---|---|---|
OptimizersWe want to update the generator and discriminator variables separately. So we need to get the variables for each part and build optimizers for the two parts. To get all the trainable variables, we use `tf.trainable_variables()`. This creates a list of all the variables we've defined in our graph.For the gene... | # Optimizers
learning_rate = 0.002
# Get the trainable_variables, split into G and D parts
t_vars = tf.trainable_variables()
g_vars = [var for var in t_vars if var.name.startswith('generator')]
d_vars = [var for var in t_vars if var.name.startswith('discriminator')]
d_train_opt = tf.train.AdamOptimizer(learning_rate)... | _____no_output_____ | Apache-2.0 | Intro_to_GANs_Exercises.ipynb | agoila/gan_mnist |
Training | batch_size = 100
epochs = 100
samples = []
losses = []
saver = tf.train.Saver(var_list = g_vars)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for e in range(epochs):
for ii in range(mnist.train.num_examples//batch_size):
batch = mnist.train.next_batch(batch_size)
... | Epoch 1/100... Discriminator Loss: 0.3678... Generator Loss: 3.5897
Epoch 2/100... Discriminator Loss: 0.3645... Generator Loss: 3.6309
Epoch 3/100... Discriminator Loss: 0.4357... Generator Loss: 3.4136
Epoch 4/100... Discriminator Loss: 0.7224... Generator Loss: 6.2652
Epoch 5/100... Discriminator Loss: 0.6075... Gen... | Apache-2.0 | Intro_to_GANs_Exercises.ipynb | agoila/gan_mnist |
Training lossHere we'll check out the training losses for the generator and discriminator. | %matplotlib inline
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
losses = np.array(losses)
plt.plot(losses.T[0], label='Discriminator')
plt.plot(losses.T[1], label='Generator')
plt.title("Training Losses")
plt.legend() | _____no_output_____ | Apache-2.0 | Intro_to_GANs_Exercises.ipynb | agoila/gan_mnist |
Generator samples from trainingHere we can view samples of images from the generator. First we'll look at images taken while training. | def view_samples(epoch, samples):
fig, axes = plt.subplots(figsize=(7,7), nrows=4, ncols=4, sharey=True, sharex=True)
for ax, img in zip(axes.flatten(), samples[epoch]):
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
im = ax.imshow(img.reshape((28,28)), cmap='Greys_r')
... | _____no_output_____ | Apache-2.0 | Intro_to_GANs_Exercises.ipynb | agoila/gan_mnist |
These are samples from the final training epoch. You can see the generator is able to reproduce numbers like 5, 7, 3, 0, 9. Since this is just a sample, it isn't representative of the full range of images this generator can make. | _ = view_samples(-1, samples) | _____no_output_____ | Apache-2.0 | Intro_to_GANs_Exercises.ipynb | agoila/gan_mnist |
Below I'm showing the generated images as the network was training, every 10 epochs. With bonus optical illusion! | rows, cols = 10, 6
fig, axes = plt.subplots(figsize=(7,12), nrows=rows, ncols=cols, sharex=True, sharey=True)
for sample, ax_row in zip(samples[::int(len(samples)/rows)], axes):
for img, ax in zip(sample[::int(len(sample)/cols)], ax_row):
ax.imshow(img.reshape((28,28)), cmap='Greys_r')
ax.xaxis.set... | _____no_output_____ | Apache-2.0 | Intro_to_GANs_Exercises.ipynb | agoila/gan_mnist |
It starts out as all noise. Then it learns to make only the center white and the rest black. You can start to see some number like structures appear out of the noise. Looks like 1, 9, and 8 show up first. Then, it learns 5 and 3. Sampling from the generatorWe can also get completely new images from the generator by us... | saver = tf.train.Saver(var_list=g_vars)
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('checkpoints'))
sample_z = np.random.uniform(-1, 1, size=(16, z_size))
gen_samples = sess.run(
generator(input_z, input_size, n_units=g_hidden_size, reuse=True, alpha=alpha),
... | INFO:tensorflow:Restoring parameters from checkpoints\generator.ckpt
| Apache-2.0 | Intro_to_GANs_Exercises.ipynb | agoila/gan_mnist |
Intro to Machine Learning with Classification Contents1. **Loading** iris dataset2. Splitting into **train**- and **test**-set3. Creating a **model** and training it4. **Predicting** test set5. **Evaluating** the result6. Selecting **features** This notebook will introduce you to Machine Learning and classification, ... | %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from sklearn import datasets | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
1. Loading iris dataset We load the dataset from the datasets module in sklearn. | iris = datasets.load_iris() | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
This dataset contains information about iris flowers. Every entry describes a flower, more specifically its - sepal length- sepal width- petal length- petal widthSo every entry has four columns. 
