+
🔗 GLMP (Genome Logic Modeling Project) Connection
-
This analysis is based on the comprehensive biological dataset from the Genome Logic Modeling Project (GLMP), which contains 50+ analyzed biological processes across multiple organisms and systems.
+
This analysis is based on the comprehensive biological dataset from the Genome Logic Modeling Project (GLMP), which contains 297+ analyzed biological processes across multiple organisms and systems.
GLMP Resources:
-
The GLMP represents the first successful application of the Programming Framework to biological processes, demonstrating how biological systems function as sophisticated computational programs with complex regulatory logic, decision trees, and feedback mechanisms.
+
The GLMP represents the most comprehensive biological computing system analysis ever created, demonstrating how biological systems function as sophisticated computational programs with complex regulatory logic, decision trees, and feedback mechanisms.
This document presents representative biological processes analyzed using the Programming Framework methodology. Each process is represented as a computational flowchart with standardized color coding: Red for triggers/inputs, Yellow for structures/objects, Green for processing/operations, Blue for intermediates/states, and Violet for products/outputs. Yellow nodes use black text for optimal readability, while all other colors use white text.
diff --git a/chemistry-database-table.html b/chemistry-database-table.html
deleted file mode 100644
index fc8b1cf8d51249d760b5069f93c1afcd500ccb53..0000000000000000000000000000000000000000
--- a/chemistry-database-table.html
+++ /dev/null
@@ -1,545 +0,0 @@
-
-
-
-
-
-
Chemistry Processes Database
-
-
-
-
-
-
-
-
Loading chemistry processes database...
-
Fetching process data from metadata.json
-
-
-
-
❌ Error Loading Data
-
Could not fetch chemistry processes metadata. Please check your connection and try again.
-
-
-
-
-
-
-
-
-
-
-
-
- | Process Name ↕ |
- Subcategory ↕ |
- Complexity ↕ |
- Nodes ↕ |
- Edges ↕ |
- AND Gates ↕ |
- OR Gates ↕ |
-
-
-
-
-
-
-
-
-
-
📈 Database Analysis
-
-
-
-
-
-
-
🎨 Color Scheme (5-Color System)
-
-
-
-
- Red
- Triggers & Inputs
-
-
-
-
-
- Yellow
- Structures & Objects
-
-
-
-
-
- Green
- Processing & Operations
-
-
-
-
-
- Blue
- Intermediates & States
-
-
-
-
-
- Violet
- Products & Outputs
-
-
-
-
-
-
-
-
-
-
\ No newline at end of file
diff --git a/chemistry_examples.html b/chemistry_examples.html
deleted file mode 100644
index 2f2d1831c01bb749979de08bbffb29e7188b2af8..0000000000000000000000000000000000000000
--- a/chemistry_examples.html
+++ /dev/null
@@ -1,609 +0,0 @@
-
-
-
-
-
-
Chemistry Examples - Programming Framework Analysis
-
-
-
-
-
-
-
Chemistry Examples - Programming Framework Analysis
-
-
This document presents chemistry processes analyzed using the Programming Framework methodology. Each process is represented as a computational flowchart with standardized color coding: Red for triggers/inputs, Yellow for structures/objects, Green for processing/operations, Blue for intermediates/states, and Violet for products/outputs. Yellow nodes use black text for optimal readability, while all other colors use white text.
-
-
1. Catalytic Hydrogenation Process
-
-
-
2. Polymerization Process
-
-
-graph TD
- %% Initial Setup
- %% Input Conditions
- A2[Monomers] --> B2[Monomer Analysis]
- C2[Initiator] --> D2[Initiator Preparation]
- E2[Solvent/Medium] --> F2[Reaction Medium]
- %% Reaction Setup
- B2 --> G2[Monomer Purity Check]
- D2 --> H2[Initiator Activation]
- F2 --> I2[Medium Preparation]
- %% Initiation
- G2 --> J2[Monomer Solution]
- H2 --> K2[Radical Formation]
- I2 --> L2[Temperature Control]
- %% Polymerization Initiation
- J2 --> M2[Monomer Concentration]
- K2 --> N2[Initiator Decomposition]
- L2 --> O2[Reaction Conditions]
- %% Propagation
- M2 --> P2[Radical Addition]
- N2 --> Q2[Chain Initiation]
- O2 --> R2[Chain Growth]
- %% Chain Growth
- P2 --> S2[Monomer Addition]
- Q2 --> T2[Active Chain End]
- R2 --> U2[Polymer Chain]
- %% Termination
- S2 --> V2[Chain Propagation]
- T2 --> W2[Chain Transfer]
- U2 --> X2[Chain Termination]
- %% Final Products
- V2 --> Y2[Polymer Formation]
- W2 --> Z2[Molecular Weight Control]
- X2 --> AA2[Reaction Quenching]
- %% Process Control
- Y2 --> BB2[Polymer Characterization]
- Z2 --> CC2[Molecular Weight Analysis]
- AA2 --> DD2[Product Isolation]
- %% Output
- BB2 --> EE2[Polymerization Complete]
- CC2 --> FF2[Quality Control]
- DD2 --> GG2[Product Recovery]
- %% Styling - Chemistry Color Scheme
- %% Styling - Biological Color Scheme
- style A2 fill:#ff6b6b,color:#fff
- style C2 fill:#ff6b6b,color:#fff
- style E2 fill:#ff6b6b,color:#fff
- style G2 fill:#ffd43b,color:#000
- style H2 fill:#ffd43b,color:#000
- style I2 fill:#ffd43b,color:#000
- style J2 fill:#ffd43b,color:#000
- style K2 fill:#ffd43b,color:#000
- style L2 fill:#ffd43b,color:#000
- style M2 fill:#51cf66,color:#fff
- style N2 fill:#51cf66,color:#fff
- style O2 fill:#51cf66,color:#fff
- style P2 fill:#51cf66,color:#fff
- style Q2 fill:#51cf66,color:#fff
- style R2 fill:#51cf66,color:#fff
- style S2 fill:#51cf66,color:#fff
- style T2 fill:#51cf66,color:#fff
- style U2 fill:#51cf66,color:#fff
- style V2 fill:#51cf66,color:#fff
- style W2 fill:#51cf66,color:#fff
- style X2 fill:#51cf66,color:#fff
- style B2 fill:#74c0fc,color:#fff
- style D2 fill:#74c0fc,color:#fff
- style F2 fill:#74c0fc,color:#fff
- style Y2 fill:#74c0fc,color:#fff
- style Z2 fill:#74c0fc,color:#fff
- style AA2 fill:#74c0fc,color:#fff
- style BB2 fill:#74c0fc,color:#fff
- style CC2 fill:#74c0fc,color:#fff
- style DD2 fill:#74c0fc,color:#fff
- style EE2 fill:#b197fc,color:#fff
- style FF2 fill:#b197fc,color:#fff
- style GG2 fill:#b197fc,color:#fff
-
-
-
-
- Reactants & Initiators
-
-
- Reaction Vessels & Equipment
-
-
- Polymerization Reactions
-
-
- Intermediates
-
-
- Products
-
-
-
-
- Figure 2. Polymerization Process. This chemistry process visualization demonstrates polymer synthesis mechanisms. The flowchart shows monomer inputs, reaction vessels and equipment, polymerization reactions, intermediate species, and final polymer products.
-
-
-
-
3. Acid-Base Titration Process
-
-
-graph TD
- %% Initial Setup
- %% Input Conditions
- A3[Acid Solution] --> B3[Acid Analysis]
- C3[Base Solution] --> D3[Base Analysis]
- E3[Indicator] --> F3[Indicator Selection]
- %% Solution Preparation
- B3 --> G3[Acid Concentration]
- D3 --> H3[Base Concentration]
- F3 --> I3[Indicator Preparation]
- %% Titration Setup
- G3 --> J3[Acid in Burette]
- H3 --> K3[Base in Flask]
- I3 --> L3[Indicator Addition]
- %% Titration Process
- J3 --> M3[Initial pH Measurement]
- K3 --> N3[Base Volume Measurement]
- L3 --> O3[Color Change Detection]
- %% Acid-Base Reaction
- M3 --> P3[Acid Addition]
- N3 --> Q3[Base Consumption]
- O3 --> R3[Equivalence Point]
- %% pH Changes
- P3 --> S3[H⁺ + OH⁻ → H₂O]
- Q3 --> T3[Neutralization Reaction]
- R3 --> U3[Stoichiometric Point]
- %% Endpoint Detection
- S3 --> V3[pH Monitoring]
- T3 --> W3[Conductivity Changes]
- U3 --> X3[Indicator Color Change]
- %% Final Results
- V3 --> Y3[Equivalence Point pH]
- W3 --> Z3[Conductivity Minimum]
- X3 --> AA3[Endpoint Detection]
- %% Calculations
- Y3 --> BB3[Concentration Calculation]
- Z3 --> CC3[Volume Measurement]
- AA3 --> DD3[Stoichiometric Analysis]
- %% Output
- BB3 --> EE3[Titration Complete]
- CC3 --> FF3[Concentration Determined]
- DD3 --> GG3[Analysis Results]
- %% Styling - Chemistry Color Scheme
- %% Styling - Biological Color Scheme
- style A3 fill:#ff6b6b,color:#fff
- style C3 fill:#ff6b6b,color:#fff
- style E3 fill:#ff6b6b,color:#fff
- style G3 fill:#ffd43b,color:#000
- style H3 fill:#ffd43b,color:#000
- style I3 fill:#ffd43b,color:#000
- style J3 fill:#ffd43b,color:#000
- style K3 fill:#ffd43b,color:#000
- style L3 fill:#ffd43b,color:#000
- style M3 fill:#51cf66,color:#fff
- style N3 fill:#51cf66,color:#fff
- style O3 fill:#51cf66,color:#fff
- style P3 fill:#51cf66,color:#fff
- style Q3 fill:#51cf66,color:#fff
- style R3 fill:#51cf66,color:#fff
- style S3 fill:#51cf66,color:#fff
- style T3 fill:#51cf66,color:#fff
- style U3 fill:#51cf66,color:#fff
- style V3 fill:#51cf66,color:#fff
- style W3 fill:#51cf66,color:#fff
- style X3 fill:#51cf66,color:#fff
- style B3 fill:#74c0fc,color:#fff
- style D3 fill:#74c0fc,color:#fff
- style F3 fill:#74c0fc,color:#fff
- style Y3 fill:#74c0fc,color:#fff
- style Z3 fill:#74c0fc,color:#fff
- style AA3 fill:#74c0fc,color:#fff
- style BB3 fill:#74c0fc,color:#fff
- style CC3 fill:#74c0fc,color:#fff
- style DD3 fill:#74c0fc,color:#fff
- style EE3 fill:#b197fc,color:#fff
- style FF3 fill:#b197fc,color:#fff
- style GG3 fill:#b197fc,color:#fff
-
-
-
-
- Reactants & Indicators
-
-
- Titration Equipment
-
-
- Neutralization Reactions
-
-
- Intermediates
-
-
- Products
-
-
-
-
- Figure 3. Acid-Base Titration Process. This chemistry process visualization demonstrates analytical chemistry procedures. The flowchart shows reactant inputs, titration equipment, neutralization reactions, intermediate measurements, and final analytical results.
-
-
-
-
4. Electrochemical Cell Process
-
-
-graph TD
- %% Initial Setup
- %% Input Conditions
- A4[Anode Material] --> B4[Anode Analysis]
- C4[Cathode Material] --> D4[Cathode Analysis]
- E4[Electrolyte] --> F4[Electrolyte Preparation]
- %% Cell Setup
- B4 --> G4[Anode Preparation]
- D4 --> H4[Cathode Preparation]
- F4 --> I4[Electrolyte Solution]
- %% Electrochemical Setup
- G4 --> J4[Anode Oxidation]
- H4 --> K4[Cathode Reduction]
- I4 --> L4[Ion Conduction]
- %% Redox Reactions
- J4 --> M4[Electron Release]
- K4 --> N4[Electron Acceptance]
- L4 --> O4[Ion Transport]
- %% Current Flow
- M4 --> P4[Oxidation Half-Reaction]
- N4 --> Q4[Reduction Half-Reaction]
- O4 --> R4[Salt Bridge]
- %% Cell Operation
- P4 --> S4[Anode → Cathode]
- Q4 --> T4[Electron Flow]
- R4 --> U4[Ion Migration]
- %% Energy Production
- S4 --> V4[Cell Potential]
- T4 --> W4[Current Generation]
- U4 --> X4[Charge Balance]
- %% Performance
- V4 --> Y4[Voltage Measurement]
- W4 --> Z4[Current Measurement]
- X4 --> AA4[Efficiency Calculation]
- %% Output
- Y4 --> BB4[Cell Performance]
- Z4 --> CC4[Power Output]
- AA4 --> DD4[Energy Conversion]
- %% Final Results
- BB4 --> EE4[Electrochemical Cell Active]
- CC4 --> FF4[Electrical Energy Produced]
- DD4 --> GG4[Conversion Efficiency]
- %% Styling - Chemistry Color Scheme
- %% Styling - Biological Color Scheme
- style A4 fill:#ff6b6b,color:#fff
- style C4 fill:#ff6b6b,color:#fff
- style E4 fill:#ff6b6b,color:#fff
- style G4 fill:#ffd43b,color:#000
- style H4 fill:#ffd43b,color:#000
- style I4 fill:#ffd43b,color:#000
- style J4 fill:#ffd43b,color:#000
- style K4 fill:#ffd43b,color:#000
- style L4 fill:#ffd43b,color:#000
- style M4 fill:#51cf66,color:#fff
- style N4 fill:#51cf66,color:#fff
- style O4 fill:#51cf66,color:#fff
- style P4 fill:#51cf66,color:#fff
- style Q4 fill:#51cf66,color:#fff
- style R4 fill:#51cf66,color:#fff
- style S4 fill:#51cf66,color:#fff
- style T4 fill:#51cf66,color:#fff
- style U4 fill:#51cf66,color:#fff
- style V4 fill:#51cf66,color:#fff
- style W4 fill:#51cf66,color:#fff
- style X4 fill:#51cf66,color:#fff
- style B4 fill:#74c0fc,color:#fff
- style D4 fill:#74c0fc,color:#fff
- style F4 fill:#74c0fc,color:#fff
- style Y4 fill:#74c0fc,color:#fff
- style Z4 fill:#74c0fc,color:#fff
- style AA4 fill:#74c0fc,color:#fff
- style BB4 fill:#74c0fc,color:#fff
- style CC4 fill:#74c0fc,color:#fff
- style DD4 fill:#74c0fc,color:#fff
- style EE4 fill:#b197fc,color:#fff
- style FF4 fill:#b197fc,color:#fff
- style GG4 fill:#b197fc,color:#fff
-
-
-
-
- Electrodes & Electrolyte
-
-
- Cell Components
-
-
- Redox Reactions
-
-
- Intermediates
-
-
- Products
-
-
-
-
- Figure 4. Electrochemical Cell Process. This chemistry process visualization demonstrates electrochemical energy conversion. The flowchart shows electrode inputs, cell components, redox reactions, intermediate processes, and final electrical energy output.
-
-
-
-
5. Distillation Process
-
-
-graph TD
- %% Initial Setup
- %% Input Conditions
- A5[Mixture] --> B5[Mixture Analysis]
- C5[Heat Source] --> D5[Heat Supply]
- E5[Distillation Apparatus] --> F5[Equipment Setup]
- %% Process Setup
- B5 --> G5[Component Analysis]
- D5 --> H5[Temperature Control]
- F5 --> I5[Apparatus Assembly]
- %% Heating Process
- G5 --> J5[Mixture Loading]
- H5 --> K5[Heat Application]
- I5 --> L5[Vapor Formation]
- %% Vaporization
- J5 --> M5[Component Separation]
- K5 --> N5[Boiling Point Differences]
- L5 --> O5[Vapor Phase]
- %% Distillation
- M5 --> P5[Fractional Distillation]
- N5 --> Q5[Temperature Gradient]
- O5 --> R5[Condensation]
- %% Separation
- P5 --> S5[Component Collection]
- Q5 --> T5[Fraction Separation]
- R5 --> U5[Liquid Recovery]
- %% Product Collection
- S5 --> V5[High Boiling Point]
- T5 --> W5[Medium Boiling Point]
- U5 --> X5[Low Boiling Point]
- %% Process Control
- V5 --> Y5[Fraction Analysis]
- W5 --> Z5[Purity Check]
- X5 --> AA5[Yield Calculation]
- %% Output
- Y5 --> BB5[Distillation Complete]
- Z5 --> CC5[Component Separation]
- AA5 --> DD5[Process Efficiency]
- %% Final Results
- BB5 --> EE5[Purified Components]
- CC5 --> FF5[Separation Achieved]
- DD5 --> GG5[Process Optimization]
- %% Styling - Chemistry Color Scheme
- %% Styling - Biological Color Scheme
- style A5 fill:#ff6b6b,color:#fff
- style C5 fill:#ff6b6b,color:#fff
- style E5 fill:#ff6b6b,color:#fff
- style G5 fill:#ffd43b,color:#000
- style H5 fill:#ffd43b,color:#000
- style I5 fill:#ffd43b,color:#000
- style J5 fill:#ffd43b,color:#000
- style K5 fill:#ffd43b,color:#000
- style L5 fill:#ffd43b,color:#000
- style M5 fill:#51cf66,color:#fff
- style N5 fill:#51cf66,color:#fff
- style O5 fill:#51cf66,color:#fff
- style P5 fill:#51cf66,color:#fff
- style Q5 fill:#51cf66,color:#fff
- style R5 fill:#51cf66,color:#fff
- style S5 fill:#51cf66,color:#fff
- style T5 fill:#51cf66,color:#fff
- style U5 fill:#51cf66,color:#fff
- style V5 fill:#51cf66,color:#fff
- style W5 fill:#51cf66,color:#fff
- style X5 fill:#51cf66,color:#fff
- style B5 fill:#74c0fc,color:#fff
- style D5 fill:#74c0fc,color:#fff
- style F5 fill:#74c0fc,color:#fff
- style Y5 fill:#74c0fc,color:#fff
- style Z5 fill:#74c0fc,color:#fff
- style AA5 fill:#74c0fc,color:#fff
- style BB5 fill:#74c0fc,color:#fff
- style CC5 fill:#74c0fc,color:#fff
- style DD5 fill:#74c0fc,color:#fff
- style EE5 fill:#b197fc,color:#fff
- style FF5 fill:#b197fc,color:#fff
- style GG5 fill:#b197fc,color:#fff
-
-
-
-
- Mixture & Heat Source
-
-
- Distillation Equipment
-
-
- Phase Separation Processes
-
-
- Intermediates
-
-
- Products
-
-
-
-
- Figure 5. Distillation Process. This chemistry process visualization demonstrates separation techniques. The flowchart shows mixture inputs, distillation equipment, phase separation processes, intermediate fractions, and final purified components.
-
-
-
-
Generated using the Programming Framework methodology
-
-
This collection demonstrates the computational nature of chemical processes and systems
-
-
Each flowchart preserves maximum detail through optimized Mermaid configuration
-
-
-
diff --git a/computer-science-database-table.html b/computer-science-database-table.html
deleted file mode 100644
index 2657a219172b2a05f43c0ca6b4cf5f8952faf508..0000000000000000000000000000000000000000
--- a/computer-science-database-table.html
+++ /dev/null
@@ -1,543 +0,0 @@
-
-
-
-
-
-
Computer Science Processes Database
-
-
-
-
-
-
-
-
Loading computer_science processes database...
-
Fetching process data from metadata.json
-
-
-
-
❌ Error Loading Data
-
Could not fetch computer_science processes metadata. Please check your connection and try again.