df["target"] = iris.target
df.sample(n=10) # show 10 random rows | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
There are 3 different species of irises in the dataset. Every species has 50 samples, so there are 150 entries in total.We can confirm this by checking the "data"-element of the iris variable. The "data"-element is a 2D-array that contains all our entries. We can use the python function `.shape` to check its dimensions... | iris.data.shape | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
To get an example of the data, we can print the first ten rows: | print(iris.data[0:10, :]) # 0:10 gets rows 0-10, : gets all the columns | [[5.1 3.5 1.4 0.2]
[4.9 3. 1.4 0.2]
[4.7 3.2 1.3 0.2]
[4.6 3.1 1.5 0.2]
[5. 3.6 1.4 0.2]
[5.4 3.9 1.7 0.4]
[4.6 3.4 1.4 0.3]
[5. 3.4 1.5 0.2]
[4.4 2.9 1.4 0.2]
[4.9 3.1 1.5 0.1]]
| MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
The labels that we're looking for are in the "target"-element of the iris variable. This 1D-array contains the iris species for each of the entries. | iris.target.shape | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
Let's have a look at the target values: | print(iris.target) | [0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
0 0 0 0 0 0 0 0 0 0 0 0 0 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 2 2 2 2 2 2 2 2 2 2 2
2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2
2 2]
| MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
There are three categories so each entry will be classified as 0, 1 or 2. To get the names of the corresponding species we can print `target_names`. | print(iris.target_names) | ['setosa' 'versicolor' 'virginica']
| MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
The iris variable is a dataset from sklearn and also contains a description of itself. We already provided the information you need to know about the data, but if you want to check, you can print the `.DESCR` method of the iris dataset. | print(iris.DESCR) | .. _iris_dataset:
Iris plants dataset
--------------------
**Data Set Characteristics:**
:Number of Instances: 150 (50 in each of three classes)
:Number of Attributes: 4 numeric, predictive attributes and the class
:Attribute Information:
- sepal length in cm
- sepal width in cm
-... | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
Now we have a good idea what our data looks like.Our task now is to solve a **supervised** learning problem: Predict the species of an iris using the measurements that serve as our so-called **features**. | # First, we store the features we use and the labels we want to predict into two different variables
X = iris.data
y = iris.target | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
2. Splitting into train- and test-set We want to evaluate our model on data with labels that our model has not seen yet. This will give us an idea on how well the model can predict new data, and makes sure we are not [overfitting](https://en.wikipedia.org/wiki/Overfitting). If we would test and train on the same data,... | from sklearn.model_selection import train_test_split
??train_test_split
X_train, X_test, y_train, y_test = train_test_split(iris.data, iris.target, test_size=0.33, stratify=iris.target)# TODO: split iris.data and iris.target into test and train | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
We can now check the size of the resulting arrays. The shapes should be `(100, 4)`, `(100,)`, `(50, 4)` and `(50,)`. | print("X_train shape: {}, y_train shape: {}".format(X_train.shape, y_train.shape))
print("X_test shape: {} , y_test shape: {}".format(X_test.shape, y_test.shape)) | X_train shape: (100, 4), y_train shape: (100,)
X_test shape: (50, 4) , y_test shape: (50,)
| MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
3. Creating a model and training it Now we will give the data to a model. We will use a Decision Tree Classifier model for this.This model will create a decision tree based on the X_train and y_train values and include decisions like this: # TODO: create a decision tree classifier | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
The model is still empty and doesn't know anything. Train (fit) it with our train-data, so that it learns things about our iris-dataset. | model = model.fit(X_train, y_train)# TODO: fit the train-data to the model | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
4. Predicting test set We now have a model that contains a decision tree. This decision tree knows how to turn our X_train values into y_train values. We will now let it run on our X_test values and have a look at the result.We don't want to overwrite our actual y_test values, so we store the predicted y_test values a... | y_pred = model.predict(X_test)# TODO: predict y_pred from X_test | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
5. Evaluating the result We now have y_test (the real values for X_test) and y_pred. We can print these values and compare them, to get an idea of how good the model predicted the data. | print(y_test)
print("-"*75) # print a line
print(y_pred) | [0 2 0 2 2 1 1 0 1 1 2 0 2 0 2 1 2 0 0 1 0 2 2 1 1 1 2 0 1 2 2 0 1 0 0 1 1
2 2 0 1 1 1 0 2 0 1 0 0 2]
---------------------------------------------------------------------------
[0 2 0 2 1 1 1 0 1 1 2 0 2 0 2 1 2 0 0 1 0 2 2 1 1 1 2 0 2 2 2 0 1 0 0 2 1
2 2 0 1 1 1 0 2 0 1 0 0 2]
| MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
If we look at the values closely, we can discover that all but two values are predicted correctly. However, it is bothersome to compare the numbers one by one. There are only thirty of them, but what if there were one hundred? We will need an easier method to compare our results.Luckily, this can also be found in sklea... | from sklearn import metrics
accuracy = metrics.accuracy_score(y_test, y_pred) # TODO: calculate accuracy score of y_test and y_pred
print(accuracy) | 0.94
| MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
That's pretty good, isn't it?To understand what our classifier actually did, have a look at the following picture: 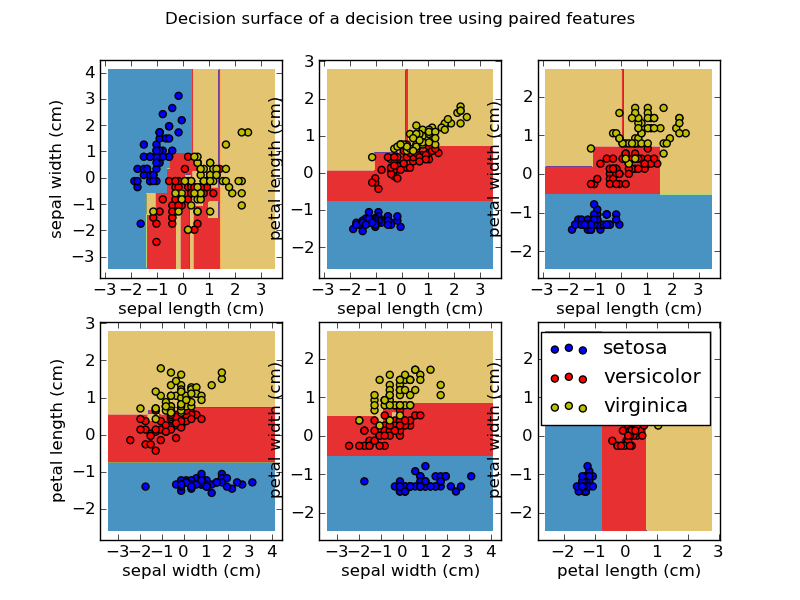 We see the distribution of all our features, compared with each other. Some have very clear distinction... | from sklearn.feature_selection import SelectKBest, chi2
selector = SelectKBest(chi2, k=2) # TODO: create a selector for the 2 best features and fit X_train and y_train to it
X_train, X_test, y_train, y_test = train_test_split(iris.data, iris.target, test_size=0.33, random_state=42)
selector = selector.fit(X_train, y_t... | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
We can check which features our selector selected, using the following function: | print(selector.get_support()) | [False False True True]
| MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
It gives us an array of True and False values that represent the columns of the original X_train. The values that are marked by True are considered the most informative by the selector. Let's use the selector to select (transform) these features from the X_train values. | X_train_new = selector.transform(X_train) # TODO: use selector to transform X_train | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
The dimensions of X_train have now changed: | X_train_new.shape | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
If we want to use these values in our model, we will need to adjust X_test as well. We would get in trouble later if X_train has only 2 columns and X_test has 4. So perform the same selection on X_test. | X_test_new = selector.transform(X_test) # TODO: use selector to transform X_test
X_test_new.shape | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
Now we can repeat the earlier steps: create a model, fit the data to it and predict our y_test values. | model = tree.DecisionTreeClassifier(random_state=0) # TODO: create model as before
model = model.fit(X_train_new, y_train) # TODO: fit model as before, but use X_train_new
y_pred = model.predict(X_test_new) # TODO: predict values as before, but use X_test_new | _____no_output_____ | MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
Let's have a look at the accuracy score of our new prediction. | accuracy = metrics.accuracy_score(y_test, y_pred) # TODO: calculate accuracy score of y_test and y_pred
print(accuracy) # TODO: calculate accuracy score as before | 1.0
| MIT | PXL_DIGITAL_JAAR_2/Data Advanced/Bestanden/notebooks_data/Machine Learning Exercises/Own solutions/Machine_Learning_1_Classification.ipynb | Limoentaart/PXL_IT_JAAR_1 |
This Dataset was taken online | import pandas as pd
import numpy as np
dataset = pd.read_csv('url_dataset.csv')
#deleting all columns except url
dataset.drop(dataset.columns.difference(['URL']), 1, inplace=True)
dataset.head(5)
dataset.to_csv('cleaned_link_dataset.csv')
#split protocol from the other part of link
cleaned_dataset = pd.read_csv('cleane... | _____no_output_____ | MIT | Link Dataset Cleaning.ipynb | geridashja/phishing-links-detection |
420-A52-SF - Algorithmes d'apprentissage supervisé - Hiver 2020 - Spécialisation technique en Intelligence Artificielle - Mikaël Swawola, M.Sc.**Objectif:** cette séance de travaux pratique est consacrée à la mise en oeuvre de l'ensemble des connaissances acq... | %reload_ext autoreload
%autoreload 2
%matplotlib inline | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
0 - Chargement des bibliothèques | # Manipulation de données