-
-
-
-
-
-
-
-
-
-
-
-
- | Process Name ↕ |
- Subcategory ↕ |
- Complexity ↕ |
- Nodes ↕ |
- Edges ↕ |
- AND Gates ↕ |
- OR Gates ↕ |
-
-
-
-
-
-
-
-
-
-
📈 Database Analysis
-
-
-
-
-
-
-
🎨 Color Scheme (5-Color System)
-
-
-
-
- Red
- Triggers & Inputs
-
-
-
-
-
- Yellow
- Structures & Objects
-
-
-
-
-
- Green
- Processing & Operations
-
-
-
-
-
- Blue
- Intermediates & States
-
-
-
-
-
- Violet
- Products & Outputs
-
-
-
-
-
-
-
-
-
-
\ No newline at end of file
diff --git a/data/aristotle-syllogistic-figure-2.mmd b/data/aristotle-syllogistic-figure-2.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..de55244a987b3a9277ca8adef3d91d09c6060c4e
--- /dev/null
+++ b/data/aristotle-syllogistic-figure-2.mmd
@@ -0,0 +1,30 @@
+graph TD
+ DefAEIO("DefAEIO Categorical forms")
+ DefFig("DefFig Figures 1,2,3")
+ Econv("Econv E-conversion")
+ Barbara("Barbara Barbara")
+ Celarent("Celarent Celarent")
+ Ferio("Ferio Ferio")
+ Cesare("Cesare Cesare")
+ Camestres("Camestres Camestres")
+ Festino("Festino Festino")
+ Baroco("Baroco Baroco")
+ DefAEIO --> Barbara
+ DefFig --> Barbara
+ DefAEIO --> Celarent
+ DefFig --> Celarent
+ DefAEIO --> Ferio
+ DefFig --> Ferio
+ Econv --> Cesare
+ Celarent --> Cesare
+ Econv --> Camestres
+ Celarent --> Camestres
+ Econv --> Festino
+ Ferio --> Festino
+ Barbara --> Baroco
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class Econv,Barbara,Celarent,Ferio axiom
+ class DefAEIO,DefFig definition
+ class Cesare,Camestres,Festino,Baroco theorem
\ No newline at end of file
diff --git a/data/aristotle-syllogistic-figure-3.mmd b/data/aristotle-syllogistic-figure-3.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..d5191ada404b7042ecaf16a16147c120c7850405
--- /dev/null
+++ b/data/aristotle-syllogistic-figure-3.mmd
@@ -0,0 +1,37 @@
+graph TD
+ DefAEIO("DefAEIO Categorical forms")
+ DefFig("DefFig Figures 1,2,3")
+ Iconv("Iconv I-conversion")
+ Aconv("Aconv A-conversion")
+ Barbara("Barbara Barbara")
+ Darii("Darii Darii")
+ Ferio("Ferio Ferio")
+ Darapti("Darapti Darapti")
+ Felapton("Felapton Felapton")
+ Disamis("Disamis Disamis")
+ Datisi("Datisi Datisi")
+ Bocardo("Bocardo Bocardo")
+ Ferison("Ferison Ferison")
+ DefAEIO --> Barbara
+ DefFig --> Barbara
+ DefAEIO --> Darii
+ DefFig --> Darii
+ DefAEIO --> Ferio
+ DefFig --> Ferio
+ Aconv --> Darapti
+ Darii --> Darapti
+ Aconv --> Felapton
+ Ferio --> Felapton
+ Iconv --> Disamis
+ Darii --> Disamis
+ Aconv --> Datisi
+ Darii --> Datisi
+ Barbara --> Bocardo
+ Aconv --> Ferison
+ Ferio --> Ferison
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class Iconv,Aconv,Barbara,Darii,Ferio axiom
+ class DefAEIO,DefFig definition
+ class Darapti,Felapton,Disamis,Datisi,Bocardo,Ferison theorem
\ No newline at end of file
diff --git a/data/aristotle-syllogistic-foundations-perfect.mmd b/data/aristotle-syllogistic-foundations-perfect.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..257a7293866f272e8800de7a029172aa3fb7ebaf
--- /dev/null
+++ b/data/aristotle-syllogistic-foundations-perfect.mmd
@@ -0,0 +1,23 @@
+graph TD
+ DefAEIO("DefAEIO Categorical forms")
+ DefFig("DefFig Figures 1,2,3")
+ Econv("Econv E-conversion")
+ Iconv("Iconv I-conversion")
+ Aconv("Aconv A-conversion")
+ Barbara("Barbara Barbara")
+ Celarent("Celarent Celarent")
+ Darii("Darii Darii")
+ Ferio("Ferio Ferio")
+ DefAEIO --> Barbara
+ DefFig --> Barbara
+ DefAEIO --> Celarent
+ DefFig --> Celarent
+ DefAEIO --> Darii
+ DefFig --> Darii
+ DefAEIO --> Ferio
+ DefFig --> Ferio
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class Econv,Iconv,Aconv,Barbara,Celarent,Darii,Ferio axiom
+ class DefAEIO,DefFig definition
\ No newline at end of file
diff --git a/data/aristotle-syllogistic.json b/data/aristotle-syllogistic.json
new file mode 100644
index 0000000000000000000000000000000000000000..f25bc81de8e9b53f04e836dcff8e53a9ce41ceab
--- /dev/null
+++ b/data/aristotle-syllogistic.json
@@ -0,0 +1,326 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "aristotle-syllogistic",
+ "name": "Aristotle Syllogistic Logic",
+ "subject": "logic",
+ "variant": "classical",
+ "description": "Aristotelian categorical syllogistic. Four perfect syllogisms (Barbara, Celarent, Darii, Ferio), three conversion rules, and ten imperfect syllogisms reduced to the perfect ones. Based on Prior Analytics.",
+ "structure": {
+ "axioms": 7,
+ "definitions": 2,
+ "theorems": 10
+ }
+ },
+ "metadata": {
+ "created": "2026-03-15",
+ "lastUpdated": "2026-03-15",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Aristotle Syllogistic Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Aristotle",
+ "syllogism",
+ "categorical logic",
+ "Prior Analytics",
+ "Barbara",
+ "Celarent"
+ ]
+ },
+ "sources": [
+ {
+ "id": "aristotle",
+ "type": "primary",
+ "authors": "Aristotle",
+ "title": "Prior Analytics",
+ "year": "c. 350 BCE",
+ "notes": "Syllogistic theory"
+ },
+ {
+ "id": "stanford",
+ "type": "secondary",
+ "authors": "Smith, R.",
+ "title": "Aristotle's Logic",
+ "url": "https://plato.stanford.edu/entries/aristotle-logic/",
+ "notes": "Stanford Encyclopedia"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "DefAEIO",
+ "type": "definition",
+ "label": "A, E, I, O: All/No/Some/Some-not",
+ "shortLabel": "DefAEIO",
+ "short": "Categorical forms",
+ "colorClass": "definition"
+ },
+ {
+ "id": "DefFig",
+ "type": "definition",
+ "label": "Three figures: middle term position",
+ "shortLabel": "DefFig",
+ "short": "Figures 1,2,3",
+ "colorClass": "definition"
+ },
+ {
+ "id": "Econv",
+ "type": "axiom",
+ "label": "Eab implies Eba (No M is P implies No P is M)",
+ "shortLabel": "Econv",
+ "short": "E-conversion",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "Iconv",
+ "type": "axiom",
+ "label": "Iab implies Iba (Some M is P implies Some P is M)",
+ "shortLabel": "Iconv",
+ "short": "I-conversion",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "Aconv",
+ "type": "axiom",
+ "label": "Aab implies Iba (All M is P implies Some P is M)",
+ "shortLabel": "Aconv",
+ "short": "A-conversion",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "Barbara",
+ "type": "axiom",
+ "label": "AAA-1: All M is P, All S is M therefore All S is P",
+ "shortLabel": "Barbara",
+ "short": "Barbara",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "Celarent",
+ "type": "axiom",
+ "label": "EAE-1: No M is P, All S is M therefore No S is P",
+ "shortLabel": "Celarent",
+ "short": "Celarent",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "Darii",
+ "type": "axiom",
+ "label": "AII-1: All M is P, Some S is M therefore Some S is P",
+ "shortLabel": "Darii",
+ "short": "Darii",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "Ferio",
+ "type": "axiom",
+ "label": "EIO-1: No M is P, Some S is M therefore Some S is not P",
+ "shortLabel": "Ferio",
+ "short": "Ferio",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "Cesare",
+ "type": "theorem",
+ "label": "EAE-2: No P is M, All S is M therefore No S is P",
+ "shortLabel": "Cesare",
+ "short": "Cesare",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Camestres",
+ "type": "theorem",
+ "label": "AEE-2: All P is M, No S is M therefore No S is P",
+ "shortLabel": "Camestres",
+ "short": "Camestres",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Festino",
+ "type": "theorem",
+ "label": "EIO-2: No P is M, Some S is M therefore Some S is not P",
+ "shortLabel": "Festino",
+ "short": "Festino",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Baroco",
+ "type": "theorem",
+ "label": "AOO-2: All P is M, Some S is not M therefore Some S is not P",
+ "shortLabel": "Baroco",
+ "short": "Baroco",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Darapti",
+ "type": "theorem",
+ "label": "AAI-3: All M is P, All M is S therefore Some S is P",
+ "shortLabel": "Darapti",
+ "short": "Darapti",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Felapton",
+ "type": "theorem",
+ "label": "EAO-3: No M is P, All M is S therefore Some S is not P",
+ "shortLabel": "Felapton",
+ "short": "Felapton",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Disamis",
+ "type": "theorem",
+ "label": "IAI-3: Some M is P, All M is S therefore Some S is P",
+ "shortLabel": "Disamis",
+ "short": "Disamis",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Datisi",
+ "type": "theorem",
+ "label": "AII-3: All M is P, Some M is S therefore Some S is P",
+ "shortLabel": "Datisi",
+ "short": "Datisi",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Bocardo",
+ "type": "theorem",
+ "label": "OAO-3: Some M is not P, All M is S therefore Some S is not P",
+ "shortLabel": "Bocardo",
+ "short": "Bocardo",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Ferison",
+ "type": "theorem",
+ "label": "EIO-3: No M is P, Some M is S therefore Some S is not P",
+ "shortLabel": "Ferison",
+ "short": "Ferison",
+ "colorClass": "theorem"
+ }
+ ],
+ "edges": [
+ {
+ "from": "DefAEIO",
+ "to": "Barbara"
+ },
+ {
+ "from": "DefFig",
+ "to": "Barbara"
+ },
+ {
+ "from": "DefAEIO",
+ "to": "Celarent"
+ },
+ {
+ "from": "DefFig",
+ "to": "Celarent"
+ },
+ {
+ "from": "DefAEIO",
+ "to": "Darii"
+ },
+ {
+ "from": "DefFig",
+ "to": "Darii"
+ },
+ {
+ "from": "DefAEIO",
+ "to": "Ferio"
+ },
+ {
+ "from": "DefFig",
+ "to": "Ferio"
+ },
+ {
+ "from": "Econv",
+ "to": "Cesare"
+ },
+ {
+ "from": "Celarent",
+ "to": "Cesare"
+ },
+ {
+ "from": "Econv",
+ "to": "Camestres"
+ },
+ {
+ "from": "Celarent",
+ "to": "Camestres"
+ },
+ {
+ "from": "Econv",
+ "to": "Festino"
+ },
+ {
+ "from": "Ferio",
+ "to": "Festino"
+ },
+ {
+ "from": "Barbara",
+ "to": "Baroco"
+ },
+ {
+ "from": "Aconv",
+ "to": "Darapti"
+ },
+ {
+ "from": "Darii",
+ "to": "Darapti"
+ },
+ {
+ "from": "Aconv",
+ "to": "Felapton"
+ },
+ {
+ "from": "Ferio",
+ "to": "Felapton"
+ },
+ {
+ "from": "Iconv",
+ "to": "Disamis"
+ },
+ {
+ "from": "Darii",
+ "to": "Disamis"
+ },
+ {
+ "from": "Aconv",
+ "to": "Datisi"
+ },
+ {
+ "from": "Darii",
+ "to": "Datisi"
+ },
+ {
+ "from": "Barbara",
+ "to": "Bocardo"
+ },
+ {
+ "from": "Aconv",
+ "to": "Ferison"
+ },
+ {
+ "from": "Ferio",
+ "to": "Ferison"
+ }
+ ],
+ "colorScheme": {
+ "axiom": {
+ "fill": "#e74c3c",
+ "stroke": "#c0392b"
+ },
+ "definition": {
+ "fill": "#3498db",
+ "stroke": "#2980b9"
+ },
+ "theorem": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/aristotle-syllogistic.mmd b/data/aristotle-syllogistic.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..5b8a89c8ef3ccbaadbdeae1bae110ec357a43852
--- /dev/null
+++ b/data/aristotle-syllogistic.mmd
@@ -0,0 +1,52 @@
+graph TD
+ DefAEIO("DefAEIO Categorical forms")
+ DefFig("DefFig Figures 1,2,3")
+ Econv("Econv E-conversion")
+ Iconv("Iconv I-conversion")
+ Aconv("Aconv A-conversion")
+ Barbara("Barbara Barbara")
+ Celarent("Celarent Celarent")
+ Darii("Darii Darii")
+ Ferio("Ferio Ferio")
+ Cesare("Cesare Cesare")
+ Camestres("Camestres Camestres")
+ Festino("Festino Festino")
+ Baroco("Baroco Baroco")
+ Darapti("Darapti Darapti")
+ Felapton("Felapton Felapton")
+ Disamis("Disamis Disamis")
+ Datisi("Datisi Datisi")
+ Bocardo("Bocardo Bocardo")
+ Ferison("Ferison Ferison")
+ DefAEIO --> Barbara
+ DefFig --> Barbara
+ DefAEIO --> Celarent
+ DefFig --> Celarent
+ DefAEIO --> Darii
+ DefFig --> Darii
+ DefAEIO --> Ferio
+ DefFig --> Ferio
+ Econv --> Cesare
+ Celarent --> Cesare
+ Econv --> Camestres
+ Celarent --> Camestres
+ Econv --> Festino
+ Ferio --> Festino
+ Barbara --> Baroco
+ Aconv --> Darapti
+ Darii --> Darapti
+ Aconv --> Felapton
+ Ferio --> Felapton
+ Iconv --> Disamis
+ Darii --> Disamis
+ Aconv --> Datisi
+ Darii --> Datisi
+ Barbara --> Bocardo
+ Aconv --> Ferison
+ Ferio --> Ferison
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class Econv,Iconv,Aconv,Barbara,Celarent,Darii,Ferio axiom
+ class DefAEIO,DefFig definition
+ class Cesare,Camestres,Festino,Baroco,Darapti,Felapton,Disamis,Datisi,Bocardo,Ferison theorem
\ No newline at end of file
diff --git a/data/combinatorics-advanced-counting.mmd b/data/combinatorics-advanced-counting.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..48e2d60940bda01dff18d3f590646cdd71f49fb1
--- /dev/null
+++ b/data/combinatorics-advanced-counting.mmd
@@ -0,0 +1,24 @@
+graph TD
+ DefFact("DefFact Factorial")
+ DefSum("DefSum Sum principle")
+ DefProd("DefProd Product principle")
+ PermNoRep("PermNoRep Permutations no rep")
+ Pigeonhole("Pigeonhole Pigeonhole principle")
+ InclExcl("InclExcl Inclusion-exclusion")
+ InclExcl3("InclExcl3 Incl-excl 3 sets")
+ Derange("Derange Derangements")
+ Stirling2("Stirling2 Stirling numbers")
+ DefFact --> PermNoRep
+ DefProd --> PermNoRep
+ DefSum --> Pigeonhole
+ DefSum --> InclExcl
+ InclExcl --> InclExcl3
+ InclExcl --> Derange
+ PermNoRep --> Derange
+ DefSum --> Stirling2
+ DefProd --> Stirling2
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class DefFact,DefSum,DefProd definition
+ class PermNoRep,Pigeonhole,InclExcl,InclExcl3,Derange,Stirling2 theorem
\ No newline at end of file
diff --git a/data/combinatorics-combinations-binomial.mmd b/data/combinatorics-combinations-binomial.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..5e54b07b55cbbac9b25b54491cfa1f8bb44c77d7
--- /dev/null
+++ b/data/combinatorics-combinations-binomial.mmd
@@ -0,0 +1,20 @@
+graph TD
+ DefFact("DefFact Factorial")
+ DefProd("DefProd Product principle")
+ PermNoRep("PermNoRep Permutations no rep")
+ CombNoRep("CombNoRep Combinations")
+ CombRep("CombRep Combinations with rep")
+ BinomThm("BinomThm Binomial theorem")
+ Pascal("Pascal Pascal identity")
+ DefFact --> PermNoRep
+ DefProd --> PermNoRep
+ PermNoRep --> CombNoRep
+ DefFact --> CombNoRep
+ CombNoRep --> CombRep
+ CombNoRep --> BinomThm
+ CombNoRep --> Pascal
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class DefFact,DefProd definition
+ class PermNoRep,CombNoRep,CombRep,BinomThm,Pascal theorem
\ No newline at end of file
diff --git a/data/combinatorics-principles-permutations.mmd b/data/combinatorics-principles-permutations.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..5c34a9628df525a83165cb4a036d723645482af5
--- /dev/null
+++ b/data/combinatorics-principles-permutations.mmd
@@ -0,0 +1,17 @@
+graph TD
+ DefFact("DefFact Factorial")
+ DefSum("DefSum Sum principle")
+ DefProd("DefProd Product principle")
+ PermNoRep("PermNoRep Permutations no rep")
+ PermRep("PermRep Permutations with rep")
+ CombNoRep("CombNoRep Combinations")
+ DefFact --> PermNoRep
+ DefProd --> PermNoRep
+ DefProd --> PermRep
+ PermNoRep --> CombNoRep
+ DefFact --> CombNoRep
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class DefFact,DefSum,DefProd definition
+ class PermNoRep,PermRep,CombNoRep theorem
\ No newline at end of file
diff --git a/data/combinatorics.json b/data/combinatorics.json
new file mode 100644
index 0000000000000000000000000000000000000000..4bd3b400f48e129dd4b24d264624613c7abae266
--- /dev/null
+++ b/data/combinatorics.json
@@ -0,0 +1,239 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "combinatorics",
+ "name": "Combinatorics",
+ "subject": "discrete_mathematics",
+ "variant": "classical",
+ "description": "Counting principles: sum and product rules, permutations (with/without repetition), combinations, binomial theorem, pigeonhole principle, inclusion-exclusion, derangements.",
+ "structure": {
+ "axioms": 0,
+ "definitions": 3,
+ "theorems": 11
+ }
+ },
+ "metadata": {
+ "created": "2026-03-15",
+ "lastUpdated": "2026-03-15",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Combinatorics Dependency Graph. Programming Framework.",
+ "keywords": [
+ "combinatorics",
+ "permutations",
+ "combinations",
+ "counting",
+ "binomial theorem"
+ ]
+ },
+ "sources": [
+ {
+ "id": "dmoi",
+ "type": "primary",
+ "title": "Discrete Mathematics: An Open Introduction",
+ "url": "https://discrete.openmathbooks.org/dmoi4/sec_counting-combperm.html",
+ "notes": "Counting principles"
+ },
+ {
+ "id": "mathisfun",
+ "type": "digital",
+ "title": "Combinations and Permutations",
+ "url": "https://www.mathsisfun.com/combinatorics/combinations-permutations.html",
+ "notes": "Formulas"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "DefFact",
+ "type": "definition",
+ "label": "Factorial: n! = n(n-1)...1, 0!=1",
+ "shortLabel": "DefFact",
+ "short": "Factorial",
+ "colorClass": "definition"
+ },
+ {
+ "id": "DefSum",
+ "type": "definition",
+ "label": "Sum principle: disjoint choices add (OR)",
+ "shortLabel": "DefSum",
+ "short": "Sum principle",
+ "colorClass": "definition"
+ },
+ {
+ "id": "DefProd",
+ "type": "definition",
+ "label": "Product principle: sequential choices multiply (AND)",
+ "shortLabel": "DefProd",
+ "short": "Product principle",
+ "colorClass": "definition"
+ },
+ {
+ "id": "PermNoRep",
+ "type": "theorem",
+ "label": "P(n,r) = n!/(n-r)! arrangements of r from n",
+ "shortLabel": "PermNoRep",
+ "short": "Permutations no rep",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "PermRep",
+ "type": "theorem",
+ "label": "n^r arrangements of r from n with repetition",
+ "shortLabel": "PermRep",
+ "short": "Permutations with rep",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "CombNoRep",
+ "type": "theorem",
+ "label": "C(n,r) = n!/(r!(n-r)!) = P(n,r)/r!",
+ "shortLabel": "CombNoRep",
+ "short": "Combinations",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "CombRep",
+ "type": "theorem",
+ "label": "C(n+r-1,r) ways to choose r from n with rep",
+ "shortLabel": "CombRep",
+ "short": "Combinations with rep",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "BinomThm",
+ "type": "theorem",
+ "label": "(a+b)^n = sum C(n,k) a^k b^(n-k)",
+ "shortLabel": "BinomThm",
+ "short": "Binomial theorem",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Pascal",
+ "type": "theorem",
+ "label": "C(n,k) = C(n-1,k-1) + C(n-1,k)",
+ "shortLabel": "Pascal",
+ "short": "Pascal identity",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Pigeonhole",
+ "type": "theorem",
+ "label": "n+1 objects in n boxes implies one box has 2+",
+ "shortLabel": "Pigeonhole",
+ "short": "Pigeonhole principle",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "InclExcl",
+ "type": "theorem",
+ "label": "|A union B| = |A| + |B| - |A intersect B|",
+ "shortLabel": "InclExcl",
+ "short": "Inclusion-exclusion",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "InclExcl3",
+ "type": "theorem",
+ "label": "Inclusion-exclusion for 3 sets",
+ "shortLabel": "InclExcl3",
+ "short": "Incl-excl 3 sets",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Derange",
+ "type": "theorem",
+ "label": "D(n) = n! sum (-1)^k/k! derangements",
+ "shortLabel": "Derange",
+ "short": "Derangements",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "Stirling2",
+ "type": "theorem",
+ "label": "S(n,k) = partitions of n into k nonempty sets",
+ "shortLabel": "Stirling2",
+ "short": "Stirling numbers",
+ "colorClass": "theorem"
+ }
+ ],
+ "edges": [
+ {
+ "from": "DefFact",
+ "to": "PermNoRep"
+ },
+ {
+ "from": "DefProd",
+ "to": "PermNoRep"
+ },
+ {
+ "from": "DefProd",
+ "to": "PermRep"
+ },
+ {
+ "from": "PermNoRep",
+ "to": "CombNoRep"
+ },
+ {
+ "from": "DefFact",
+ "to": "CombNoRep"
+ },
+ {
+ "from": "CombNoRep",
+ "to": "CombRep"
+ },
+ {
+ "from": "CombNoRep",
+ "to": "BinomThm"
+ },
+ {
+ "from": "CombNoRep",
+ "to": "Pascal"
+ },
+ {
+ "from": "DefSum",
+ "to": "Pigeonhole"
+ },
+ {
+ "from": "DefSum",
+ "to": "InclExcl"
+ },
+ {
+ "from": "InclExcl",
+ "to": "InclExcl3"
+ },
+ {
+ "from": "InclExcl",
+ "to": "Derange"
+ },
+ {
+ "from": "PermNoRep",
+ "to": "Derange"
+ },
+ {
+ "from": "DefSum",
+ "to": "Stirling2"
+ },
+ {
+ "from": "DefProd",
+ "to": "Stirling2"
+ }
+ ],
+ "colorScheme": {
+ "axiom": {
+ "fill": "#e74c3c",
+ "stroke": "#c0392b"
+ },
+ "definition": {
+ "fill": "#3498db",
+ "stroke": "#2980b9"
+ },
+ "theorem": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/combinatorics.mmd b/data/combinatorics.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..dd7cf62d5e3024dc8e9cfd0ce6177385b03fdc39
--- /dev/null
+++ b/data/combinatorics.mmd
@@ -0,0 +1,35 @@
+graph TD
+ DefFact("DefFact Factorial")
+ DefSum("DefSum Sum principle")
+ DefProd("DefProd Product principle")
+ PermNoRep("PermNoRep Permutations no rep")
+ PermRep("PermRep Permutations with rep")
+ CombNoRep("CombNoRep Combinations")
+ CombRep("CombRep Combinations with rep")
+ BinomThm("BinomThm Binomial theorem")
+ Pascal("Pascal Pascal identity")
+ Pigeonhole("Pigeonhole Pigeonhole principle")
+ InclExcl("InclExcl Inclusion-exclusion")
+ InclExcl3("InclExcl3 Incl-excl 3 sets")
+ Derange("Derange Derangements")
+ Stirling2("Stirling2 Stirling numbers")
+ DefFact --> PermNoRep
+ DefProd --> PermNoRep
+ DefProd --> PermRep
+ PermNoRep --> CombNoRep
+ DefFact --> CombNoRep
+ CombNoRep --> CombRep
+ CombNoRep --> BinomThm
+ CombNoRep --> Pascal
+ DefSum --> Pigeonhole
+ DefSum --> InclExcl
+ InclExcl --> InclExcl3
+ InclExcl --> Derange
+ PermNoRep --> Derange
+ DefSum --> Stirling2
+ DefProd --> Stirling2
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class DefFact,DefSum,DefProd definition
+ class PermNoRep,PermRep,CombNoRep,CombRep,BinomThm,Pascal,Pigeonhole,InclExcl,InclExcl3,Derange,Stirling2 theorem
\ No newline at end of file
diff --git a/data/euclid-elements-book-i-props-1-10.mmd b/data/euclid-elements-book-i-props-1-10.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..2adc89278e23a45a078b05db3d0f6b4228c7d869
--- /dev/null
+++ b/data/euclid-elements-book-i-props-1-10.mmd
@@ -0,0 +1,49 @@
+graph TD
+ P1["Post. 1\nDraw a straight line from any po..."]