import numpy as np
import pandas as pd
# Visualisation de données
import matplotlib.pyplot as plt
import seaborn as sns
# Configuration de la visualisation
sns.set(style="darkgrid", rc={'figure.figsize':(11.7,8.27)}) | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
1 - Lecture du jeu de données *NBA* **Lire le fichier `NBA_train.csv`** | # Compléter le code ci-dessous ~ 1 ligne
NBA = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Afficher les dix premières lignes de la trame de données** | # Compléter le code ci-dessous ~ 1 ligne
None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
Ci-dessous, la description des différentes variables explicatives du jeu de données| Variable | Description || ------------- |:-------------------------------------------------------------:|| SeasonEnd | Année de fin de la saison ... | # Compléter le code ci-dessous ~ 1 ligne
None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Stocker le nombre de lignes du jeu de donnée (nombre d'exemples d'entraînement) dans la variable `m`** | # Compléter le code ci-dessous ~ 1 ligne
m = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Stocker le nombre de victoires au cours de la saison dans la variable `y`. Il s'agira de la variable que l'on cherche à prédire** | # Compléter le code ci-dessous ~ 1 ligne
y = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Créer la matrice des prédicteurs `X`.** Indice: `X` doit avoir 2 colonnes... | # Compléter le code ci-dessous ~ 3 lignes
X = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Vérifier la dimension de la matrice des prédicteurs `X`. Quelle est la dimension de `X` ?** | # Compléter le code ci-dessous ~ 1 ligne
None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Créer le modèle de référence (baseline)** | # Compléter le code ci-dessous ~ 1 ligne
y_baseline = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**À l'aide de l'équation normale, trouver les paramètres optimaux du modèle de régression linéaire simple** | # Compléter le code ci-dessous ~ 1 ligne
theta = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Calculer la somme des carrées des erreurs (SSE)** | # Compléter le code ci-dessous ~ 1 ligne
SSE = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Calculer la racine carrée de l'erreur quadratique moyenne (RMSE)** | # Compléter le code ci-dessous ~ 1 ligne
RMSE = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Calculer le coefficient de détermination $R^2$** | # Compléter le code ci-dessous ~ 1-2 lignes
R2 = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Affichage des résultats** | fig, ax = plt.subplots()
ax.scatter(x1, y,label="Data points")
reg_x = np.linspace(-1000,1000,50)
reg_y = theta[0] + np.linspace(-1000,1000,50)* theta[1]
ax.plot(reg_x, np.repeat(y_baseline,50), color='#777777', label="Baseline", lw=2)
ax.plot(reg_x, reg_y, color="g", lw=2, label="Modèle")
ax.set_xlabel("Différence de ... | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
3 - Régression linéaire multiple Nous allons maintenant tenter de prédire le nombre de points obtenus par une équipe donnée au cours de la saison régulière en fonction des autres variables explicatives disponibles. Nous allons mettre en oeuvre plusieurs modèles de régression linéaire multiple **Stocker le nombre de po... | # Compléter le code ci-dessous ~ 1 ligne
y = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Créer la matrice des prédicteurs `X` à partir des variables `2PA` et `3PA`** | # Compléter le code ci-dessous ~ 3 lignes
X = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Vérifier la dimension de la matrice des prédicteurs `X`. Quelle est la dimension de `X` ?** | # Compléter le code ci-dessous ~ 1 ligne
None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Créer le modèle de référence (baseline)** | # Compléter le code ci-dessous ~ 1 ligne
y_baseline = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**À l'aide de l'équation normale, trouver les paramètres optimaux du modèle de régression linéaire** | # Compléter le code ci-dessous ~ 1 ligne
theta = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Calculer la somme des carrées des erreurs (SSE)** | # Compléter le code ci-dessous ~ 1 ligne
SSE = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Calculer la racine carrée de l'erreur quadratique moyenne (RMSE)** | # Compléter le code ci-dessous ~ 1 ligne
RMSE = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
**Calculer le coefficient de détermination $R^2$** | # Compléter le code ci-dessous ~ 1-2 lignes
R2 = None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