+ P2["Post. 2\nProduce a finite straight line c..."]
+ P3["Post. 3\nDescribe a circle with any cente..."]
+ P4["Post. 4\nAll right angles equal one another"]
+ P5["Post. 5\nParallel postulate: if interior ..."]
+ CN1["CN 1\nThings equal to the same thing a..."]
+ CN2["CN 2\nIf equals are added to equals, t..."]
+ CN3["CN 3\nIf equals are subtracted from eq..."]
+ CN4["CN 4\nThings coinciding with one anoth..."]
+ CN5["CN 5\nThe whole is greater than the part"]
+ Prop1["Prop. I.1\nEquilateral triangle on given line"]
+ Prop2["Prop. I.2\nPlace line equal to given at point"]
+ Prop3["Prop. I.3\nCut off from greater segment equal to less"]
+ Prop4["Prop. I.4\nSAS congruence"]
+ Prop5["Prop. I.5\nBase angles of isosceles equal"]
+ Prop6["Prop. I.6\nSides opposite equal angles equal"]
+ Prop7["Prop. I.7\nUniqueness of triangle from ends"]
+ Prop8["Prop. I.8\nSSS congruence"]
+ Prop9["Prop. I.9\nBisect angle"]
+ Prop10["Prop. I.10\nBisect line"]
+ P1 --> Prop1
+ P3 --> Prop1
+ Prop1 --> Prop2
+ P1 --> Prop2
+ P2 --> Prop2
+ P3 --> Prop2
+ Prop2 --> Prop3
+ P3 --> Prop3
+ CN4 --> Prop4
+ CN5 --> Prop4
+ Prop3 --> Prop5
+ Prop4 --> Prop5
+ Prop3 --> Prop6
+ Prop4 --> Prop6
+ Prop5 --> Prop7
+ Prop7 --> Prop8
+ Prop1 --> Prop9
+ Prop3 --> Prop9
+ Prop8 --> Prop9
+ Prop1 --> Prop10
+ Prop4 --> Prop10
+ Prop9 --> Prop10
+ classDef postulate fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef commonNotion fill:#9b59b6,color:#fff,stroke:#8e44ad
+ classDef proposition fill:#1abc9c,color:#fff,stroke:#16a085
+ class P1,P2,P3,P4,P5 postulate
+ class CN1,CN2,CN3,CN4,CN5 commonNotion
+ class Prop1,Prop2,Prop3,Prop4,Prop5,Prop6,Prop7,Prop8,Prop9,Prop10 proposition
\ No newline at end of file
diff --git a/data/euclid-elements-book-i-props-11-20.mmd b/data/euclid-elements-book-i-props-11-20.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..684379d465fbbfe1826543247ee3cd759ae17028
--- /dev/null
+++ b/data/euclid-elements-book-i-props-11-20.mmd
@@ -0,0 +1,78 @@
+graph TD
+ P1["Post. 1\nDraw a straight line from any po..."]
+ P2["Post. 2\nProduce a finite straight line c..."]
+ P3["Post. 3\nDescribe a circle with any cente..."]
+ P4["Post. 4\nAll right angles equal one another"]
+ P5["Post. 5\nParallel postulate: if interior ..."]
+ CN1["CN 1\nThings equal to the same thing a..."]
+ CN2["CN 2\nIf equals are added to equals, t..."]
+ CN3["CN 3\nIf equals are subtracted from eq..."]
+ CN4["CN 4\nThings coinciding with one anoth..."]
+ CN5["CN 5\nThe whole is greater than the part"]
+ Prop1["Prop. I.1\nEquilateral triangle on given line"]
+ Prop2["Prop. I.2\nPlace line equal to given at point"]
+ Prop3["Prop. I.3\nCut off from greater segment equal to less"]
+ Prop4["Prop. I.4\nSAS congruence"]
+ Prop5["Prop. I.5\nBase angles of isosceles equal"]
+ Prop7["Prop. I.7\nUniqueness of triangle from ends"]
+ Prop8["Prop. I.8\nSSS congruence"]
+ Prop9["Prop. I.9\nBisect angle"]
+ Prop10["Prop. I.10\nBisect line"]
+ Prop11["Prop. I.11\nPerpendicular from point on line"]
+ Prop12["Prop. I.12\nPerpendicular from point not on line"]
+ Prop13["Prop. I.13\nAngles on line sum to two right"]
+ Prop14["Prop. I.14\nIf angles sum to two right, straight line"]
+ Prop15["Prop. I.15\nVertical angles equal"]
+ Prop16["Prop. I.16\nExterior angle > interior opposite"]
+ Prop17["Prop. I.17\nSum of two angles < two right"]
+ Prop18["Prop. I.18\nAngle opposite greater side greater"]
+ Prop19["Prop. I.19\nSide opposite greater angle greater"]
+ Prop20["Prop. I.20\nTriangle inequality"]
+ P1 --> Prop1
+ P3 --> Prop1
+ Prop1 --> Prop2
+ P1 --> Prop2
+ P2 --> Prop2
+ P3 --> Prop2
+ Prop2 --> Prop3
+ P3 --> Prop3
+ CN4 --> Prop4
+ CN5 --> Prop4
+ Prop3 --> Prop5
+ Prop4 --> Prop5
+ Prop5 --> Prop7
+ Prop7 --> Prop8
+ Prop1 --> Prop9
+ Prop3 --> Prop9
+ Prop8 --> Prop9
+ Prop1 --> Prop10
+ Prop4 --> Prop10
+ Prop9 --> Prop10
+ Prop1 --> Prop11
+ Prop3 --> Prop11
+ Prop8 --> Prop11
+ Prop8 --> Prop12
+ Prop10 --> Prop12
+ Prop11 --> Prop13
+ Prop13 --> Prop14
+ Prop13 --> Prop15
+ Prop3 --> Prop16
+ Prop4 --> Prop16
+ Prop10 --> Prop16
+ Prop15 --> Prop16
+ Prop13 --> Prop17
+ Prop16 --> Prop17
+ Prop3 --> Prop18
+ Prop5 --> Prop18
+ Prop16 --> Prop18
+ Prop5 --> Prop19
+ Prop18 --> Prop19
+ Prop3 --> Prop20
+ Prop5 --> Prop20
+ Prop19 --> Prop20
+ classDef postulate fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef commonNotion fill:#9b59b6,color:#fff,stroke:#8e44ad
+ classDef proposition fill:#1abc9c,color:#fff,stroke:#16a085
+ class P1,P2,P3,P4,P5 postulate
+ class CN1,CN2,CN3,CN4,CN5 commonNotion
+ class Prop1,Prop2,Prop3,Prop4,Prop5,Prop7,Prop8,Prop9,Prop10,Prop11,Prop12,Prop13,Prop14,Prop15,Prop16,Prop17,Prop18,Prop19,Prop20 proposition
\ No newline at end of file
diff --git a/data/euclid-elements-book-i-props-21-30.mmd b/data/euclid-elements-book-i-props-21-30.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..79cc4c69f5bc05b4f89dbfd21a49ae04d3a2595b
--- /dev/null
+++ b/data/euclid-elements-book-i-props-21-30.mmd
@@ -0,0 +1,105 @@
+graph TD
+ P1["Post. 1\nDraw a straight line from any po..."]
+ P2["Post. 2\nProduce a finite straight line c..."]
+ P3["Post. 3\nDescribe a circle with any cente..."]
+ P4["Post. 4\nAll right angles equal one another"]
+ P5["Post. 5\nParallel postulate: if interior ..."]
+ CN1["CN 1\nThings equal to the same thing a..."]
+ CN2["CN 2\nIf equals are added to equals, t..."]
+ CN3["CN 3\nIf equals are subtracted from eq..."]
+ CN4["CN 4\nThings coinciding with one anoth..."]
+ CN5["CN 5\nThe whole is greater than the part"]
+ Prop1["Prop. I.1\nEquilateral triangle on given line"]
+ Prop2["Prop. I.2\nPlace line equal to given at point"]
+ Prop3["Prop. I.3\nCut off from greater segment equal to less"]
+ Prop4["Prop. I.4\nSAS congruence"]
+ Prop5["Prop. I.5\nBase angles of isosceles equal"]
+ Prop7["Prop. I.7\nUniqueness of triangle from ends"]
+ Prop8["Prop. I.8\nSSS congruence"]
+ Prop9["Prop. I.9\nBisect angle"]
+ Prop10["Prop. I.10\nBisect line"]
+ Prop11["Prop. I.11\nPerpendicular from point on line"]
+ Prop13["Prop. I.13\nAngles on line sum to two right"]
+ Prop15["Prop. I.15\nVertical angles equal"]
+ Prop16["Prop. I.16\nExterior angle > interior opposite"]
+ Prop18["Prop. I.18\nAngle opposite greater side greater"]
+ Prop19["Prop. I.19\nSide opposite greater angle greater"]
+ Prop20["Prop. I.20\nTriangle inequality"]
+ Prop21["Prop. I.21\nLines from ends within triangle"]
+ Prop22["Prop. I.22\nConstruct triangle from three lines"]
+ Prop23["Prop. I.23\nConstruct angle equal to given"]
+ Prop24["Prop. I.24\nSAS for greater angle => greater base"]
+ Prop25["Prop. I.25\nSAS for greater base => greater angle"]
+ Prop26["Prop. I.26\nAAS congruence"]
+ Prop27["Prop. I.27\nAlternate angles equal => parallel"]
+ Prop28["Prop. I.28\nExterior = interior opposite => parallel"]
+ Prop29["Prop. I.29\nParallel => alternate angles equal"]
+ Prop30["Prop. I.30\nTransitivity of parallel"]
+ P1 --> Prop1
+ P3 --> Prop1
+ Prop1 --> Prop2
+ P1 --> Prop2
+ P2 --> Prop2
+ P3 --> Prop2
+ Prop2 --> Prop3
+ P3 --> Prop3
+ CN4 --> Prop4
+ CN5 --> Prop4
+ Prop3 --> Prop5
+ Prop4 --> Prop5
+ Prop5 --> Prop7
+ Prop7 --> Prop8
+ Prop1 --> Prop9
+ Prop3 --> Prop9
+ Prop8 --> Prop9
+ Prop1 --> Prop10
+ Prop4 --> Prop10
+ Prop9 --> Prop10
+ Prop1 --> Prop11
+ Prop3 --> Prop11
+ Prop8 --> Prop11
+ Prop11 --> Prop13
+ Prop13 --> Prop15
+ Prop3 --> Prop16
+ Prop4 --> Prop16
+ Prop10 --> Prop16
+ Prop15 --> Prop16
+ Prop3 --> Prop18
+ Prop5 --> Prop18
+ Prop16 --> Prop18
+ Prop5 --> Prop19
+ Prop18 --> Prop19
+ Prop3 --> Prop20
+ Prop5 --> Prop20
+ Prop19 --> Prop20
+ Prop16 --> Prop21
+ Prop20 --> Prop21
+ Prop3 --> Prop22
+ Prop20 --> Prop22
+ Prop8 --> Prop23
+ Prop22 --> Prop23
+ Prop3 --> Prop24
+ Prop4 --> Prop24
+ Prop5 --> Prop24
+ Prop19 --> Prop24
+ Prop23 --> Prop24
+ Prop4 --> Prop25
+ Prop24 --> Prop25
+ Prop3 --> Prop26
+ Prop4 --> Prop26
+ Prop16 --> Prop26
+ Prop16 --> Prop27
+ Prop13 --> Prop28
+ Prop15 --> Prop28
+ Prop27 --> Prop28
+ Prop13 --> Prop29
+ Prop15 --> Prop29
+ Prop27 --> Prop29
+ P5 --> Prop29
+ Prop29 --> Prop30
+ classDef postulate fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef commonNotion fill:#9b59b6,color:#fff,stroke:#8e44ad
+ classDef proposition fill:#1abc9c,color:#fff,stroke:#16a085
+ class P1,P2,P3,P4,P5 postulate
+ class CN1,CN2,CN3,CN4,CN5 commonNotion
+ class Prop1,Prop2,Prop3,Prop4,Prop5,Prop7,Prop8,Prop9,Prop10,Prop11,Prop13,Prop15,Prop16,Prop18,Prop19,Prop20,Prop21,Prop22,Prop23,Prop24,Prop25,Prop26,Prop27,Prop28,Prop29,Prop30 proposition
\ No newline at end of file
diff --git a/data/euclid-elements-book-i-props-31-41.mmd b/data/euclid-elements-book-i-props-31-41.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..b2a6e5ce28d248b018fac455cd03bfee57571032
--- /dev/null
+++ b/data/euclid-elements-book-i-props-31-41.mmd
@@ -0,0 +1,127 @@
+graph TD
+ P1["Post. 1\nDraw a straight line from any po..."]
+ P2["Post. 2\nProduce a finite straight line c..."]
+ P3["Post. 3\nDescribe a circle with any cente..."]
+ P4["Post. 4\nAll right angles equal one another"]
+ P5["Post. 5\nParallel postulate: if interior ..."]
+ CN1["CN 1\nThings equal to the same thing a..."]
+ CN2["CN 2\nIf equals are added to equals, t..."]
+ CN3["CN 3\nIf equals are subtracted from eq..."]
+ CN4["CN 4\nThings coinciding with one anoth..."]
+ CN5["CN 5\nThe whole is greater than the part"]
+ Prop1["Prop. I.1\nEquilateral triangle on given line"]
+ Prop2["Prop. I.2\nPlace line equal to given at point"]
+ Prop3["Prop. I.3\nCut off from greater segment equal to less"]
+ Prop4["Prop. I.4\nSAS congruence"]
+ Prop5["Prop. I.5\nBase angles of isosceles equal"]
+ Prop7["Prop. I.7\nUniqueness of triangle from ends"]
+ Prop8["Prop. I.8\nSSS congruence"]
+ Prop9["Prop. I.9\nBisect angle"]
+ Prop10["Prop. I.10\nBisect line"]
+ Prop11["Prop. I.11\nPerpendicular from point on line"]
+ Prop13["Prop. I.13\nAngles on line sum to two right"]
+ Prop15["Prop. I.15\nVertical angles equal"]
+ Prop16["Prop. I.16\nExterior angle > interior opposite"]
+ Prop18["Prop. I.18\nAngle opposite greater side greater"]
+ Prop19["Prop. I.19\nSide opposite greater angle greater"]
+ Prop20["Prop. I.20\nTriangle inequality"]
+ Prop22["Prop. I.22\nConstruct triangle from three lines"]
+ Prop23["Prop. I.23\nConstruct angle equal to given"]
+ Prop26["Prop. I.26\nAAS congruence"]
+ Prop27["Prop. I.27\nAlternate angles equal => parallel"]
+ Prop29["Prop. I.29\nParallel => alternate angles equal"]
+ Prop31["Prop. I.31\nDraw parallel through point"]
+ Prop32["Prop. I.32\nExterior angle = sum interior opposite"]
+ Prop33["Prop. I.33\nJoining ends of equal parallel lines"]
+ Prop34["Prop. I.34\nParallelogram properties"]
+ Prop35["Prop. I.35\nParallelograms same base equal"]
+ Prop36["Prop. I.36\nParallelograms equal bases equal"]
+ Prop37["Prop. I.37\nTriangles same base equal"]
+ Prop38["Prop. I.38\nTriangles equal bases equal"]
+ Prop39["Prop. I.39\nEqual triangles same base same side"]
+ Prop40["Prop. I.40\nEqual triangles equal bases same side"]
+ Prop41["Prop. I.41\nParallelogram = 2× triangle"]
+ P1 --> Prop1
+ P3 --> Prop1
+ Prop1 --> Prop2
+ P1 --> Prop2
+ P2 --> Prop2
+ P3 --> Prop2
+ Prop2 --> Prop3
+ P3 --> Prop3
+ CN4 --> Prop4
+ CN5 --> Prop4
+ Prop3 --> Prop5
+ Prop4 --> Prop5
+ Prop5 --> Prop7
+ Prop7 --> Prop8
+ Prop1 --> Prop9
+ Prop3 --> Prop9
+ Prop8 --> Prop9
+ Prop1 --> Prop10
+ Prop4 --> Prop10
+ Prop9 --> Prop10
+ Prop1 --> Prop11
+ Prop3 --> Prop11
+ Prop8 --> Prop11
+ Prop11 --> Prop13
+ Prop13 --> Prop15
+ Prop3 --> Prop16
+ Prop4 --> Prop16
+ Prop10 --> Prop16
+ Prop15 --> Prop16
+ Prop3 --> Prop18
+ Prop5 --> Prop18
+ Prop16 --> Prop18
+ Prop5 --> Prop19
+ Prop18 --> Prop19
+ Prop3 --> Prop20
+ Prop5 --> Prop20
+ Prop19 --> Prop20
+ Prop3 --> Prop22
+ Prop20 --> Prop22
+ Prop8 --> Prop23
+ Prop22 --> Prop23
+ Prop3 --> Prop26
+ Prop4 --> Prop26
+ Prop16 --> Prop26
+ Prop16 --> Prop27
+ Prop13 --> Prop29
+ Prop15 --> Prop29
+ Prop27 --> Prop29
+ P5 --> Prop29
+ Prop23 --> Prop31
+ Prop27 --> Prop31
+ Prop13 --> Prop32
+ Prop29 --> Prop32
+ Prop31 --> Prop32
+ Prop4 --> Prop33
+ Prop27 --> Prop33
+ Prop29 --> Prop33
+ Prop4 --> Prop34
+ Prop26 --> Prop34
+ Prop29 --> Prop34
+ Prop4 --> Prop35
+ Prop29 --> Prop35
+ Prop34 --> Prop35
+ Prop33 --> Prop36
+ Prop34 --> Prop36
+ Prop35 --> Prop36
+ Prop31 --> Prop37
+ Prop34 --> Prop37
+ Prop35 --> Prop37
+ Prop31 --> Prop38
+ Prop34 --> Prop38
+ Prop36 --> Prop38
+ Prop31 --> Prop39
+ Prop37 --> Prop39
+ Prop31 --> Prop40
+ Prop38 --> Prop40
+ Prop34 --> Prop41
+ Prop37 --> Prop41
+ classDef postulate fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef commonNotion fill:#9b59b6,color:#fff,stroke:#8e44ad
+ classDef proposition fill:#1abc9c,color:#fff,stroke:#16a085
+ class P1,P2,P3,P4,P5 postulate
+ class CN1,CN2,CN3,CN4,CN5 commonNotion
+ class Prop1,Prop2,Prop3,Prop4,Prop5,Prop7,Prop8,Prop9,Prop10,Prop11,Prop13,Prop15,Prop16,Prop18,Prop19,Prop20,Prop22,Prop23,Prop26,Prop27,Prop29,Prop31,Prop32,Prop33,Prop34,Prop35,Prop36,Prop37,Prop38,Prop39,Prop40,Prop41 proposition
\ No newline at end of file
diff --git a/data/euclid-elements-book-i-props-42-48.mmd b/data/euclid-elements-book-i-props-42-48.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..ecb6934b94cacf8426406f860cbd17c9ff589a5f
--- /dev/null
+++ b/data/euclid-elements-book-i-props-42-48.mmd
@@ -0,0 +1,160 @@
+graph TD
+ P1["Post. 1\nDraw a straight line from any po..."]
+ P2["Post. 2\nProduce a finite straight line c..."]
+ P3["Post. 3\nDescribe a circle with any cente..."]
+ P4["Post. 4\nAll right angles equal one another"]
+ P5["Post. 5\nParallel postulate: if interior ..."]
+ CN1["CN 1\nThings equal to the same thing a..."]
+ CN2["CN 2\nIf equals are added to equals, t..."]
+ CN3["CN 3\nIf equals are subtracted from eq..."]
+ CN4["CN 4\nThings coinciding with one anoth..."]
+ CN5["CN 5\nThe whole is greater than the part"]
+ Prop1["Prop. I.1\nEquilateral triangle on given line"]
+ Prop2["Prop. I.2\nPlace line equal to given at point"]
+ Prop3["Prop. I.3\nCut off from greater segment equal to less"]
+ Prop4["Prop. I.4\nSAS congruence"]
+ Prop5["Prop. I.5\nBase angles of isosceles equal"]
+ Prop7["Prop. I.7\nUniqueness of triangle from ends"]
+ Prop8["Prop. I.8\nSSS congruence"]
+ Prop9["Prop. I.9\nBisect angle"]
+ Prop10["Prop. I.10\nBisect line"]
+ Prop11["Prop. I.11\nPerpendicular from point on line"]
+ Prop13["Prop. I.13\nAngles on line sum to two right"]
+ Prop14["Prop. I.14\nIf angles sum to two right, straight line"]
+ Prop15["Prop. I.15\nVertical angles equal"]
+ Prop16["Prop. I.16\nExterior angle > interior opposite"]
+ Prop18["Prop. I.18\nAngle opposite greater side greater"]
+ Prop19["Prop. I.19\nSide opposite greater angle greater"]
+ Prop20["Prop. I.20\nTriangle inequality"]
+ Prop22["Prop. I.22\nConstruct triangle from three lines"]
+ Prop23["Prop. I.23\nConstruct angle equal to given"]
+ Prop26["Prop. I.26\nAAS congruence"]
+ Prop27["Prop. I.27\nAlternate angles equal => parallel"]
+ Prop29["Prop. I.29\nParallel => alternate angles equal"]
+ Prop30["Prop. I.30\nTransitivity of parallel"]
+ Prop31["Prop. I.31\nDraw parallel through point"]
+ Prop33["Prop. I.33\nJoining ends of equal parallel lines"]
+ Prop34["Prop. I.34\nParallelogram properties"]
+ Prop35["Prop. I.35\nParallelograms same base equal"]
+ Prop36["Prop. I.36\nParallelograms equal bases equal"]
+ Prop37["Prop. I.37\nTriangles same base equal"]
+ Prop38["Prop. I.38\nTriangles equal bases equal"]
+ Prop41["Prop. I.41\nParallelogram = 2× triangle"]
+ Prop42["Prop. I.42\nConstruct parallelogram = triangle"]
+ Prop43["Prop. I.43\nComplements of parallelogram"]
+ Prop44["Prop. I.44\nApply parallelogram to line"]
+ Prop45["Prop. I.45\nConstruct parallelogram = rectilinear figure"]
+ Prop46["Prop. I.46\nConstruct square on line"]
+ Prop47["Prop. I.47\nPythagorean theorem"]
+ Prop48["Prop. I.48\nConverse Pythagorean"]
+ P1 --> Prop1
+ P3 --> Prop1
+ Prop1 --> Prop2
+ P1 --> Prop2
+ P2 --> Prop2
+ P3 --> Prop2
+ Prop2 --> Prop3
+ P3 --> Prop3
+ CN4 --> Prop4
+ CN5 --> Prop4
+ Prop3 --> Prop5
+ Prop4 --> Prop5
+ Prop5 --> Prop7
+ Prop7 --> Prop8
+ Prop1 --> Prop9
+ Prop3 --> Prop9
+ Prop8 --> Prop9
+ Prop1 --> Prop10
+ Prop4 --> Prop10
+ Prop9 --> Prop10
+ Prop1 --> Prop11
+ Prop3 --> Prop11
+ Prop8 --> Prop11
+ Prop11 --> Prop13
+ Prop13 --> Prop14
+ Prop13 --> Prop15
+ Prop3 --> Prop16
+ Prop4 --> Prop16
+ Prop10 --> Prop16
+ Prop15 --> Prop16
+ Prop3 --> Prop18
+ Prop5 --> Prop18
+ Prop16 --> Prop18
+ Prop5 --> Prop19
+ Prop18 --> Prop19
+ Prop3 --> Prop20
+ Prop5 --> Prop20
+ Prop19 --> Prop20
+ Prop3 --> Prop22
+ Prop20 --> Prop22
+ Prop8 --> Prop23
+ Prop22 --> Prop23
+ Prop3 --> Prop26
+ Prop4 --> Prop26
+ Prop16 --> Prop26
+ Prop16 --> Prop27
+ Prop13 --> Prop29
+ Prop15 --> Prop29
+ Prop27 --> Prop29
+ P5 --> Prop29
+ Prop29 --> Prop30
+ Prop23 --> Prop31
+ Prop27 --> Prop31
+ Prop4 --> Prop33
+ Prop27 --> Prop33
+ Prop29 --> Prop33
+ Prop4 --> Prop34
+ Prop26 --> Prop34
+ Prop29 --> Prop34
+ Prop4 --> Prop35
+ Prop29 --> Prop35
+ Prop34 --> Prop35
+ Prop33 --> Prop36
+ Prop34 --> Prop36
+ Prop35 --> Prop36
+ Prop31 --> Prop37
+ Prop34 --> Prop37
+ Prop35 --> Prop37
+ Prop31 --> Prop38
+ Prop34 --> Prop38
+ Prop36 --> Prop38
+ Prop34 --> Prop41
+ Prop37 --> Prop41
+ Prop10 --> Prop42
+ Prop23 --> Prop42
+ Prop31 --> Prop42
+ Prop38 --> Prop42
+ Prop41 --> Prop42
+ Prop34 --> Prop43
+ Prop15 --> Prop44
+ Prop29 --> Prop44
+ Prop31 --> Prop44
+ Prop42 --> Prop44
+ Prop43 --> Prop44
+ Prop14 --> Prop45
+ Prop29 --> Prop45
+ Prop30 --> Prop45
+ Prop33 --> Prop45
+ Prop34 --> Prop45
+ Prop42 --> Prop45
+ Prop44 --> Prop45
+ Prop3 --> Prop46
+ Prop11 --> Prop46
+ Prop29 --> Prop46
+ Prop31 --> Prop46
+ Prop34 --> Prop46
+ Prop4 --> Prop47
+ Prop14 --> Prop47
+ Prop31 --> Prop47
+ Prop41 --> Prop47
+ Prop46 --> Prop47
+ Prop3 --> Prop48
+ Prop8 --> Prop48
+ Prop11 --> Prop48
+ Prop47 --> Prop48
+ classDef postulate fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef commonNotion fill:#9b59b6,color:#fff,stroke:#8e44ad
+ classDef proposition fill:#1abc9c,color:#fff,stroke:#16a085
+ class P1,P2,P3,P4,P5 postulate
+ class CN1,CN2,CN3,CN4,CN5 commonNotion
+ class Prop1,Prop2,Prop3,Prop4,Prop5,Prop7,Prop8,Prop9,Prop10,Prop11,Prop13,Prop14,Prop15,Prop16,Prop18,Prop19,Prop20,Prop22,Prop23,Prop26,Prop27,Prop29,Prop30,Prop31,Prop33,Prop34,Prop35,Prop36,Prop37,Prop38,Prop41,Prop42,Prop43,Prop44,Prop45,Prop46,Prop47,Prop48 proposition
\ No newline at end of file
diff --git a/data/euclid-elements-book-i.json b/data/euclid-elements-book-i.json
new file mode 100644
index 0000000000000000000000000000000000000000..45d9a093b1e394b4c358067a2c00a1b649b601e4
--- /dev/null
+++ b/data/euclid-elements-book-i.json
@@ -0,0 +1,1167 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-i",
+ "name": "Euclid's Elements, Book I",
+ "subject": "geometry",
+ "variant": "classical",
+ "description": "The 48 propositions of Book I with dependencies on postulates (P1–P5), common notions (CN1–CN5), and prior propositions. Source: David E. Joyce, Clark University.",
+ "structure": {
+ "books": 1,
+ "propositions": 48,
+ "foundationTypes": [
+ "postulate",
+ "commonNotion"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-15",
+ "lastUpdated": "2026-03-15",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book I Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book I",
+ "plane geometry",
+ "constructions",
+ "Pythagorean theorem"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book I",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookI/bookI.html",
+ "notes": "Clark University; dependency table from Guide"
+ },
+ {
+ "id": "euclid-heath",
+ "type": "primary",
+ "authors": "Heath, T.L.",
+ "title": "The Thirteen Books of Euclid's Elements",
+ "year": "1908",
+ "edition": "2nd",
+ "publisher": "Cambridge University Press",
+ "url": "https://archive.org/details/euclidheath00heatiala",
+ "notes": "Standard English translation"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "P1",
+ "type": "postulate",
+ "label": "Draw a straight line from any point to any point",
+ "shortLabel": "Post. 1",
+ "book": 1,
+ "number": 1,
+ "colorClass": "postulate"
+ },
+ {
+ "id": "P2",
+ "type": "postulate",
+ "label": "Produce a finite straight line continuously in a straight line",
+ "shortLabel": "Post. 2",
+ "book": 1,
+ "number": 2,
+ "colorClass": "postulate"
+ },
+ {
+ "id": "P3",
+ "type": "postulate",
+ "label": "Describe a circle with any center and radius",
+ "shortLabel": "Post. 3",
+ "book": 1,
+ "number": 3,
+ "colorClass": "postulate"
+ },
+ {
+ "id": "P4",
+ "type": "postulate",
+ "label": "All right angles equal one another",
+ "shortLabel": "Post. 4",
+ "book": 1,
+ "number": 4,
+ "colorClass": "postulate"
+ },
+ {
+ "id": "P5",
+ "type": "postulate",
+ "label": "Parallel postulate: if interior angles < two right, lines meet",
+ "shortLabel": "Post. 5",
+ "book": 1,
+ "number": 5,
+ "colorClass": "postulate"
+ },
+ {
+ "id": "CN1",
+ "type": "commonNotion",
+ "label": "Things equal to the same thing are equal to each other",
+ "shortLabel": "CN 1",
+ "book": 1,
+ "number": 1,
+ "colorClass": "commonNotion"
+ },
+ {
+ "id": "CN2",
+ "type": "commonNotion",
+ "label": "If equals are added to equals, the wholes are equal",
+ "shortLabel": "CN 2",
+ "book": 1,
+ "number": 2,
+ "colorClass": "commonNotion"
+ },
+ {
+ "id": "CN3",
+ "type": "commonNotion",
+ "label": "If equals are subtracted from equals, the remainders are equal",
+ "shortLabel": "CN 3",
+ "book": 1,
+ "number": 3,
+ "colorClass": "commonNotion"
+ },
+ {
+ "id": "CN4",
+ "type": "commonNotion",
+ "label": "Things coinciding with one another are equal",
+ "shortLabel": "CN 4",
+ "book": 1,
+ "number": 4,
+ "colorClass": "commonNotion"
+ },
+ {
+ "id": "CN5",
+ "type": "commonNotion",
+ "label": "The whole is greater than the part",
+ "shortLabel": "CN 5",
+ "book": 1,
+ "number": 5,
+ "colorClass": "commonNotion"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "To construct an equilateral triangle on a given finite straight line",
+ "shortLabel": "Prop. I.1",
+ "short": "Equilateral triangle on given line",
+ "book": 1,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "To place a straight line equal to a given straight line with one end at a given point",
+ "shortLabel": "Prop. I.2",
+ "short": "Place line equal to given at point",
+ "book": 1,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "To cut off from the greater of two given unequal straight lines a straight line equal to the less",
+ "shortLabel": "Prop. I.3",
+ "short": "Cut off from greater segment equal to less",
+ "book": 1,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "If two triangles have two sides equal to two sides respectively, and the angles contained equal, then bases and remaining angles equal",
+ "shortLabel": "Prop. I.4",
+ "short": "SAS congruence",
+ "book": 1,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "In isosceles triangles the angles at the base equal one another",
+ "shortLabel": "Prop. I.5",
+ "short": "Base angles of isosceles equal",
+ "book": 1,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "If in a triangle two angles equal one another, then the sides opposite the equal angles also equal one another",
+ "shortLabel": "Prop. I.6",
+ "short": "Sides opposite equal angles equal",
+ "book": 1,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "Given two lines from ends of a line meeting at a point, no other such pair from same ends on same side",
+ "shortLabel": "Prop. I.7",
+ "short": "Uniqueness of triangle from ends",
+ "book": 1,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "If two triangles have two sides equal to two sides respectively, and the base equal to the base, then the angles contained are equal",
+ "shortLabel": "Prop. I.8",
+ "short": "SSS congruence",
+ "book": 1,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "To bisect a given rectilinear angle",
+ "shortLabel": "Prop. I.9",
+ "short": "Bisect angle",
+ "book": 1,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "To bisect a given finite straight line",
+ "shortLabel": "Prop. I.10",
+ "short": "Bisect line",
+ "book": 1,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "To draw a straight line at right angles to a given straight line from a given point on it",
+ "shortLabel": "Prop. I.11",
+ "short": "Perpendicular from point on line",
+ "book": 1,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "To draw a straight line perpendicular to a given infinite straight line from a given point not on it",
+ "shortLabel": "Prop. I.12",
+ "short": "Perpendicular from point not on line",
+ "book": 1,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "If a straight line stands on a straight line, it makes either two right angles or angles whose sum equals two right angles",
+ "shortLabel": "Prop. I.13",
+ "short": "Angles on line sum to two right",
+ "book": 1,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "If with any straight line, at a point, two lines not on same side make adjacent angles equal to two right, they are in a straight line",
+ "shortLabel": "Prop. I.14",
+ "short": "If angles sum to two right, straight line",
+ "book": 1,
+ "number": 14,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop15",
+ "type": "proposition",
+ "label": "If two straight lines cut one another, they make the vertical angles equal to one another",
+ "shortLabel": "Prop. I.15",
+ "short": "Vertical angles equal",
+ "book": 1,
+ "number": 15,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop16",
+ "type": "proposition",
+ "label": "In any triangle, if one side is produced, the exterior angle is greater than either interior opposite angle",
+ "shortLabel": "Prop. I.16",
+ "short": "Exterior angle > interior opposite",
+ "book": 1,
+ "number": 16,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop17",
+ "type": "proposition",
+ "label": "In any triangle the sum of any two angles is less than two right angles",
+ "shortLabel": "Prop. I.17",
+ "short": "Sum of two angles < two right",
+ "book": 1,
+ "number": 17,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop18",
+ "type": "proposition",
+ "label": "In any triangle the angle opposite the greater side is greater",
+ "shortLabel": "Prop. I.18",
+ "short": "Angle opposite greater side greater",
+ "book": 1,
+ "number": 18,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop19",
+ "type": "proposition",
+ "label": "In any triangle the side opposite the greater angle is greater",
+ "shortLabel": "Prop. I.19",
+ "short": "Side opposite greater angle greater",
+ "book": 1,
+ "number": 19,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop20",
+ "type": "proposition",
+ "label": "In any triangle the sum of any two sides is greater than the remaining one",
+ "shortLabel": "Prop. I.20",
+ "short": "Triangle inequality",
+ "book": 1,
+ "number": 20,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop21",
+ "type": "proposition",
+ "label": "If from ends of one side two lines meet within the triangle, their sum < sum of other two sides",
+ "shortLabel": "Prop. I.21",
+ "short": "Lines from ends within triangle",
+ "book": 1,
+ "number": 21,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop22",
+ "type": "proposition",
+ "label": "To construct a triangle out of three straight lines which equal three given straight lines",
+ "shortLabel": "Prop. I.22",
+ "short": "Construct triangle from three lines",
+ "book": 1,
+ "number": 22,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop23",
+ "type": "proposition",
+ "label": "To construct a rectilinear angle equal to a given rectilinear angle on a given straight line",
+ "shortLabel": "Prop. I.23",
+ "short": "Construct angle equal to given",
+ "book": 1,
+ "number": 23,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop24",
+ "type": "proposition",
+ "label": "If two triangles have two sides equal but one contained angle greater, the base is greater",
+ "shortLabel": "Prop. I.24",
+ "short": "SAS for greater angle => greater base",
+ "book": 1,
+ "number": 24,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop25",
+ "type": "proposition",
+ "label": "If two triangles have two sides equal but base greater, the contained angle is greater",
+ "shortLabel": "Prop. I.25",
+ "short": "SAS for greater base => greater angle",
+ "book": 1,
+ "number": 25,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop26",
+ "type": "proposition",
+ "label": "If two triangles have two angles equal and one side equal, the remaining sides and angle equal",
+ "shortLabel": "Prop. I.26",
+ "short": "AAS congruence",
+ "book": 1,
+ "number": 26,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop27",
+ "type": "proposition",
+ "label": "If a line falling on two lines makes alternate angles equal, the lines are parallel",
+ "shortLabel": "Prop. I.27",
+ "short": "Alternate angles equal => parallel",
+ "book": 1,
+ "number": 27,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop28",
+ "type": "proposition",
+ "label": "If exterior angle equals interior opposite, or interior same-side sum to two right, lines parallel",
+ "shortLabel": "Prop. I.28",
+ "short": "Exterior = interior opposite => parallel",
+ "book": 1,
+ "number": 28,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop29",
+ "type": "proposition",
+ "label": "A line falling on parallel lines makes alternate angles equal, exterior = interior opposite",
+ "shortLabel": "Prop. I.29",
+ "short": "Parallel => alternate angles equal",
+ "book": 1,
+ "number": 29,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop30",
+ "type": "proposition",
+ "label": "Straight lines parallel to the same straight line are also parallel to one another",
+ "shortLabel": "Prop. I.30",
+ "short": "Transitivity of parallel",
+ "book": 1,
+ "number": 30,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop31",
+ "type": "proposition",
+ "label": "To draw a straight line through a given point parallel to a given straight line",
+ "shortLabel": "Prop. I.31",
+ "short": "Draw parallel through point",
+ "book": 1,
+ "number": 31,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop32",
+ "type": "proposition",
+ "label": "In any triangle, exterior angle equals sum of two interior opposite; three angles = two right",
+ "shortLabel": "Prop. I.32",
+ "short": "Exterior angle = sum interior opposite",
+ "book": 1,
+ "number": 32,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop33",
+ "type": "proposition",
+ "label": "Straight lines which join the ends of equal and parallel straight lines in same directions are equal and parallel",
+ "shortLabel": "Prop. I.33",
+ "short": "Joining ends of equal parallel lines",
+ "book": 1,
+ "number": 33,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop34",
+ "type": "proposition",
+ "label": "In parallelogrammic areas the opposite sides and angles equal one another, diameter bisects",
+ "shortLabel": "Prop. I.34",
+ "short": "Parallelogram properties",
+ "book": 1,
+ "number": 34,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop35",
+ "type": "proposition",
+ "label": "Parallelograms which are on the same base and in the same parallels equal one another",
+ "shortLabel": "Prop. I.35",
+ "short": "Parallelograms same base equal",
+ "book": 1,
+ "number": 35,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop36",
+ "type": "proposition",
+ "label": "Parallelograms which are on equal bases and in the same parallels equal one another",
+ "shortLabel": "Prop. I.36",
+ "short": "Parallelograms equal bases equal",
+ "book": 1,
+ "number": 36,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop37",
+ "type": "proposition",
+ "label": "Triangles which are on the same base and in the same parallels equal one another",
+ "shortLabel": "Prop. I.37",
+ "short": "Triangles same base equal",
+ "book": 1,
+ "number": 37,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop38",
+ "type": "proposition",
+ "label": "Triangles which are on equal bases and in the same parallels equal one another",
+ "shortLabel": "Prop. I.38",
+ "short": "Triangles equal bases equal",
+ "book": 1,
+ "number": 38,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop39",
+ "type": "proposition",
+ "label": "Equal triangles on same base and same side are in the same parallels",
+ "shortLabel": "Prop. I.39",
+ "short": "Equal triangles same base same side",
+ "book": 1,
+ "number": 39,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop40",
+ "type": "proposition",
+ "label": "Equal triangles on equal bases and same side are in the same parallels",
+ "shortLabel": "Prop. I.40",
+ "short": "Equal triangles equal bases same side",
+ "book": 1,
+ "number": 40,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop41",
+ "type": "proposition",
+ "label": "If a parallelogram has same base with triangle and same parallels, parallelogram is double the triangle",
+ "shortLabel": "Prop. I.41",
+ "short": "Parallelogram = 2× triangle",
+ "book": 1,
+ "number": 41,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop42",
+ "type": "proposition",
+ "label": "To construct a parallelogram equal to a given triangle in a given rectilinear angle",
+ "shortLabel": "Prop. I.42",
+ "short": "Construct parallelogram = triangle",
+ "book": 1,
+ "number": 42,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop43",
+ "type": "proposition",
+ "label": "In any parallelogram the complements of the parallelograms about the diameter equal one another",
+ "shortLabel": "Prop. I.43",
+ "short": "Complements of parallelogram",
+ "book": 1,
+ "number": 43,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop44",
+ "type": "proposition",
+ "label": "To a given straight line in a given angle, to apply a parallelogram equal to a given triangle",
+ "shortLabel": "Prop. I.44",
+ "short": "Apply parallelogram to line",
+ "book": 1,
+ "number": 44,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop45",
+ "type": "proposition",
+ "label": "To construct a parallelogram equal to a given rectilinear figure in a given rectilinear angle",
+ "shortLabel": "Prop. I.45",
+ "short": "Construct parallelogram = rectilinear figure",
+ "book": 1,
+ "number": 45,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop46",
+ "type": "proposition",