3 - Ajouter les variables explicatives FTA et AST **Recommencer les étapes ci-dessus en incluant les variables FTA et AST** | None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
4 - Ajouter les variables explicatives ORB et STL **Recommencer les étapes ci-dessus en incluant les variables ORB et STL** | None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
5 - Ajouter les variables explicatives DRB et BLK **Recommencer les étapes ci-dessus en incluant les variables DRB et BLK** | None | _____no_output_____ | MIT | nbs/06-metriques-et-evaluation-des-modeles-de-regression/06-TP.ipynb | tiombo/TP12 |
Multi-Perceptor VQGAN + CLIP (v.3.2021.11.29)by [@remi_durant](https://twitter.com/remi_durant)Lots drawn from or inspired by other colabs, chief among them is [@jbusted1](https://twitter.com/jbusted1)'s MSE regularized VQGAN + Clip, and [@RiversHaveWings](https://twitter.com/RiversHaveWings) VQGAN + Clip with Z+Quan... | #@title First check what GPU you got and make sure it's a good one.
#@markdown - Tier List: (K80 < T4 < P100 < V100 < A100)
from subprocess import getoutput
!nvidia-smi --query-gpu=name,memory.total,memory.free --format=csv,noheader | _____no_output_____ | MIT | Multi_Perceptor_VQGAN_+_CLIP_[Public].ipynb | keirwilliamsxyz/keirxyz |
Setup | #@title memory footprint support libraries/code
!ln -sf /opt/bin/nvidia-smi /usr/bin/nvidia-smi
!pip install gputil
!pip install psutil
!pip install humanize
import psutil
import humanize
import os
import GPUtil as GPU
#@title Print GPU details
!nvidia-smi
GPUs = GPU.getGPUs()
# XXX: only one GPU on Colab and isn’... | _____no_output_____ | MIT | Multi_Perceptor_VQGAN_+_CLIP_[Public].ipynb | keirwilliamsxyz/keirxyz |
Make some Art! | #@title Set VQGAN Model Save Location
#@markdown It's a lot faster to load model files from google drive than to download them every time you want to use this notebook.
save_vqgan_models_to_drive = True #@param {type: 'boolean'}
download_all = False
vqgan_path_on_google_drive = "/content/drive/MyDrive/Art/Models/VQGAN... | _____no_output_____ | MIT | Multi_Perceptor_VQGAN_+_CLIP_[Public].ipynb | keirwilliamsxyz/keirxyz |
Before generating, the rest of the setup steps must first be executed by pressing **`Runtime > Run All`**. This only needs to be done once. | #@title Do the Run
#@markdown What do you want to see?
text_prompt = 'made of buildings:200 | Ethiopian flags:40 | pollution:30 | 4k:20 | Unreal engine:20 | V-ray:20 | Cryengine:20 | Ray tracing:20 | Photorealistic:20 | Hyper-realistic:20'#@param {type:'string'}
gen_seed = -1#@param {type:'number'}
#@markdown - If you... | _____no_output_____ | MIT | Multi_Perceptor_VQGAN_+_CLIP_[Public].ipynb | keirwilliamsxyz/keirxyz |
Vertex Pipelines: Lightweight Python function-based components, and component I/O Run in Colab View on GitHub Open in Google Cloud Notebooks OverviewThis notebooks shows how to use [the Kubeflow Pipelines (KFP) SDK](https://www.kubeflow.org/docs/components/pip... | import os
# Google Cloud Notebook
if os.path.exists("/opt/deeplearning/metadata/env_version"):
USER_FLAG = "--user"
else:
USER_FLAG = ""
! pip3 install --upgrade google-cloud-aiplatform $USER_FLAG | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Install the latest GA version of *google-cloud-storage* library as well. | ! pip3 install -U google-cloud-storage $USER_FLAG | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Install the latest GA version of *google-cloud-pipeline-components* library as well. | ! pip3 install $USER kfp google-cloud-pipeline-components --upgrade | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Restart the kernelOnce you've installed the additional packages, you need to restart the notebook kernel so it can find the packages. | import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True) | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Check the versions of the packages you installed. The KFP SDK version should be >=1.6. | ! python3 -c "import kfp; print('KFP SDK version: {}'.format(kfp.__version__))"
! python3 -c "import google_cloud_pipeline_components; print('google_cloud_pipeline_components version: {}'.format(google_cloud_pipeline_components.__version__))" | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Before you begin GPU runtimeThis tutorial does not require a GPU runtime. Set up your Google Cloud project**The following steps are required, regardless of your notebook environment.**1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account... | PROJECT_ID = "[your-project-id]" # @param {type:"string"}
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = ! gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:"... | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