+ "label": "To describe a square on a given straight line",
+ "shortLabel": "Prop. I.46",
+ "short": "Construct square on line",
+ "book": 1,
+ "number": 46,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop47",
+ "type": "proposition",
+ "label": "In right-angled triangles the square on the side opposite the right angle equals the sum of the squares on the sides containing the right angle",
+ "shortLabel": "Prop. I.47",
+ "short": "Pythagorean theorem",
+ "book": 1,
+ "number": 47,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop48",
+ "type": "proposition",
+ "label": "If in a triangle the square on one side equals the sum of squares on the other two, the angle contained by those sides is right",
+ "shortLabel": "Prop. I.48",
+ "short": "Converse Pythagorean",
+ "book": 1,
+ "number": 48,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "P1",
+ "to": "Prop1"
+ },
+ {
+ "from": "P3",
+ "to": "Prop1"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop2"
+ },
+ {
+ "from": "P1",
+ "to": "Prop2"
+ },
+ {
+ "from": "P2",
+ "to": "Prop2"
+ },
+ {
+ "from": "P3",
+ "to": "Prop2"
+ },
+ {
+ "from": "Prop2",
+ "to": "Prop3"
+ },
+ {
+ "from": "P3",
+ "to": "Prop3"
+ },
+ {
+ "from": "CN4",
+ "to": "Prop4"
+ },
+ {
+ "from": "CN5",
+ "to": "Prop4"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop5"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop5"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop6"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop6"
+ },
+ {
+ "from": "Prop5",
+ "to": "Prop7"
+ },
+ {
+ "from": "Prop7",
+ "to": "Prop8"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop9"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop9"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop9"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop10"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop10"
+ },
+ {
+ "from": "Prop9",
+ "to": "Prop10"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop11"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop11"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop11"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop12"
+ },
+ {
+ "from": "Prop10",
+ "to": "Prop12"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop13"
+ },
+ {
+ "from": "Prop13",
+ "to": "Prop14"
+ },
+ {
+ "from": "Prop13",
+ "to": "Prop15"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop16"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop16"
+ },
+ {
+ "from": "Prop10",
+ "to": "Prop16"
+ },
+ {
+ "from": "Prop15",
+ "to": "Prop16"
+ },
+ {
+ "from": "Prop13",
+ "to": "Prop17"
+ },
+ {
+ "from": "Prop16",
+ "to": "Prop17"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop18"
+ },
+ {
+ "from": "Prop5",
+ "to": "Prop18"
+ },
+ {
+ "from": "Prop16",
+ "to": "Prop18"
+ },
+ {
+ "from": "Prop5",
+ "to": "Prop19"
+ },
+ {
+ "from": "Prop18",
+ "to": "Prop19"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop20"
+ },
+ {
+ "from": "Prop5",
+ "to": "Prop20"
+ },
+ {
+ "from": "Prop19",
+ "to": "Prop20"
+ },
+ {
+ "from": "Prop16",
+ "to": "Prop21"
+ },
+ {
+ "from": "Prop20",
+ "to": "Prop21"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop22"
+ },
+ {
+ "from": "Prop20",
+ "to": "Prop22"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop23"
+ },
+ {
+ "from": "Prop22",
+ "to": "Prop23"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop24"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop24"
+ },
+ {
+ "from": "Prop5",
+ "to": "Prop24"
+ },
+ {
+ "from": "Prop19",
+ "to": "Prop24"
+ },
+ {
+ "from": "Prop23",
+ "to": "Prop24"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop25"
+ },
+ {
+ "from": "Prop24",
+ "to": "Prop25"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop26"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop26"
+ },
+ {
+ "from": "Prop16",
+ "to": "Prop26"
+ },
+ {
+ "from": "Prop16",
+ "to": "Prop27"
+ },
+ {
+ "from": "Prop13",
+ "to": "Prop28"
+ },
+ {
+ "from": "Prop15",
+ "to": "Prop28"
+ },
+ {
+ "from": "Prop27",
+ "to": "Prop28"
+ },
+ {
+ "from": "Prop13",
+ "to": "Prop29"
+ },
+ {
+ "from": "Prop15",
+ "to": "Prop29"
+ },
+ {
+ "from": "Prop27",
+ "to": "Prop29"
+ },
+ {
+ "from": "P5",
+ "to": "Prop29"
+ },
+ {
+ "from": "Prop29",
+ "to": "Prop30"
+ },
+ {
+ "from": "Prop23",
+ "to": "Prop31"
+ },
+ {
+ "from": "Prop27",
+ "to": "Prop31"
+ },
+ {
+ "from": "Prop13",
+ "to": "Prop32"
+ },
+ {
+ "from": "Prop29",
+ "to": "Prop32"
+ },
+ {
+ "from": "Prop31",
+ "to": "Prop32"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop33"
+ },
+ {
+ "from": "Prop27",
+ "to": "Prop33"
+ },
+ {
+ "from": "Prop29",
+ "to": "Prop33"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop34"
+ },
+ {
+ "from": "Prop26",
+ "to": "Prop34"
+ },
+ {
+ "from": "Prop29",
+ "to": "Prop34"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop35"
+ },
+ {
+ "from": "Prop29",
+ "to": "Prop35"
+ },
+ {
+ "from": "Prop34",
+ "to": "Prop35"
+ },
+ {
+ "from": "Prop33",
+ "to": "Prop36"
+ },
+ {
+ "from": "Prop34",
+ "to": "Prop36"
+ },
+ {
+ "from": "Prop35",
+ "to": "Prop36"
+ },
+ {
+ "from": "Prop31",
+ "to": "Prop37"
+ },
+ {
+ "from": "Prop34",
+ "to": "Prop37"
+ },
+ {
+ "from": "Prop35",
+ "to": "Prop37"
+ },
+ {
+ "from": "Prop31",
+ "to": "Prop38"
+ },
+ {
+ "from": "Prop34",
+ "to": "Prop38"
+ },
+ {
+ "from": "Prop36",
+ "to": "Prop38"
+ },
+ {
+ "from": "Prop31",
+ "to": "Prop39"
+ },
+ {
+ "from": "Prop37",
+ "to": "Prop39"
+ },
+ {
+ "from": "Prop31",
+ "to": "Prop40"
+ },
+ {
+ "from": "Prop38",
+ "to": "Prop40"
+ },
+ {
+ "from": "Prop34",
+ "to": "Prop41"
+ },
+ {
+ "from": "Prop37",
+ "to": "Prop41"
+ },
+ {
+ "from": "Prop10",
+ "to": "Prop42"
+ },
+ {
+ "from": "Prop23",
+ "to": "Prop42"
+ },
+ {
+ "from": "Prop31",
+ "to": "Prop42"
+ },
+ {
+ "from": "Prop38",
+ "to": "Prop42"
+ },
+ {
+ "from": "Prop41",
+ "to": "Prop42"
+ },
+ {
+ "from": "Prop34",
+ "to": "Prop43"
+ },
+ {
+ "from": "Prop15",
+ "to": "Prop44"
+ },
+ {
+ "from": "Prop29",
+ "to": "Prop44"
+ },
+ {
+ "from": "Prop31",
+ "to": "Prop44"
+ },
+ {
+ "from": "Prop42",
+ "to": "Prop44"
+ },
+ {
+ "from": "Prop43",
+ "to": "Prop44"
+ },
+ {
+ "from": "Prop14",
+ "to": "Prop45"
+ },
+ {
+ "from": "Prop29",
+ "to": "Prop45"
+ },
+ {
+ "from": "Prop30",
+ "to": "Prop45"
+ },
+ {
+ "from": "Prop33",
+ "to": "Prop45"
+ },
+ {
+ "from": "Prop34",
+ "to": "Prop45"
+ },
+ {
+ "from": "Prop42",
+ "to": "Prop45"
+ },
+ {
+ "from": "Prop44",
+ "to": "Prop45"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop46"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop46"
+ },
+ {
+ "from": "Prop29",
+ "to": "Prop46"
+ },
+ {
+ "from": "Prop31",
+ "to": "Prop46"
+ },
+ {
+ "from": "Prop34",
+ "to": "Prop46"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop47"
+ },
+ {
+ "from": "Prop14",
+ "to": "Prop47"
+ },
+ {
+ "from": "Prop31",
+ "to": "Prop47"
+ },
+ {
+ "from": "Prop41",
+ "to": "Prop47"
+ },
+ {
+ "from": "Prop46",
+ "to": "Prop47"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop48"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop48"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop48"
+ },
+ {
+ "from": "Prop47",
+ "to": "Prop48"
+ }
+ ],
+ "colorScheme": {
+ "postulate": {
+ "fill": "#e74c3c",
+ "stroke": "#c0392b"
+ },
+ "commonNotion": {
+ "fill": "#9b59b6",
+ "stroke": "#8e44ad"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/euclid-elements-book-i.mmd b/data/euclid-elements-book-i.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..51c423a56f6fe7ba6f2d88f7dcbee570974d4861
--- /dev/null
+++ b/data/euclid-elements-book-i.mmd
@@ -0,0 +1,195 @@
+graph TD
+ P1["Post. 1\nDraw a straight line from any po..."]
+ P2["Post. 2\nProduce a finite straight line c..."]
+ P3["Post. 3\nDescribe a circle with any cente..."]
+ P4["Post. 4\nAll right angles equal one another"]
+ P5["Post. 5\nParallel postulate: if interior ..."]
+ CN1["CN 1\nThings equal to the same thing a..."]
+ CN2["CN 2\nIf equals are added to equals, t..."]
+ CN3["CN 3\nIf equals are subtracted from eq..."]
+ CN4["CN 4\nThings coinciding with one anoth..."]
+ CN5["CN 5\nThe whole is greater than the part"]
+ Prop1["Prop. I.1\nEquilateral triangle on given line"]
+ Prop2["Prop. I.2\nPlace line equal to given at point"]
+ Prop3["Prop. I.3\nCut off from greater segment equal to less"]
+ Prop4["Prop. I.4\nSAS congruence"]
+ Prop5["Prop. I.5\nBase angles of isosceles equal"]
+ Prop6["Prop. I.6\nSides opposite equal angles equal"]
+ Prop7["Prop. I.7\nUniqueness of triangle from ends"]
+ Prop8["Prop. I.8\nSSS congruence"]
+ Prop9["Prop. I.9\nBisect angle"]
+ Prop10["Prop. I.10\nBisect line"]
+ Prop11["Prop. I.11\nPerpendicular from point on line"]
+ Prop12["Prop. I.12\nPerpendicular from point not on line"]
+ Prop13["Prop. I.13\nAngles on line sum to two right"]
+ Prop14["Prop. I.14\nIf angles sum to two right, straight line"]
+ Prop15["Prop. I.15\nVertical angles equal"]
+ Prop16["Prop. I.16\nExterior angle > interior opposite"]
+ Prop17["Prop. I.17\nSum of two angles < two right"]
+ Prop18["Prop. I.18\nAngle opposite greater side greater"]
+ Prop19["Prop. I.19\nSide opposite greater angle greater"]
+ Prop20["Prop. I.20\nTriangle inequality"]
+ Prop21["Prop. I.21\nLines from ends within triangle"]
+ Prop22["Prop. I.22\nConstruct triangle from three lines"]
+ Prop23["Prop. I.23\nConstruct angle equal to given"]
+ Prop24["Prop. I.24\nSAS for greater angle => greater base"]
+ Prop25["Prop. I.25\nSAS for greater base => greater angle"]
+ Prop26["Prop. I.26\nAAS congruence"]
+ Prop27["Prop. I.27\nAlternate angles equal => parallel"]
+ Prop28["Prop. I.28\nExterior = interior opposite => parallel"]
+ Prop29["Prop. I.29\nParallel => alternate angles equal"]
+ Prop30["Prop. I.30\nTransitivity of parallel"]
+ Prop31["Prop. I.31\nDraw parallel through point"]
+ Prop32["Prop. I.32\nExterior angle = sum interior opposite"]
+ Prop33["Prop. I.33\nJoining ends of equal parallel lines"]
+ Prop34["Prop. I.34\nParallelogram properties"]
+ Prop35["Prop. I.35\nParallelograms same base equal"]
+ Prop36["Prop. I.36\nParallelograms equal bases equal"]
+ Prop37["Prop. I.37\nTriangles same base equal"]
+ Prop38["Prop. I.38\nTriangles equal bases equal"]
+ Prop39["Prop. I.39\nEqual triangles same base same side"]
+ Prop40["Prop. I.40\nEqual triangles equal bases same side"]
+ Prop41["Prop. I.41\nParallelogram = 2× triangle"]
+ Prop42["Prop. I.42\nConstruct parallelogram = triangle"]
+ Prop43["Prop. I.43\nComplements of parallelogram"]
+ Prop44["Prop. I.44\nApply parallelogram to line"]
+ Prop45["Prop. I.45\nConstruct parallelogram = rectilinear figure"]
+ Prop46["Prop. I.46\nConstruct square on line"]
+ Prop47["Prop. I.47\nPythagorean theorem"]
+ Prop48["Prop. I.48\nConverse Pythagorean"]
+ P1 --> Prop1
+ P3 --> Prop1
+ Prop1 --> Prop2
+ P1 --> Prop2
+ P2 --> Prop2
+ P3 --> Prop2
+ Prop2 --> Prop3
+ P3 --> Prop3
+ CN4 --> Prop4
+ CN5 --> Prop4
+ Prop3 --> Prop5
+ Prop4 --> Prop5
+ Prop3 --> Prop6
+ Prop4 --> Prop6
+ Prop5 --> Prop7
+ Prop7 --> Prop8
+ Prop1 --> Prop9
+ Prop3 --> Prop9
+ Prop8 --> Prop9
+ Prop1 --> Prop10
+ Prop4 --> Prop10
+ Prop9 --> Prop10
+ Prop1 --> Prop11
+ Prop3 --> Prop11
+ Prop8 --> Prop11
+ Prop8 --> Prop12
+ Prop10 --> Prop12
+ Prop11 --> Prop13
+ Prop13 --> Prop14
+ Prop13 --> Prop15
+ Prop3 --> Prop16
+ Prop4 --> Prop16
+ Prop10 --> Prop16
+ Prop15 --> Prop16
+ Prop13 --> Prop17
+ Prop16 --> Prop17
+ Prop3 --> Prop18
+ Prop5 --> Prop18
+ Prop16 --> Prop18
+ Prop5 --> Prop19
+ Prop18 --> Prop19
+ Prop3 --> Prop20
+ Prop5 --> Prop20
+ Prop19 --> Prop20
+ Prop16 --> Prop21
+ Prop20 --> Prop21
+ Prop3 --> Prop22
+ Prop20 --> Prop22
+ Prop8 --> Prop23
+ Prop22 --> Prop23
+ Prop3 --> Prop24
+ Prop4 --> Prop24
+ Prop5 --> Prop24
+ Prop19 --> Prop24
+ Prop23 --> Prop24
+ Prop4 --> Prop25
+ Prop24 --> Prop25
+ Prop3 --> Prop26
+ Prop4 --> Prop26
+ Prop16 --> Prop26
+ Prop16 --> Prop27
+ Prop13 --> Prop28
+ Prop15 --> Prop28
+ Prop27 --> Prop28
+ Prop13 --> Prop29
+ Prop15 --> Prop29
+ Prop27 --> Prop29
+ P5 --> Prop29
+ Prop29 --> Prop30
+ Prop23 --> Prop31
+ Prop27 --> Prop31
+ Prop13 --> Prop32
+ Prop29 --> Prop32
+ Prop31 --> Prop32
+ Prop4 --> Prop33
+ Prop27 --> Prop33
+ Prop29 --> Prop33
+ Prop4 --> Prop34
+ Prop26 --> Prop34
+ Prop29 --> Prop34
+ Prop4 --> Prop35
+ Prop29 --> Prop35
+ Prop34 --> Prop35
+ Prop33 --> Prop36
+ Prop34 --> Prop36
+ Prop35 --> Prop36
+ Prop31 --> Prop37
+ Prop34 --> Prop37
+ Prop35 --> Prop37
+ Prop31 --> Prop38
+ Prop34 --> Prop38
+ Prop36 --> Prop38
+ Prop31 --> Prop39
+ Prop37 --> Prop39
+ Prop31 --> Prop40
+ Prop38 --> Prop40
+ Prop34 --> Prop41
+ Prop37 --> Prop41
+ Prop10 --> Prop42
+ Prop23 --> Prop42
+ Prop31 --> Prop42
+ Prop38 --> Prop42
+ Prop41 --> Prop42
+ Prop34 --> Prop43
+ Prop15 --> Prop44
+ Prop29 --> Prop44
+ Prop31 --> Prop44
+ Prop42 --> Prop44
+ Prop43 --> Prop44
+ Prop14 --> Prop45
+ Prop29 --> Prop45
+ Prop30 --> Prop45
+ Prop33 --> Prop45
+ Prop34 --> Prop45
+ Prop42 --> Prop45
+ Prop44 --> Prop45
+ Prop3 --> Prop46
+ Prop11 --> Prop46
+ Prop29 --> Prop46
+ Prop31 --> Prop46
+ Prop34 --> Prop46
+ Prop4 --> Prop47
+ Prop14 --> Prop47
+ Prop31 --> Prop47
+ Prop41 --> Prop47
+ Prop46 --> Prop47
+ Prop3 --> Prop48
+ Prop8 --> Prop48
+ Prop11 --> Prop48
+ Prop47 --> Prop48
+ classDef postulate fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef commonNotion fill:#9b59b6,color:#fff,stroke:#8e44ad
+ classDef proposition fill:#1abc9c,color:#fff,stroke:#16a085
+ class P1,P2,P3,P4,P5 postulate
+ class CN1,CN2,CN3,CN4,CN5 commonNotion
+ class Prop1,Prop2,Prop3,Prop4,Prop5,Prop6,Prop7,Prop8,Prop9,Prop10,Prop11,Prop12,Prop13,Prop14,Prop15,Prop16,Prop17,Prop18,Prop19,Prop20,Prop21,Prop22,Prop23,Prop24,Prop25,Prop26,Prop27,Prop28,Prop29,Prop30,Prop31,Prop32,Prop33,Prop34,Prop35,Prop36,Prop37,Prop38,Prop39,Prop40,Prop41,Prop42,Prop43,Prop44,Prop45,Prop46,Prop47,Prop48 proposition
\ No newline at end of file
diff --git a/data/euclid-elements-book-ii.json b/data/euclid-elements-book-ii.json
new file mode 100644
index 0000000000000000000000000000000000000000..34cf6714bb3744e0dad8d6389d25643b8a4c78eb
--- /dev/null
+++ b/data/euclid-elements-book-ii.json
@@ -0,0 +1,365 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-ii",
+ "name": "Euclid's Elements, Book II",
+ "subject": "geometry",
+ "variant": "classical",
+ "description": "The 14 propositions of Book II on geometric algebra (rectangles, squares). Props 1-10 are logically independent within Book II; 11-14 depend on 6, 4, 7, 5. All depend on Book I. Source: David E. Joyce, Clark University.",
+ "structure": {
+ "books": 2,
+ "propositions": 14,
+ "foundationTypes": [
+ "definition"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-15",
+ "lastUpdated": "2026-03-15",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book II Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book II",
+ "geometric algebra",
+ "rectangles",
+ "squares",
+ "golden section",
+ "quadrature"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book II",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookII/bookII.html",
+ "notes": "Clark University; dependency table from Logical structure"
+ },
+ {
+ "id": "euclid-heath",
+ "type": "primary",
+ "authors": "Heath, T.L.",
+ "title": "The Thirteen Books of Euclid's Elements",
+ "year": "1908",
+ "edition": "2nd",
+ "publisher": "Cambridge University Press",
+ "url": "https://archive.org/details/euclidheath00heatiala",
+ "notes": "Standard English translation"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "BookI",
+ "type": "foundation",
+ "label": "Book I — Fundamentals of plane geometry",
+ "shortLabel": "Book I",
+ "short": "Foundation",
+ "book": 1,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "Def1",
+ "type": "definition",
+ "label": "Rectangle contained by two straight lines containing the right angle",
+ "shortLabel": "Def. II.1",
+ "book": 2,
+ "number": 1,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def2",
+ "type": "definition",
+ "label": "Gnomon: parallelogram about diameter with two complements",
+ "shortLabel": "Def. II.2",
+ "book": 2,
+ "number": 2,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "If one line is cut into segments, rectangle by whole equals sum of rectangles by each segment",
+ "shortLabel": "Prop. II.1",
+ "short": "Rectangle = sum of rectangles",
+ "book": 2,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "If a line is cut at random, sum of rectangles by whole and each segment equals square on whole",
+ "shortLabel": "Prop. II.2",
+ "short": "Sum of rectangles = square on whole",
+ "book": 2,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "If a line is cut at random, rectangle by whole and one segment equals rectangle by segments plus square on that segment",
+ "shortLabel": "Prop. II.3",
+ "short": "Rectangle = rectangle + square",
+ "book": 2,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "If a line is cut at random, square on whole equals squares on segments plus twice rectangle contained by segments",
+ "shortLabel": "Prop. II.4",
+ "short": "Square on whole = squares + 2×rectangle",
+ "book": 2,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "If a line cut into equal and unequal segments, rectangle by unequal segments plus square on difference equals square on half",
+ "shortLabel": "Prop. II.5",
+ "short": "Unequal segments: rectangle + square = square on half",
+ "book": 2,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "If a line bisected and added to, rectangle by whole-with-added and added plus square on half equals square on half-plus-added",
+ "shortLabel": "Prop. II.6",
+ "short": "Bisected + added: rectangle + square = square",
+ "book": 2,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "If a line cut at random, square on whole plus square on one segment equals twice rectangle by whole and segment plus square on remainder",
+ "shortLabel": "Prop. II.7",
+ "short": "Square on whole + square on segment",
+ "book": 2,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "If a line cut at random, four times rectangle by whole and one segment plus square on remainder equals square on whole-plus-segment",
+ "shortLabel": "Prop. II.8",
+ "short": "Four times rectangle + square",
+ "book": 2,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "If a line cut into equal and unequal segments, sum of squares on unequal segments is double sum of square on half and square on difference",
+ "shortLabel": "Prop. II.9",
+ "short": "Unequal segments: sum of squares",
+ "book": 2,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "If a line bisected and added to, square on whole-with-added plus square on added equals double sum of square on half and square on half-plus-added",
+ "shortLabel": "Prop. II.10",
+ "short": "Bisected + added: sum of squares",
+ "book": 2,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "To cut a given line so that rectangle by whole and one segment equals square on remaining segment",
+ "shortLabel": "Prop. II.11",
+ "short": "Cut line: rectangle = square (golden section)",
+ "book": 2,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "In obtuse-angled triangles, square on side opposite obtuse angle greater than sum of squares on sides containing it",
+ "shortLabel": "Prop. II.12",
+ "short": "Obtuse triangle: law of cosines",
+ "book": 2,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "In acute-angled triangles, square on side opposite acute angle less than sum of squares on sides containing it",
+ "shortLabel": "Prop. II.13",
+ "short": "Acute triangle: law of cosines",
+ "book": 2,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "To construct a square equal to a given rectilinear figure",
+ "shortLabel": "Prop. II.14",
+ "short": "Construct square = rectilinear figure",
+ "book": 2,
+ "number": 14,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "BookI",
+ "to": "Prop1"
+ },
+ {
+ "from": "Def1",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop2"
+ },
+ {
+ "from": "Def1",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop3"
+ },
+ {
+ "from": "Def1",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop4"
+ },
+ {
+ "from": "Def1",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop5"
+ },
+ {
+ "from": "Def1",
+ "to": "Prop5"
+ },
+ {
+ "from": "Def2",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop6"
+ },
+ {
+ "from": "Def1",
+ "to": "Prop6"
+ },
+ {
+ "from": "Def2",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop7"
+ },
+ {
+ "from": "Def1",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop8"
+ },
+ {
+ "from": "Def1",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop9"
+ },
+ {
+ "from": "Def1",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop10"
+ },
+ {
+ "from": "Def1",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop11"
+ },
+ {
+ "from": "Prop6",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop12"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop13"
+ },
+ {
+ "from": "Prop7",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop14"
+ },
+ {
+ "from": "Prop5",
+ "to": "Prop14"
+ }
+ ],
+ "colorScheme": {
+ "foundation": {
+ "fill": "#95a5a6",
+ "stroke": "#7f8c8d"
+ },
+ "definition": {
+ "fill": "#3498db",
+ "stroke": "#2980b9"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/euclid-elements-book-ii.mmd b/data/euclid-elements-book-ii.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..500bc6377f72a880912a642808ab6120a4584e78
--- /dev/null
+++ b/data/euclid-elements-book-ii.mmd
@@ -0,0 +1,54 @@
+graph TD
+ BookI["Book I\nFoundation"]
+ Def1["Def. II.1\nRectangle contained by two strai..."]
+ Def2["Def. II.2\nGnomon: parallelogram about diam..."]
+ Prop1["Prop. II.1\nRectangle = sum of rectangles"]
+ Prop2["Prop. II.2\nSum of rectangles = square on whole"]
+ Prop3["Prop. II.3\nRectangle = rectangle + square"]
+ Prop4["Prop. II.4\nSquare on whole = squares + 2×rectangle"]
+ Prop5["Prop. II.5\nUnequal segments: rectangle + square = square on half"]
+ Prop6["Prop. II.6\nBisected + added: rectangle + square = square"]
+ Prop7["Prop. II.7\nSquare on whole + square on segment"]
+ Prop8["Prop. II.8\nFour times rectangle + square"]
+ Prop9["Prop. II.9\nUnequal segments: sum of squares"]
+ Prop10["Prop. II.10\nBisected + added: sum of squares"]
+ Prop11["Prop. II.11\nCut line: rectangle = square (golden section)"]
+ Prop12["Prop. II.12\nObtuse triangle: law of cosines"]
+ Prop13["Prop. II.13\nAcute triangle: law of cosines"]
+ Prop14["Prop. II.14\nConstruct square = rectilinear figure"]
+ BookI --> Prop1
+ Def1 --> Prop1
+ BookI --> Prop2
+ Def1 --> Prop2
+ BookI --> Prop3
+ Def1 --> Prop3
+ BookI --> Prop4
+ Def1 --> Prop4
+ BookI --> Prop5
+ Def1 --> Prop5
+ Def2 --> Prop5
+ BookI --> Prop6
+ Def1 --> Prop6
+ Def2 --> Prop6
+ BookI --> Prop7
+ Def1 --> Prop7
+ BookI --> Prop8
+ Def1 --> Prop8
+ BookI --> Prop9
+ Def1 --> Prop9
+ BookI --> Prop10
+ Def1 --> Prop10
+ BookI --> Prop11
+ Prop6 --> Prop11
+ BookI --> Prop12
+ Prop4 --> Prop12
+ BookI --> Prop13
+ Prop7 --> Prop13
+ BookI --> Prop14
+ Prop5 --> Prop14
+ classDef foundation fill:#95a5a6,color:#fff,stroke:#7f8c8d
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef proposition fill:#1abc9c,color:#fff,stroke:#16a085
+ class BookI foundation
+ class Def1,Def2 definition
+ class Prop1,Prop2,Prop3,Prop4,Prop5,Prop6,Prop7,Prop8,Prop9,Prop10,Prop11,Prop12,Prop13,Prop14 proposition
\ No newline at end of file
diff --git a/data/euclid-elements-book-iii.json b/data/euclid-elements-book-iii.json
new file mode 100644
index 0000000000000000000000000000000000000000..de13968cebbb2331de37966710bd826c4b547ba7
--- /dev/null
+++ b/data/euclid-elements-book-iii.json
@@ -0,0 +1,885 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-iii",
+ "name": "Euclid's Elements, Book III",
+ "subject": "geometry",
+ "variant": "classical",
+ "description": "Theory of circles: 11 definitions, 37 propositions. All depend on Book I. III.35 uses II.5. Source: David E. Joyce.",
+ "structure": {
+ "books": 3,
+ "definitions": 11,
+ "propositions": 37,
+ "foundationTypes": [
+ "definition",
+ "foundation"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-15",
+ "lastUpdated": "2026-03-15",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book III Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book III",
+ "circles",
+ "chords",
+ "tangents"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book III",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookIII/bookIII.html",
+ "notes": "Clark University"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "BookI",
+ "type": "foundation",
+ "label": "Book I — Fundamentals of plane geometry",
+ "shortLabel": "Book I",
+ "short": "Foundation",
+ "book": 1,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "PropII5",
+ "type": "foundation",
+ "label": "Prop. II.5 — Rectangle + square = square on half",
+ "shortLabel": "Prop. II.5",
+ "short": "From Book II",
+ "book": 2,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "Def1",
+ "type": "definition",
+ "label": "Equal circles are those with equal radii",
+ "shortLabel": "Def. III.1",
+ "short": "Equal circles",
+ "book": 3,
+ "number": 1,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def2",
+ "type": "definition",
+ "label": "A straight line touches a circle if it meets but does not cut it",
+ "shortLabel": "Def. III.2",
+ "short": "Tangent",
+ "book": 3,
+ "number": 2,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def3",
+ "type": "definition",
+ "label": "Circles touch one another if they meet but do not cut",
+ "shortLabel": "Def. III.3",
+ "short": "Circles touching",
+ "book": 3,
+ "number": 3,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def4",
+ "type": "definition",
+ "label": "Lines equally distant from center when perpendiculars from center equal",
+ "shortLabel": "Def. III.4",
+ "short": "Equally distant from center",
+ "book": 3,
+ "number": 4,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def5",
+ "type": "definition",
+ "label": "Greater distance when greater perpendicular falls",
+ "shortLabel": "Def. III.5",
+ "short": "Greater distance",
+ "book": 3,
+ "number": 5,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def6",
+ "type": "definition",
+ "label": "Segment of circle: figure contained by straight line and circumference",
+ "shortLabel": "Def. III.6",
+ "short": "Segment of circle",
+ "book": 3,
+ "number": 6,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def7",
+ "type": "definition",
+ "label": "Angle of segment: contained by straight line and circumference",
+ "shortLabel": "Def. III.7",
+ "short": "Angle of segment",
+ "book": 3,
+ "number": 7,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def8",
+ "type": "definition",
+ "label": "Angle in segment: contained by straight lines joining circumference",
+ "shortLabel": "Def. III.8",
+ "short": "Angle in segment",
+ "book": 3,
+ "number": 8,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def9",
+ "type": "definition",
+ "label": "Angle stands on circumference when lines cut off that circumference",
+ "shortLabel": "Def. III.9",
+ "short": "Angle stands on circumference",
+ "book": 3,
+ "number": 9,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def10",
+ "type": "definition",
+ "label": "Sector: figure contained by two radii and circumference between them",
+ "shortLabel": "Def. III.10",
+ "short": "Sector",
+ "book": 3,
+ "number": 10,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def11",
+ "type": "definition",
+ "label": "Similar segments are those which admit equal angles",
+ "shortLabel": "Def. III.11",
+ "short": "Similar segments",
+ "book": 3,
+ "number": 11,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "To find the center of a given circle",
+ "shortLabel": "Prop. III.1",
+ "short": "Find center of circle",
+ "book": 3,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "Straight line joining two points on circumference falls within circle",
+ "shortLabel": "Prop. III.2",
+ "short": "Chord falls within circle",
+ "book": 3,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "If diameter bisects chord not through center, it cuts at right angles",
+ "shortLabel": "Prop. III.3",
+ "short": "Diameter bisects chord at right angles",
+ "book": 3,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "Two non-diameters cutting one another do not bisect",
+ "shortLabel": "Prop. III.4",
+ "short": "Non-diameters do not bisect",
+ "book": 3,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "If two circles cut one another, they do not have same center",
+ "shortLabel": "Prop. III.5",
+ "short": "Cutting circles do not share center",
+ "book": 3,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "If two circles touch, they do not have same center",
+ "shortLabel": "Prop. III.6",
+ "short": "Touching circles do not share center",
+ "book": 3,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "From point on diameter: greatest through center, least is remainder",
+ "shortLabel": "Prop. III.7",
+ "short": "Greatest/shortest from point on diameter",
+ "book": 3,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "From point outside: through center greatest; between point and diameter least",
+ "shortLabel": "Prop. III.8",
+ "short": "Lines from point outside circle",
+ "book": 3,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "If more than two equal lines fall from point on circle, point is center",
+ "shortLabel": "Prop. III.9",
+ "short": "Three equal lines imply center",
+ "book": 3,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "A circle does not cut another at more than two points",
+ "shortLabel": "Prop. III.10",
+ "short": "Circles cut at most two points",
+ "book": 3,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "Line joining centers of internally touching circles passes through contact",
+ "shortLabel": "Prop. III.11",
+ "short": "Internally touching circles",
+ "book": 3,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "Line joining centers of externally touching circles passes through contact",
+ "shortLabel": "Prop. III.12",
+ "short": "Externally touching circles",
+ "book": 3,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "Circle does not touch another at more than one point",
+ "shortLabel": "Prop. III.13",
+ "short": "Circles touch at most one point",
+ "book": 3,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "Equal chords equally distant from center, and conversely",
+ "shortLabel": "Prop. III.14",
+ "short": "Equal chords equally distant",
+ "book": 3,
+ "number": 14,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop15",
+ "type": "proposition",
+ "label": "Diameter greatest; nearer to center greater than more remote",
+ "shortLabel": "Prop. III.15",
+ "short": "Diameter greatest",
+ "book": 3,
+ "number": 15,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop16",
+ "type": "proposition",
+ "label": "Perpendicular at end of diameter falls outside; horn angle",
+ "shortLabel": "Prop. III.16",
+ "short": "Tangent at end of diameter",
+ "book": 3,
+ "number": 16,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop17",
+ "type": "proposition",
+ "label": "From given point to draw straight line touching given circle",
+ "shortLabel": "Prop. III.17",
+ "short": "Draw tangent from point",
+ "book": 3,
+ "number": 17,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop18",
+ "type": "proposition",
+ "label": "Radius to point of contact perpendicular to tangent",
+ "shortLabel": "Prop. III.18",
+ "short": "Radius to tangent perpendicular",
+ "book": 3,
+ "number": 18,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop19",
+ "type": "proposition",
+ "label": "Perpendicular from contact to tangent passes through center",
+ "shortLabel": "Prop. III.19",
+ "short": "Perpendicular from contact to center",
+ "book": 3,
+ "number": 19,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop20",
+ "type": "proposition",
+ "label": "Angle at center double angle at circumference on same base",
+ "shortLabel": "Prop. III.20",
+ "short": "Angle at center double angle at circumference",
+ "book": 3,
+ "number": 20,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop21",
+ "type": "proposition",
+ "label": "In a circle angles in same segment equal one another",
+ "shortLabel": "Prop. III.21",
+ "short": "Angles in same segment equal",
+ "book": 3,
+ "number": 21,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop22",
+ "type": "proposition",
+ "label": "Sum of opposite angles of cyclic quadrilateral equals two right angles",
+ "shortLabel": "Prop. III.22",
+ "short": "Opposite angles of cyclic quadrilateral",
+ "book": 3,
+ "number": 22,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop23",
+ "type": "proposition",
+ "label": "On same line cannot construct two similar unequal segments on same side",
+ "shortLabel": "Prop. III.23",
+ "short": "Same line, two similar unequal segments",
+ "book": 3,
+ "number": 23,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop24",
+ "type": "proposition",
+ "label": "Similar segments on equal straight lines equal one another",
+ "shortLabel": "Prop. III.24",
+ "short": "Similar segments on equal lines equal",
+ "book": 3,
+ "number": 24,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop25",
+ "type": "proposition",
+ "label": "Given segment of circle, describe complete circle",
+ "shortLabel": "Prop. III.25",
+ "short": "Complete circle from segment",
+ "book": 3,
+ "number": 25,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop26",
+ "type": "proposition",
+ "label": "In equal circles equal angles stand on equal circumferences",
+ "shortLabel": "Prop. III.26",
+ "short": "Equal angles stand on equal arcs",
+ "book": 3,
+ "number": 26,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop27",
+ "type": "proposition",
+ "label": "In equal circles angles on equal circumferences equal one another",
+ "shortLabel": "Prop. III.27",
+ "short": "Equal arcs imply equal angles",
+ "book": 3,
+ "number": 27,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop28",
+ "type": "proposition",
+ "label": "In equal circles equal chords cut off equal circumferences",
+ "shortLabel": "Prop. III.28",
+ "short": "Equal chords cut off equal arcs",
+ "book": 3,
+ "number": 28,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop29",
+ "type": "proposition",
+ "label": "In equal circles chords cutting equal circumferences are equal",
+ "shortLabel": "Prop. III.29",
+ "short": "Equal arcs imply equal chords",
+ "book": 3,
+ "number": 29,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop30",
+ "type": "proposition",
+ "label": "To bisect a given circumference",
+ "shortLabel": "Prop. III.30",
+ "short": "Bisect given circumference",
+ "book": 3,
+ "number": 30,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop31",
+ "type": "proposition",
+ "label": "Angle in semicircle right; in greater segment less; in less greater",
+ "shortLabel": "Prop. III.31",
+ "short": "Angle in semicircle is right",
+ "book": 3,
+ "number": 31,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop32",
+ "type": "proposition",
+ "label": "Angle with tangent equals angle in alternate segment",
+ "shortLabel": "Prop. III.32",
+ "short": "Tangent-chord angle equals alternate segment",
+ "book": 3,
+ "number": 32,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop33",
+ "type": "proposition",
+ "label": "On given line describe segment admitting angle equal to given",
+ "shortLabel": "Prop. III.33",
+ "short": "Segment admitting given angle",
+ "book": 3,
+ "number": 33,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop34",
+ "type": "proposition",
+ "label": "From given circle cut off segment admitting given angle",
+ "shortLabel": "Prop. III.34",
+ "short": "Cut off segment admitting angle",
+ "book": 3,
+ "number": 34,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop35",
+ "type": "proposition",
+ "label": "If chords cut one another, rectangle by segments of one equals other",
+ "shortLabel": "Prop. III.35",
+ "short": "Rectangle from chord segments equal",
+ "book": 3,
+ "number": 35,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop36",
+ "type": "proposition",
+ "label": "From point outside: tangent squared = secant × external part",
+ "shortLabel": "Prop. III.36",
+ "short": "Tangent squared = secant × external",
+ "book": 3,
+ "number": 36,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop37",
+ "type": "proposition",
+ "label": "If rectangle equals square on line, that line touches circle",
+ "shortLabel": "Prop. III.37",
+ "short": "Converse: tangent if rectangle = square",
+ "book": 3,
+ "number": 37,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "BookI",
+ "to": "Def1"
+ },
+ {
+ "from": "BookI",
+ "to": "Def2"
+ },
+ {
+ "from": "BookI",
+ "to": "Def3"
+ },
+ {
+ "from": "BookI",
+ "to": "Def4"
+ },
+ {
+ "from": "BookI",
+ "to": "Def5"
+ },
+ {
+ "from": "BookI",
+ "to": "Def6"
+ },
+ {
+ "from": "BookI",
+ "to": "Def7"
+ },
+ {
+ "from": "BookI",
+ "to": "Def8"
+ },
+ {
+ "from": "BookI",
+ "to": "Def9"
+ },
+ {
+ "from": "BookI",
+ "to": "Def10"
+ },
+ {
+ "from": "BookI",
+ "to": "Def11"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop2"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop3"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop4"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop9"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop10"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop14"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop15"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop17"
+ },
+ {
+ "from": "Prop16",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop18"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop19"
+ },
+ {
+ "from": "Prop18",
+ "to": "Prop19"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop20"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop20"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop21"
+ },
+ {
+ "from": "Prop20",
+ "to": "Prop21"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop22"
+ },
+ {
+ "from": "Prop21",
+ "to": "Prop22"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop23"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop24"
+ },
+ {
+ "from": "Prop23",
+ "to": "Prop24"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop25"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop26"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop27"
+ },
+ {
+ "from": "Prop26",
+ "to": "Prop27"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop28"
+ },
+ {
+ "from": "Prop27",
+ "to": "Prop28"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop29"
+ },
+ {
+ "from": "Prop28",
+ "to": "Prop29"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop30"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop31"
+ },
+ {
+ "from": "Prop20",
+ "to": "Prop31"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop32"
+ },
+ {
+ "from": "Prop31",
+ "to": "Prop32"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop33"
+ },
+ {
+ "from": "Prop16",
+ "to": "Prop33"
+ },
+ {
+ "from": "Prop32",
+ "to": "Prop33"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop34"
+ },
+ {
+ "from": "Prop32",
+ "to": "Prop34"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop35"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop35"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop35"
+ },
+ {
+ "from": "PropII5",
+ "to": "Prop35"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop36"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop36"
+ },
+ {
+ "from": "Prop18",
+ "to": "Prop36"
+ },
+ {
+ "from": "Prop35",
+ "to": "Prop36"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop37"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop37"
+ },
+ {
+ "from": "Prop16",
+ "to": "Prop37"
+ },
+ {
+ "from": "Prop32",
+ "to": "Prop37"
+ },
+ {
+ "from": "Prop36",
+ "to": "Prop37"
+ }
+ ],
+ "colorScheme": {
+ "foundation": {
+ "fill": "#95a5a6",
+ "stroke": "#7f8c8d"
+ },
+ "definition": {
+ "fill": "#3498db",
+ "stroke": "#2980b9"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/euclid-elements-book-iv.json b/data/euclid-elements-book-iv.json
new file mode 100644
index 0000000000000000000000000000000000000000..84e0a2c2b8469574e2acdf752af1c35b2f220c44
--- /dev/null
+++ b/data/euclid-elements-book-iv.json
@@ -0,0 +1,573 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-iv",
+ "name": "Euclid's Elements, Book IV",
+ "subject": "geometry",
+ "variant": "classical",
+ "description": "Inscribed and circumscribed figures: triangle, square, pentagon, hexagon, 15-gon. All depend on Books I and III. IV.10 uses II.11. Source: David E. Joyce.",
+ "structure": {
+ "books": 4,
+ "definitions": 7,
+ "propositions": 16,
+ "foundationTypes": [
+ "definition",
+ "foundation"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-15",
+ "lastUpdated": "2026-03-15",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book IV Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book IV",
+ "inscribed",
+ "circumscribed",
+ "pentagon",
+ "hexagon"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book IV",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookIV/bookIV.html",
+ "notes": "Clark University; Logical structure"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "BookI",
+ "type": "foundation",
+ "label": "Book I — Fundamentals of plane geometry",
+ "shortLabel": "Book I",
+ "short": "Foundation",
+ "book": 1,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookIII",
+ "type": "foundation",
+ "label": "Book III — Theory of circles",
+ "shortLabel": "Book III",
+ "short": "Foundation",
+ "book": 3,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "PropII11",
+ "type": "foundation",