RegionYou can also change the `REGION` variable, which is used for operationsthroughout the rest of this notebook. Below are regions supported for Vertex AI. We recommend that you choose the region closest to you.- Americas: `us-central1`- Europe: `europe-west4`- Asia Pacific: `asia-east1`You may not use a multi-regi... | REGION = "us-central1" # @param {type: "string"} | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
TimestampIf you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append the timestamp onto the name of resources you create in this tutorial. | from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S") | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Authenticate your Google Cloud account**If you are using Google Cloud Notebook**, your environment is already authenticated. Skip this step.**If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.**Otherwise**, follow these steps:In the Cloud Cons... | # If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
import os
import sys
# If on Google Cloud Notebook, then don't execute this code
i... | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Create a Cloud Storage bucket**The following steps are required, regardless of your notebook environment.**When you initialize the Vertex SDK for Python, you specify a Cloud Storage staging bucket. The staging bucket is where all the data associated with your dataset and model resources are retained across sessions.Se... | BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket. | ! gsutil mb -l $REGION $BUCKET_NAME | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Finally, validate access to your Cloud Storage bucket by examining its contents: | ! gsutil ls -al $BUCKET_NAME | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Service Account**If you don't know your service account**, try to get your service account using `gcloud` command by executing the second cell below. | SERVICE_ACCOUNT = "[your-service-account]" # @param {type:"string"}
if (
SERVICE_ACCOUNT == ""
or SERVICE_ACCOUNT is None
or SERVICE_ACCOUNT == "[your-service-account]"
):
# Get your GCP project id from gcloud
shell_output = !gcloud auth list 2>/dev/null
SERVICE_ACCOUNT = shell_output[2].strip(... | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Set service account access for Vertex PipelinesRun the following commands to grant your service account access to read and write pipeline artifacts in the bucket that you created in the previous step -- you only need to run these once per service account. | ! gsutil iam ch serviceAccount:{SERVICE_ACCOUNT}:roles/storage.objectCreator $BUCKET_NAME
! gsutil iam ch serviceAccount:{SERVICE_ACCOUNT}:roles/storage.objectViewer $BUCKET_NAME | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Set up variablesNext, set up some variables used throughout the tutorial. Import libraries and define constants | import google.cloud.aiplatform as aip | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Vertex Pipelines constantsSetup up the following constants for Vertex Pipelines: | PIPELINE_ROOT = "{}/pipeline_root/shakespeare".format(BUCKET_NAME) | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Additional imports. | from typing import NamedTuple
import kfp
from kfp.v2 import dsl
from kfp.v2.dsl import (Artifact, Dataset, Input, InputPath, Model, Output,
OutputPath, component) | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Initialize Vertex SDK for PythonInitialize the Vertex SDK for Python for your project and corresponding bucket. | aip.init(project=PROJECT_ID, staging_bucket=BUCKET_NAME) | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Define Python function-based pipeline componentsIn this tutorial, you define function-based components that consume parameters and produce (typed) Artifacts and parameters. Functions can produce Artifacts in three ways:* Accept an output local path using `OutputPath`* Accept an `OutputArtifact` which gives the functio... | @component
def preprocess(
# An input parameter of type string.
message: str,
# Use Output to get a metadata-rich handle to the output artifact
# of type `Dataset`.
output_dataset_one: Output[Dataset],
# A locally accessible filepath for another output artifact of type
# `Dataset`.
outpu... | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Define train componentThe second component definition, `train`, defines as input both an `InputPath` of type `Dataset`, and an `InputArtifact` of type `Dataset` (as well as other parameter inputs). It uses the `NamedTuple` format for function output. As shown, these outputs can be Artifacts as well as parameters.Addi... | @component(
base_image="python:3.9", # Use a different base image.