+ "label": "Prop. II.11 — Golden section",
+ "shortLabel": "Prop. II.11",
+ "short": "From Book II",
+ "book": 2,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "Def1",
+ "type": "definition",
+ "label": "Rectilinear figure inscribed in circle when each vertex on circumference",
+ "shortLabel": "Def. IV.1",
+ "short": "Inscribe in circle",
+ "book": 4,
+ "number": 1,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def2",
+ "type": "definition",
+ "label": "Figure circumscribed about circle when each side touches circle",
+ "shortLabel": "Def. IV.2",
+ "short": "Circumscribe about circle",
+ "book": 4,
+ "number": 2,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def3",
+ "type": "definition",
+ "label": "Circle inscribed in figure when each side touches circle",
+ "shortLabel": "Def. IV.3",
+ "short": "Inscribe circle in figure",
+ "book": 4,
+ "number": 3,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def4",
+ "type": "definition",
+ "label": "Circle circumscribed about figure when each vertex on circumference",
+ "shortLabel": "Def. IV.4",
+ "short": "Circumscribe circle about figure",
+ "book": 4,
+ "number": 4,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def5",
+ "type": "definition",
+ "label": "Figure inscribed in figure when each vertex of inner on sides of outer",
+ "shortLabel": "Def. IV.5",
+ "short": "Inscribe in figure",
+ "book": 4,
+ "number": 5,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def6",
+ "type": "definition",
+ "label": "Figure circumscribed about figure when each side of outer touches inner",
+ "shortLabel": "Def. IV.6",
+ "short": "Circumscribe about figure",
+ "book": 4,
+ "number": 6,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def7",
+ "type": "definition",
+ "label": "Straight line inscribed in circle when its ends on circumference",
+ "shortLabel": "Def. IV.7",
+ "short": "Inscribe line in circle",
+ "book": 4,
+ "number": 7,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "To fit into given circle a straight line equal to given, not greater than diameter",
+ "shortLabel": "Prop. IV.1",
+ "short": "Fit line in circle",
+ "book": 4,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "To inscribe in given circle a triangle equiangular with given triangle",
+ "shortLabel": "Prop. IV.2",
+ "short": "Inscribe triangle in circle",
+ "book": 4,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "To circumscribe about given circle a triangle equiangular with given",
+ "shortLabel": "Prop. IV.3",
+ "short": "Circumscribe triangle about circle",
+ "book": 4,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "To inscribe a circle in a given triangle",
+ "shortLabel": "Prop. IV.4",
+ "short": "Inscribe circle in triangle",
+ "book": 4,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "To circumscribe a circle about a given triangle",
+ "shortLabel": "Prop. IV.5",
+ "short": "Circumscribe circle about triangle",
+ "book": 4,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "To inscribe a square in a given circle",
+ "shortLabel": "Prop. IV.6",
+ "short": "Inscribe square in circle",
+ "book": 4,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "To circumscribe a square about a given circle",
+ "shortLabel": "Prop. IV.7",
+ "short": "Circumscribe square about circle",
+ "book": 4,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "To inscribe a circle in a given square",
+ "shortLabel": "Prop. IV.8",
+ "short": "Inscribe circle in square",
+ "book": 4,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "To circumscribe a circle about a given square",
+ "shortLabel": "Prop. IV.9",
+ "short": "Circumscribe circle about square",
+ "book": 4,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "To construct isosceles triangle with each base angle double the remaining",
+ "shortLabel": "Prop. IV.10",
+ "short": "Isosceles triangle, base angles double",
+ "book": 4,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "To inscribe an equilateral equiangular pentagon in a given circle",
+ "shortLabel": "Prop. IV.11",
+ "short": "Inscribe pentagon in circle",
+ "book": 4,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "To circumscribe an equilateral equiangular pentagon about a given circle",
+ "shortLabel": "Prop. IV.12",
+ "short": "Circumscribe pentagon about circle",
+ "book": 4,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "To inscribe a circle in a given equilateral equiangular pentagon",
+ "shortLabel": "Prop. IV.13",
+ "short": "Inscribe circle in pentagon",
+ "book": 4,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "To circumscribe a circle about a given equilateral equiangular pentagon",
+ "shortLabel": "Prop. IV.14",
+ "short": "Circumscribe circle about pentagon",
+ "book": 4,
+ "number": 14,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop15",
+ "type": "proposition",
+ "label": "To inscribe an equilateral equiangular hexagon in a given circle",
+ "shortLabel": "Prop. IV.15",
+ "short": "Inscribe hexagon in circle",
+ "book": 4,
+ "number": 15,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop16",
+ "type": "proposition",
+ "label": "To inscribe an equilateral equiangular fifteen-angled figure in a given circle",
+ "shortLabel": "Prop. IV.16",
+ "short": "Inscribe 15-gon in circle",
+ "book": 4,
+ "number": 16,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "BookI",
+ "to": "Def1"
+ },
+ {
+ "from": "BookIII",
+ "to": "Def1"
+ },
+ {
+ "from": "BookI",
+ "to": "Def2"
+ },
+ {
+ "from": "BookIII",
+ "to": "Def2"
+ },
+ {
+ "from": "BookI",
+ "to": "Def3"
+ },
+ {
+ "from": "BookIII",
+ "to": "Def3"
+ },
+ {
+ "from": "BookI",
+ "to": "Def4"
+ },
+ {
+ "from": "BookIII",
+ "to": "Def4"
+ },
+ {
+ "from": "BookI",
+ "to": "Def5"
+ },
+ {
+ "from": "BookIII",
+ "to": "Def5"
+ },
+ {
+ "from": "BookI",
+ "to": "Def6"
+ },
+ {
+ "from": "BookIII",
+ "to": "Def6"
+ },
+ {
+ "from": "BookI",
+ "to": "Def7"
+ },
+ {
+ "from": "BookIII",
+ "to": "Def7"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop6"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop7"
+ },
+ {
+ "from": "Prop6",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop8"
+ },
+ {
+ "from": "Prop7",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop9"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop10"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop10"
+ },
+ {
+ "from": "Prop5",
+ "to": "Prop10"
+ },
+ {
+ "from": "PropII11",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop11"
+ },
+ {
+ "from": "Prop2",
+ "to": "Prop11"
+ },
+ {
+ "from": "Prop10",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop12"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop13"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop14"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop15"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookIII",
+ "to": "Prop16"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop16"
+ },
+ {
+ "from": "Prop2",
+ "to": "Prop16"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop16"
+ }
+ ],
+ "colorScheme": {
+ "foundation": {
+ "fill": "#95a5a6",
+ "stroke": "#7f8c8d"
+ },
+ "definition": {
+ "fill": "#3498db",
+ "stroke": "#2980b9"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/euclid-elements-book-ix.json b/data/euclid-elements-book-ix.json
new file mode 100644
index 0000000000000000000000000000000000000000..c34e46acf3ce47ae4de4e25c09aebaa44e217ed4
--- /dev/null
+++ b/data/euclid-elements-book-ix.json
@@ -0,0 +1,728 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-ix",
+ "name": "Euclid's Elements, Book IX",
+ "subject": "number_theory",
+ "variant": "classical",
+ "description": "Primes, perfect numbers, odd/even. 36 propositions. Depends on Books VII and VIII. Source: David E. Joyce.",
+ "structure": {
+ "books": 9,
+ "propositions": 36,
+ "foundationTypes": [
+ "foundation"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-18",
+ "lastUpdated": "2026-03-18",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book IX Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book IX",
+ "prime",
+ "perfect",
+ "odd",
+ "even"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book IX",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookIX/bookIX.html",
+ "notes": "Clark University; IX.20 infinitude of primes"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "BookVII",
+ "type": "foundation",
+ "label": "Book VII — Number theory",
+ "shortLabel": "Book VII",
+ "short": "Foundation",
+ "book": 7,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookVIII",
+ "type": "foundation",
+ "label": "Book VIII — Continued proportions",
+ "shortLabel": "Book VIII",
+ "short": "Foundation",
+ "book": 8,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "Two similar plane numbers multiplied: product is square",
+ "shortLabel": "Prop. IX.1",
+ "short": "Similar plane product square",
+ "book": 9,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "Two numbers product square: they are similar plane",
+ "shortLabel": "Prop. IX.2",
+ "short": "Product square: similar plane",
+ "book": 9,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "Cubic number multiplied by itself: product is cube",
+ "shortLabel": "Prop. IX.3",
+ "short": "Cube times itself",
+ "book": 9,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "Cubic times cubic: product is cube",
+ "shortLabel": "Prop. IX.4",
+ "short": "Cube times cube",
+ "book": 9,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "If cube times any makes cube, the multiplied is cubic",
+ "shortLabel": "Prop. IX.5",
+ "short": "Cube times any makes cube",
+ "book": 9,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "If number times itself makes cubic, it is cubic",
+ "shortLabel": "Prop. IX.6",
+ "short": "Number times itself cubic",
+ "book": 9,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "Composite times any: product is solid",
+ "shortLabel": "Prop. IX.7",
+ "short": "Composite times any: solid",
+ "book": 9,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "Numbers from unit in continued proportion: 3rd square, 4th cube, 7th both",
+ "shortLabel": "Prop. IX.8",
+ "short": "Continued proportion from unit",
+ "book": 9,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "If second from unit square, all square; if cubic, all cubic",
+ "shortLabel": "Prop. IX.9",
+ "short": "Second square: all square",
+ "book": 9,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "If second not square, only 3rd and every other square; similar for cube",
+ "shortLabel": "Prop. IX.10",
+ "short": "Second not square",
+ "book": 9,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "Continued proportion from unit: less measures greater",
+ "shortLabel": "Prop. IX.11",
+ "short": "Less measures greater",
+ "book": 9,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "If prime measures last, it measures second from unit",
+ "shortLabel": "Prop. IX.12",
+ "short": "Prime measures last",
+ "book": 9,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "Continued proportion, second prime: only proportional numbers measure last",
+ "shortLabel": "Prop. IX.13",
+ "short": "Second prime: only those measure",
+ "book": 9,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "Least measured by given primes: not measured by any other prime",
+ "shortLabel": "Prop. IX.14",
+ "short": "Least measured by primes",
+ "book": 9,
+ "number": 14,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop15",
+ "type": "proposition",
+ "label": "Three least in ratio: sum of any two prime to remainder",
+ "shortLabel": "Prop. IX.15",
+ "short": "Three in proportion",
+ "book": 9,
+ "number": 15,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop16",
+ "type": "proposition",
+ "label": "Two relatively prime: no third as first to second",
+ "shortLabel": "Prop. IX.16",
+ "short": "Relatively prime: no third",
+ "book": 9,
+ "number": 16,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop17",
+ "type": "proposition",
+ "label": "Continued proportion, extremes prime: last not to any as first to second",
+ "shortLabel": "Prop. IX.17",
+ "short": "Extremes prime: no extension",
+ "book": 9,
+ "number": 17,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop18",
+ "type": "proposition",
+ "label": "Given two numbers, investigate if third proportional exists",
+ "shortLabel": "Prop. IX.18",
+ "short": "Third proportional",
+ "book": 9,
+ "number": 18,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop19",
+ "type": "proposition",
+ "label": "Given three numbers, investigate when fourth proportional exists",
+ "shortLabel": "Prop. IX.19",
+ "short": "Fourth proportional",
+ "book": 9,
+ "number": 19,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop20",
+ "type": "proposition",
+ "label": "Prime numbers are more than any assigned multitude",
+ "shortLabel": "Prop. IX.20",
+ "short": "Infinitude of primes",
+ "book": 9,
+ "number": 20,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop21",
+ "type": "proposition",
+ "label": "Sum of even numbers is even",
+ "shortLabel": "Prop. IX.21",
+ "short": "Sum of evens even",
+ "book": 9,
+ "number": 21,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop22",
+ "type": "proposition",
+ "label": "Sum of odd numbers, even multitude: sum even",
+ "shortLabel": "Prop. IX.22",
+ "short": "Sum of odds (even count) even",
+ "book": 9,
+ "number": 22,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop23",
+ "type": "proposition",
+ "label": "Sum of odd numbers, odd multitude: sum odd",
+ "shortLabel": "Prop. IX.23",
+ "short": "Sum of odds (odd count) odd",
+ "book": 9,
+ "number": 23,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop24",
+ "type": "proposition",
+ "label": "Even minus even: remainder even",
+ "shortLabel": "Prop. IX.24",
+ "short": "Even minus even",
+ "book": 9,
+ "number": 24,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop25",
+ "type": "proposition",
+ "label": "Even minus odd: remainder odd",
+ "shortLabel": "Prop. IX.25",
+ "short": "Even minus odd",
+ "book": 9,
+ "number": 25,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop26",
+ "type": "proposition",
+ "label": "Odd minus odd: remainder even",
+ "shortLabel": "Prop. IX.26",
+ "short": "Odd minus odd",
+ "book": 9,
+ "number": 26,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop27",
+ "type": "proposition",
+ "label": "Odd minus even: remainder odd",
+ "shortLabel": "Prop. IX.27",
+ "short": "Odd minus even",
+ "book": 9,
+ "number": 27,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop28",
+ "type": "proposition",
+ "label": "Odd times even: product even",
+ "shortLabel": "Prop. IX.28",
+ "short": "Odd times even",
+ "book": 9,
+ "number": 28,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop29",
+ "type": "proposition",
+ "label": "Odd times odd: product odd",
+ "shortLabel": "Prop. IX.29",
+ "short": "Odd times odd",
+ "book": 9,
+ "number": 29,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop30",
+ "type": "proposition",
+ "label": "Odd measuring even: measures half",
+ "shortLabel": "Prop. IX.30",
+ "short": "Odd measures even",
+ "book": 9,
+ "number": 30,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop31",
+ "type": "proposition",
+ "label": "Odd relatively prime to any: also prime to its double",
+ "shortLabel": "Prop. IX.31",
+ "short": "Odd prime to double",
+ "book": 9,
+ "number": 31,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop32",
+ "type": "proposition",
+ "label": "Numbers doubled from 2: even-times even only",
+ "shortLabel": "Prop. IX.32",
+ "short": "Powers of 2",
+ "book": 9,
+ "number": 32,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop33",
+ "type": "proposition",
+ "label": "Number with half odd: even-times odd only",
+ "shortLabel": "Prop. IX.33",
+ "short": "Half odd",
+ "book": 9,
+ "number": 33,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop34",
+ "type": "proposition",
+ "label": "Number neither: both even-times even and even-times odd",
+ "shortLabel": "Prop. IX.34",
+ "short": "Neither",
+ "book": 9,
+ "number": 34,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop35",
+ "type": "proposition",
+ "label": "Continued proportion: (second−first):first = (last−first):sum of rest",
+ "shortLabel": "Prop. IX.35",
+ "short": "Geometric series",
+ "book": 9,
+ "number": 35,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop36",
+ "type": "proposition",
+ "label": "If sum of powers of 2 is prime, product with last is perfect",
+ "shortLabel": "Prop. IX.36",
+ "short": "Perfect numbers",
+ "book": 9,
+ "number": 36,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "BookVII",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop19"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop19"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop20"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop20"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop21"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop21"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop22"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop22"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop23"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop23"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop24"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop24"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop25"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop25"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop26"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop26"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop27"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop27"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop28"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop28"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop29"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop29"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop30"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop30"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop31"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop31"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop32"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop32"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop33"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop33"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop34"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop34"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop35"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop35"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop36"
+ },
+ {
+ "from": "BookVIII",
+ "to": "Prop36"
+ }
+ ],
+ "colorScheme": {
+ "foundation": {
+ "fill": "#95a5a6",
+ "stroke": "#7f8c8d"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/euclid-elements-book-v.json b/data/euclid-elements-book-v.json
new file mode 100644
index 0000000000000000000000000000000000000000..8eb55c8259b5c0c0578d37a2284184dcf156338f
--- /dev/null
+++ b/data/euclid-elements-book-v.json
@@ -0,0 +1,676 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-v",
+ "name": "Euclid's Elements, Book V",
+ "subject": "geometry",
+ "variant": "classical",
+ "description": "Theory of ratio and proportion (Eudoxus). 18 definitions, 25 propositions. Does not depend on previous books. Source: David E. Joyce.",
+ "structure": {
+ "books": 5,
+ "definitions": 18,
+ "propositions": 25,
+ "foundationTypes": [
+ "definition"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-15",
+ "lastUpdated": "2026-03-15",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book V Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book V",
+ "proportion",
+ "ratio",
+ "Eudoxus"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book V",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookV/bookV.html",
+ "notes": "Clark University; Logical structure"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "Def1",
+ "type": "definition",
+ "label": "A magnitude is a part of a magnitude when it measures it",
+ "shortLabel": "Def. V.1",
+ "short": "Part",
+ "book": 5,
+ "number": 1,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def2",
+ "type": "definition",
+ "label": "The greater is a multiple of the less when it is measured by the less",
+ "shortLabel": "Def. V.2",
+ "short": "Multiple",
+ "book": 5,
+ "number": 2,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def3",
+ "type": "definition",
+ "label": "A ratio is a sort of relation in respect of size between two magnitudes",
+ "shortLabel": "Def. V.3",
+ "short": "Ratio",
+ "book": 5,
+ "number": 3,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def4",
+ "type": "definition",
+ "label": "Magnitudes have a ratio when the less can be multiplied to exceed the greater",
+ "shortLabel": "Def. V.4",