)
def train(
# An input parameter of type string.
message: str,
# Use InputPath to get a locally accessible path for the input artifact
# of type `Dataset`.
dataset_one_path: InputPath("Dataset"),
# Use InputArtifact to get ... | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Define read_artifact_input componentFinally, you define a small component that takes as input the `generic_artifact` returned by the `train` component function, and reads and prints the Artifact's contents. | @component
def read_artifact_input(
generic: Input[Artifact],
):
with open(generic.path, "r") as input_file:
generic_contents = input_file.read()
print(f"generic contents: {generic_contents}") | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Define a pipeline that uses your components and the ImporterNext, define a pipeline that uses the components that were built in the previous section, and also shows the use of the `kfp.dsl.importer`.This example uses the `importer` to create, in this case, a `Dataset` artifact from an existing URI.Note that the `train... | @dsl.pipeline(
# Default pipeline root. You can override it when submitting the pipeline.
pipeline_root=PIPELINE_ROOT,
# A name for the pipeline. Use to determine the pipeline Context.
name="metadata-pipeline-v2",
)
def pipeline(message: str):
importer = kfp.dsl.importer(
artifact_uri="gs://... | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Compile the pipelineNext, compile the pipeline. | from kfp.v2 import compiler # noqa: F811
compiler.Compiler().compile(
pipeline_func=pipeline, package_path="lightweight_pipeline.json".replace(" ", "_")
) | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Run the pipelineNext, run the pipeline. | DISPLAY_NAME = "shakespeare_" + TIMESTAMP
job = aip.PipelineJob(
display_name=DISPLAY_NAME,
template_path="lightweight_pipeline.json".replace(" ", "_"),
pipeline_root=PIPELINE_ROOT,
parameter_values={"message": "Hello, World"},
)
job.run() | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
Click on the generated link to see your run in the Cloud Console.<!-- It should look something like this as it is running: -->In the UI, many of the pipeline DAG nodes will expand or collapse when you click on them. Here is a partially-expanded view of the DAG (click image to see larger version). Cleaning upTo clean u... | delete_dataset = True
delete_pipeline = True
delete_model = True
delete_endpoint = True
delete_batchjob = True
delete_customjob = True
delete_hptjob = True
delete_bucket = True
try:
if delete_model and "DISPLAY_NAME" in globals():
models = aip.Model.list(
filter=f"display_name={DISPLAY_NAME}", ... | _____no_output_____ | Apache-2.0 | notebooks/official/pipelines/lightweight_functions_component_io_kfp.ipynb | diemtvu/vertex-ai-samples |
WikiPathways and py4cytoscape Yihang Xin and Alex Pico 2020-11-10 WikiPathways is a well-known repository for biological pathways that provides unique tools to the research community for content creation, editing and utilization [@Pico2008].Python is an interpreted, high-level and general-purpose programming language.... | %%capture
!python3 -m pip install python-igraph requests pandas networkx
!python3 -m pip install py4cytoscape | _____no_output_____ | CC0-1.0 | for-scripters/Python/wikiPathways-and-py4cytoscape.ipynb | kozo2/cytoscape-automation |
Prerequisites In addition to this package (py4cytoscape latest version 0.0.7), you will need:* Latest version of Cytoscape, which can be downloaded from https://cytoscape.org/download.html. Simply follow the installation instructions on screen.* Complete installation wizard* Launch CytoscapeFor this vignette, you’ll a... | import os
import sys
import requests
import pandas as pd
from lxml import etree as ET
from collections import OrderedDict
import py4cytoscape as p4c
# Check Version
p4c.cytoscape_version_info() | _____no_output_____ | CC0-1.0 | for-scripters/Python/wikiPathways-and-py4cytoscape.ipynb | kozo2/cytoscape-automation |
Working togetherOk, with all of these components loaded and launched, you can now perform some nifty sequences. For example, search for a pathway based on a keyword search and then load it into Cytoscape. | def find_pathways_by_text(query, species):
base_iri = 'http://webservice.wikipathways.org/'
request_params = {'query':query, 'species':species}
response = requests.get(base_iri + 'findPathwaysByText', params=request_params)
return response
response = find_pathways_by_text("colon cancer", "Homo sapiens")... | _____no_output_____ | CC0-1.0 | for-scripters/Python/wikiPathways-and-py4cytoscape.ipynb | kozo2/cytoscape-automation |
We have a list of human pathways that mention “Colon Cancer”. The results include lots of information, so let’s get a unique list of just the WPIDs. | unique_id = list(OrderedDict.fromkeys(df["id"]))
unique_id[0] | _____no_output_____ | CC0-1.0 | for-scripters/Python/wikiPathways-and-py4cytoscape.ipynb | kozo2/cytoscape-automation |
Let’s import the first one of these into Cytoscape! | cmd_list = ['wikipathways','import-as-pathway','id="',unique_id[0],'"']
cmd = " ".join(cmd_list)
p4c.commands.commands_get(cmd) | _____no_output_____ | CC0-1.0 | for-scripters/Python/wikiPathways-and-py4cytoscape.ipynb | kozo2/cytoscape-automation |
Once in Cytoscape, you can load data, apply visual style mappings, perform analyses, and export images and data formats. See py4cytoscape package for details. From networks to pathwaysIf you are already with with networks and data in Cytoscape, you may end up focusing on one or few particular genes, proteins or metabo... | p4c.session.open_session()
net_data = p4c.tables.get_table_columns(columns=['name','degree.layout','COMMON'])
max_gene = net_data[net_data["degree.layout"] == net_data["degree.layout"].max()]
max_gene | _____no_output_____ | CC0-1.0 | for-scripters/Python/wikiPathways-and-py4cytoscape.ipynb | kozo2/cytoscape-automation |
Great. It looks like MCM1 has the larget number of connections (18) in this network. Let’s use it’s identifier (YMR043W) to query WikiPathways to learn more about the gene and its biological role, and load it into Cytoscape.Pro-tip: We need to know the datasource that provides a given identifier. In this case, it’s sor... | def find_pathways_by_xref(ids, codes):
base_iri = 'http://webservice.wikipathways.org/'
request_params = {'ids':ids, 'codes':codes}
response = requests.get(base_iri + 'findPathwaysByXref', params=request_params)
return response
response = find_pathways_by_xref('YMR043W','En')
mcm1_pathways = find_pathwa... | _____no_output_____ | CC0-1.0 | for-scripters/Python/wikiPathways-and-py4cytoscape.ipynb | kozo2/cytoscape-automation |
And we can easily select the MCM1 node by name in the newly imported pathway to help see where exactly it plays its role. | p4c.network_selection.select_nodes(['Mcm1'], by_col='name') | _____no_output_____ | CC0-1.0 | for-scripters/Python/wikiPathways-and-py4cytoscape.ipynb | kozo2/cytoscape-automation |
Plotting with Matplotlib What is `matplotlib`?* `matplotlib` is a 2D plotting library for Python* It provides quick way to visualize data from Python* It comes with a set plots* We can import its functions through the command```Pythonimport matplotlib.pyplot as plt``` | import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline | _____no_output_____ | Apache-2.0 | 02-plotting-with-matplotlib.ipynb | theed-ml/notebooks |
Basic plotsYour goal is to plot the cosine and the sine functions on the same plot, using the default `matplotlib` settings. Generating the data | x = np.linspace(-np.pi, np.pi, 256, endpoint=True)
c, s = np.cos(x), np.sin(x) | _____no_output_____ | Apache-2.0 | 02-plotting-with-matplotlib.ipynb | theed-ml/notebooks |
where, * x is a vector with 256 values ranging from $-\pi$ to $\pi$ included * c and s are vectors with the cosine and the sine values of X | plt.plot(x, c)
plt.plot(x, s)
plt.show() | _____no_output_____ | Apache-2.0 | 02-plotting-with-matplotlib.ipynb | theed-ml/notebooks |
Rather than creating a plot with default size, we want to specify:* the size of the figure* the colors and type of the lines* the limits of the axes | # Create a figure of size 8x6 inches, 80 dots per inch
plt.figure(figsize=(8, 6), dpi=80)
# Create a new subplot from a grid of 1x1
plt.subplot(1, 1, 1)
# Plot cosine with a blue continuous line of width 1 (pixels)
plt.plot(x, c, color="blue", linewidth=2.5, linestyle="-")
# Plot sine with a green dotted line of wid... | _____no_output_____ | Apache-2.0 | 02-plotting-with-matplotlib.ipynb | theed-ml/notebooks |
Changing colors and line widths* We can to: - make the figure more horizontal - change the color of the lines to blue and red - have slighty thicker lines | plt.figure(figsize=(10, 6), dpi=80)
plt.plot(x, c, color="blue", linewidth=2.5, linestyle="-")
plt.plot(x, s, color="red", linewidth=2.5, linestyle="solid") | _____no_output_____ | Apache-2.0 | 02-plotting-with-matplotlib.ipynb | theed-ml/notebooks |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.