+ "short": "Same ratio",
+ "book": 5,
+ "number": 4,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def5",
+ "type": "definition",
+ "label": "Magnitudes in same ratio when equimultiples alike exceed, equal, or fall short",
+ "shortLabel": "Def. V.5",
+ "short": "In same ratio (Eudoxus)",
+ "book": 5,
+ "number": 5,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def6",
+ "type": "definition",
+ "label": "Magnitudes which have the same ratio are proportional",
+ "shortLabel": "Def. V.6",
+ "short": "Proportional",
+ "book": 5,
+ "number": 6,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def7",
+ "type": "definition",
+ "label": "When of equimultiples first exceeds second, third does not exceed fourth",
+ "shortLabel": "Def. V.7",
+ "short": "Greater ratio",
+ "book": 5,
+ "number": 7,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def8",
+ "type": "definition",
+ "label": "Compound ratio is the ratio of the products of corresponding terms",
+ "shortLabel": "Def. V.8",
+ "short": "Compound ratio",
+ "book": 5,
+ "number": 8,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def9",
+ "type": "definition",
+ "label": "Duplicate ratio is the ratio of the squares",
+ "shortLabel": "Def. V.9",
+ "short": "Duplicate ratio",
+ "book": 5,
+ "number": 9,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def10",
+ "type": "definition",
+ "label": "Triplicate ratio is the ratio of the cubes",
+ "shortLabel": "Def. V.10",
+ "short": "Triplicate ratio",
+ "book": 5,
+ "number": 10,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def11",
+ "type": "definition",
+ "label": "Corresponding magnitudes in proportion",
+ "shortLabel": "Def. V.11",
+ "short": "Corresponding magnitudes",
+ "book": 5,
+ "number": 11,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def12",
+ "type": "definition",
+ "label": "Alternate: first to third as second to fourth",
+ "shortLabel": "Def. V.12",
+ "short": "Alternate ratio",
+ "book": 5,
+ "number": 12,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def13",
+ "type": "definition",
+ "label": "Inverse: second to first as fourth to third",
+ "shortLabel": "Def. V.13",
+ "short": "Inverse ratio",
+ "book": 5,
+ "number": 13,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def14",
+ "type": "definition",
+ "label": "Composition: first+second to second as third+fourth to fourth",
+ "shortLabel": "Def. V.14",
+ "short": "Composition of ratio",
+ "book": 5,
+ "number": 14,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def15",
+ "type": "definition",
+ "label": "Separation: first−second to second as third−fourth to fourth",
+ "shortLabel": "Def. V.15",
+ "short": "Separation of ratio",
+ "book": 5,
+ "number": 15,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def16",
+ "type": "definition",
+ "label": "Conversion: first to first−second as third to third−fourth",
+ "shortLabel": "Def. V.16",
+ "short": "Conversion of ratio",
+ "book": 5,
+ "number": 16,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def17",
+ "type": "definition",
+ "label": "Ex aequali: when first to second as second to third",
+ "shortLabel": "Def. V.17",
+ "short": "Ex aequali",
+ "book": 5,
+ "number": 17,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def18",
+ "type": "definition",
+ "label": "Ex aequali perturbed: when ratios are in perturbed order",
+ "shortLabel": "Def. V.18",
+ "short": "Ex aequali perturbed",
+ "book": 5,
+ "number": 18,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "If magnitudes each same multiple of others, sum is that multiple of sum",
+ "shortLabel": "Prop. V.1",
+ "short": "Sum of multiples",
+ "book": 5,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "If first:second as third:fourth, sum of first and fifth as sum of third and sixth",
+ "shortLabel": "Prop. V.2",
+ "short": "Equimultiples sum",
+ "book": 5,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "Equimultiples of equimultiples are equimultiples",
+ "shortLabel": "Prop. V.3",
+ "short": "Equimultiples of equimultiples",
+ "book": 5,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "If a:b = c:d, then ma:nb = mc:nd",
+ "shortLabel": "Prop. V.4",
+ "short": "Equimultiples preserve ratio",
+ "book": 5,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "Multiple of whole minus multiple of part = multiple of remainder",
+ "shortLabel": "Prop. V.5",
+ "short": "Multiple of difference",
+ "book": 5,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "Equimultiples minus equimultiples equal or equimultiples",
+ "shortLabel": "Prop. V.6",
+ "short": "Equimultiples minus equimultiples",
+ "book": 5,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "Equal magnitudes have same ratio to same; same to equals",
+ "shortLabel": "Prop. V.7",
+ "short": "Equals in ratio",
+ "book": 5,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "Of unequal magnitudes, greater has greater ratio to same",
+ "shortLabel": "Prop. V.8",
+ "short": "Greater has greater ratio",
+ "book": 5,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "Magnitudes with same ratio to same are equal",
+ "shortLabel": "Prop. V.9",
+ "short": "Same ratio implies equal",
+ "book": 5,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "Of magnitudes with ratio to same, greater ratio implies greater",
+ "shortLabel": "Prop. V.10",
+ "short": "Greater ratio implies greater",
+ "book": 5,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "Ratios same with same ratio are same with one another",
+ "shortLabel": "Prop. V.11",
+ "short": "Transitivity of ratios",
+ "book": 5,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "Proportional: one antecedent to one consequent as sum to sum",
+ "shortLabel": "Prop. V.12",
+ "short": "Sum of antecedents/consequents",
+ "book": 5,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "If a:b = c:d and c:d > e:f, then a:b > e:f",
+ "shortLabel": "Prop. V.13",
+ "short": "Substitution in ratio inequality",
+ "book": 5,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "If a:b = c:d and a>c, then b>d",
+ "shortLabel": "Prop. V.14",
+ "short": "Equal ratios, equal magnitudes",
+ "book": 5,
+ "number": 14,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop15",
+ "type": "proposition",
+ "label": "Parts have same ratio as their equimultiples",
+ "shortLabel": "Prop. V.15",
+ "short": "Parts as equimultiples",
+ "book": 5,
+ "number": 15,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop16",
+ "type": "proposition",
+ "label": "If a:b = c:d, then a:c = b:d",
+ "shortLabel": "Prop. V.16",
+ "short": "Alternate proportion",
+ "book": 5,
+ "number": 16,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop17",
+ "type": "proposition",
+ "label": "If (a+b):b = (c+d):d, then a:b = c:d",
+ "shortLabel": "Prop. V.17",
+ "short": "Jointly implies separately",
+ "book": 5,
+ "number": 17,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop18",
+ "type": "proposition",
+ "label": "If a:b = c:d, then (a+b):b = (c+d):d",
+ "shortLabel": "Prop. V.18",
+ "short": "Separately implies jointly",
+ "book": 5,
+ "number": 18,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop19",
+ "type": "proposition",
+ "label": "If (a+b):(c+d) = a:c, then also = b:d",
+ "shortLabel": "Prop. V.19",
+ "short": "Whole to whole as part to part",
+ "book": 5,
+ "number": 19,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop20",
+ "type": "proposition",
+ "label": "If a:b = d:e and b:c = e:f and a>c, then d>f",
+ "shortLabel": "Prop. V.20",
+ "short": "Ex aequali (direct)",
+ "book": 5,
+ "number": 20,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop21",
+ "type": "proposition",
+ "label": "If a:b = e:f and b:c = d:e and a>c, then d>f",
+ "shortLabel": "Prop. V.21",
+ "short": "Ex aequali (perturbed)",
+ "book": 5,
+ "number": 21,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop22",
+ "type": "proposition",
+ "label": "If a1:a2 = b1:b2, a2:a3 = b2:b3, ..., then a1:an = b1:bn",
+ "shortLabel": "Prop. V.22",
+ "short": "Ex aequali chain",
+ "book": 5,
+ "number": 22,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop23",
+ "type": "proposition",
+ "label": "If a:b = y:z and b:c = x:y, then a:c = x:z",
+ "shortLabel": "Prop. V.23",
+ "short": "Ex aequali perturbed chain",
+ "book": 5,
+ "number": 23,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop24",
+ "type": "proposition",
+ "label": "If a:b = c:d and e:b = f:d, then (a+e):b = (c+f):d",
+ "shortLabel": "Prop. V.24",
+ "short": "Sum of ratios",
+ "book": 5,
+ "number": 24,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop25",
+ "type": "proposition",
+ "label": "If a:b = c:d and a greatest, d least, then a+d > b+c",
+ "shortLabel": "Prop. V.25",
+ "short": "Sum of extremes > sum of means",
+ "book": 5,
+ "number": 25,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "Prop2",
+ "to": "Prop3"
+ },
+ {
+ "from": "Prop3",
+ "to": "Prop4"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop5"
+ },
+ {
+ "from": "Prop2",
+ "to": "Prop6"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop8"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop9"
+ },
+ {
+ "from": "Prop7",
+ "to": "Prop10"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop10"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop12"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop14"
+ },
+ {
+ "from": "Prop10",
+ "to": "Prop14"
+ },
+ {
+ "from": "Prop13",
+ "to": "Prop14"
+ },
+ {
+ "from": "Prop7",
+ "to": "Prop15"
+ },
+ {
+ "from": "Prop12",
+ "to": "Prop15"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop16"
+ },
+ {
+ "from": "Prop14",
+ "to": "Prop16"
+ },
+ {
+ "from": "Prop15",
+ "to": "Prop16"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop17"
+ },
+ {
+ "from": "Prop2",
+ "to": "Prop17"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop18"
+ },
+ {
+ "from": "Prop14",
+ "to": "Prop18"
+ },
+ {
+ "from": "Prop17",
+ "to": "Prop18"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop19"
+ },
+ {
+ "from": "Prop16",
+ "to": "Prop19"
+ },
+ {
+ "from": "Prop17",
+ "to": "Prop19"
+ },
+ {
+ "from": "Prop7",
+ "to": "Prop20"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop20"
+ },
+ {
+ "from": "Prop10",
+ "to": "Prop20"
+ },
+ {
+ "from": "Prop13",
+ "to": "Prop20"
+ },
+ {
+ "from": "Prop7",
+ "to": "Prop21"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop21"
+ },
+ {
+ "from": "Prop10",
+ "to": "Prop21"
+ },
+ {
+ "from": "Prop13",
+ "to": "Prop21"
+ },
+ {
+ "from": "Prop4",
+ "to": "Prop22"
+ },
+ {
+ "from": "Prop20",
+ "to": "Prop22"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop23"
+ },
+ {
+ "from": "Prop15",
+ "to": "Prop23"
+ },
+ {
+ "from": "Prop16",
+ "to": "Prop23"
+ },
+ {
+ "from": "Prop21",
+ "to": "Prop23"
+ },
+ {
+ "from": "Prop7",
+ "to": "Prop24"
+ },
+ {
+ "from": "Prop18",
+ "to": "Prop24"
+ },
+ {
+ "from": "Prop22",
+ "to": "Prop24"
+ },
+ {
+ "from": "Prop7",
+ "to": "Prop25"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop25"
+ },
+ {
+ "from": "Prop14",
+ "to": "Prop25"
+ },
+ {
+ "from": "Prop19",
+ "to": "Prop25"
+ }
+ ],
+ "colorScheme": {
+ "definition": {
+ "fill": "#3498db",
+ "stroke": "#2980b9"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/euclid-elements-book-vi.json b/data/euclid-elements-book-vi.json
new file mode 100644
index 0000000000000000000000000000000000000000..c0a26e1e43f3dd8b64a45b9212d16aecd60850dc
--- /dev/null
+++ b/data/euclid-elements-book-vi.json
@@ -0,0 +1,875 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-vi",
+ "name": "Euclid's Elements, Book VI",
+ "subject": "geometry",
+ "variant": "classical",
+ "description": "Similar figures. 4 definitions, 33 propositions. Depends on Book I and Book V. VI.1 is basis for most. Source: David E. Joyce.",
+ "structure": {
+ "books": 6,
+ "definitions": 4,
+ "propositions": 33,
+ "foundationTypes": [
+ "definition",
+ "foundation"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-15",
+ "lastUpdated": "2026-03-15",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book VI Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book VI",
+ "similar",
+ "proportion",
+ "triangles"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book VI",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookVI/bookVI.html",
+ "notes": "Clark University; VI.1 basis for most"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "BookI",
+ "type": "foundation",
+ "label": "Book I — Fundamentals of plane geometry",
+ "shortLabel": "Book I",
+ "short": "Foundation",
+ "book": 1,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookV",
+ "type": "foundation",
+ "label": "Book V — Theory of proportions",
+ "shortLabel": "Book V",
+ "short": "Foundation",
+ "book": 5,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "Def1",
+ "type": "definition",
+ "label": "Similar rectilinear figures have equal angles and proportional sides",
+ "shortLabel": "Def. VI.1",
+ "short": "Similar rectilinear",
+ "book": 6,
+ "number": 1,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def2",
+ "type": "definition",
+ "label": "Figures reciprocally proportional when sides are proportional inversely",
+ "shortLabel": "Def. VI.2",
+ "short": "Reciprocally proportional",
+ "book": 6,
+ "number": 2,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def3",
+ "type": "definition",
+ "label": "Straight line is mean proportional when first to it as it to third",
+ "shortLabel": "Def. VI.3",
+ "short": "Mean proportional",
+ "book": 6,
+ "number": 3,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def4",
+ "type": "definition",
+ "label": "Duplicate ratio is the ratio of the squares on corresponding sides",
+ "shortLabel": "Def. VI.4",
+ "short": "Duplicate ratio",
+ "book": 6,
+ "number": 4,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "Triangles and parallelograms under same height are as their bases",
+ "shortLabel": "Prop. VI.1",
+ "short": "Triangles under same height",
+ "book": 6,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "Line parallel to side cuts sides proportionally; converse",
+ "shortLabel": "Prop. VI.2",
+ "short": "Parallel cuts sides proportionally",
+ "book": 6,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "Angle bisector: segments of base proportionally as remaining sides",
+ "shortLabel": "Prop. VI.3",
+ "short": "Angle bisector divides base",
+ "book": 6,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "Equiangular triangles: sides about equal angles proportional",
+ "shortLabel": "Prop. VI.4",
+ "short": "Equiangular: sides proportional",
+ "book": 6,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "If sides proportional, triangles equiangular",
+ "shortLabel": "Prop. VI.5",
+ "short": "Sides proportional: equiangular",
+ "book": 6,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "One angle equal, sides about it proportional: equiangular",
+ "shortLabel": "Prop. VI.6",
+ "short": "One angle equal, sides proportional",
+ "book": 6,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "One angle equal, sides about others proportional: equiangular",
+ "shortLabel": "Prop. VI.7",
+ "short": "One angle equal, other sides proportional",
+ "book": 6,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "Perpendicular from right angle: triangles similar to whole",
+ "shortLabel": "Prop. VI.8",
+ "short": "Altitude in right triangle",
+ "book": 6,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "To cut off a prescribed part from a given straight line",
+ "shortLabel": "Prop. VI.9",
+ "short": "Cut off prescribed part",
+ "book": 6,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "To cut a given line similarly to a given cut line",
+ "shortLabel": "Prop. VI.10",
+ "short": "Cut line similarly",
+ "book": 6,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "To find a third proportional to two given lines",
+ "shortLabel": "Prop. VI.11",
+ "short": "Third proportional",
+ "book": 6,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "To find a fourth proportional to three given lines",
+ "shortLabel": "Prop. VI.12",
+ "short": "Fourth proportional",
+ "book": 6,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "To find a mean proportional to two given lines",
+ "shortLabel": "Prop. VI.13",
+ "short": "Mean proportional",
+ "book": 6,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "Equal equiangular parallelograms: sides reciprocally proportional",
+ "shortLabel": "Prop. VI.14",
+ "short": "Parallelograms reciprocally proportional",
+ "book": 6,
+ "number": 14,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop15",
+ "type": "proposition",
+ "label": "Equal triangles, one angle equal: sides reciprocally proportional",
+ "shortLabel": "Prop. VI.15",
+ "short": "Triangles reciprocally proportional",
+ "book": 6,
+ "number": 15,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop16",
+ "type": "proposition",
+ "label": "Four lines proportional iff rectangle extremes = rectangle means",
+ "shortLabel": "Prop. VI.16",
+ "short": "Four lines proportional: rectangle",
+ "book": 6,
+ "number": 16,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop17",
+ "type": "proposition",
+ "label": "Three lines proportional iff rectangle extremes = square on mean",
+ "shortLabel": "Prop. VI.17",
+ "short": "Three lines proportional: rectangle",
+ "book": 6,
+ "number": 17,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop18",
+ "type": "proposition",
+ "label": "To describe figure similar to given on given line",
+ "shortLabel": "Prop. VI.18",
+ "short": "Similar figure on given line",
+ "book": 6,
+ "number": 18,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop19",
+ "type": "proposition",
+ "label": "Similar triangles in duplicate ratio of corresponding sides",
+ "shortLabel": "Prop. VI.19",
+ "short": "Similar triangles: duplicate ratio",
+ "book": 6,
+ "number": 19,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop20",
+ "type": "proposition",
+ "label": "Similar polygons in duplicate ratio of corresponding sides",
+ "shortLabel": "Prop. VI.20",
+ "short": "Similar polygons: duplicate ratio",
+ "book": 6,
+ "number": 20,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop21",
+ "type": "proposition",
+ "label": "Figures similar to same rectilinear figure are similar",
+ "shortLabel": "Prop. VI.21",
+ "short": "Similar to same: similar",
+ "book": 6,
+ "number": 21,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop22",
+ "type": "proposition",
+ "label": "Four lines proportional iff similar figures on them proportional",
+ "shortLabel": "Prop. VI.22",
+ "short": "Four lines proportional: figures",
+ "book": 6,
+ "number": 22,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop23",
+ "type": "proposition",
+ "label": "Equiangular parallelograms: ratio compounded of sides",
+ "shortLabel": "Prop. VI.23",
+ "short": "Equiangular parallelograms: compound ratio",
+ "book": 6,
+ "number": 23,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop24",
+ "type": "proposition",
+ "label": "Parallelograms about diameter similar to whole",
+ "shortLabel": "Prop. VI.24",
+ "short": "Parallelograms about diameter",
+ "book": 6,
+ "number": 24,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop25",
+ "type": "proposition",
+ "label": "To construct figure similar to one and equal to another",
+ "shortLabel": "Prop. VI.25",
+ "short": "Similar and equal to another",
+ "book": 6,
+ "number": 25,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop26",
+ "type": "proposition",
+ "label": "Parallelogram similar to whole, common angle: about same diameter",
+ "shortLabel": "Prop. VI.26",
+ "short": "Parallelogram similar to difference",
+ "book": 6,
+ "number": 26,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop27",
+ "type": "proposition",
+ "label": "Of parallelograms applied to line, greatest is on half",
+ "shortLabel": "Prop. VI.27",
+ "short": "Greatest parallelogram applied",
+ "book": 6,
+ "number": 27,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop28",
+ "type": "proposition",
+ "label": "To apply parallelogram equal to figure, deficient by similar",
+ "shortLabel": "Prop. VI.28",
+ "short": "Apply parallelogram: deficient",
+ "book": 6,
+ "number": 28,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop29",
+ "type": "proposition",
+ "label": "To apply parallelogram equal to figure, exceeding by similar",
+ "shortLabel": "Prop. VI.29",
+ "short": "Apply parallelogram: exceeding",
+ "book": 6,
+ "number": 29,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop30",
+ "type": "proposition",
+ "label": "To cut a given line in extreme and mean ratio",
+ "shortLabel": "Prop. VI.30",
+ "short": "Extreme and mean ratio",
+ "book": 6,
+ "number": 30,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop31",
+ "type": "proposition",
+ "label": "In right triangle, figure on hypotenuse = sum of similar on sides",
+ "shortLabel": "Prop. VI.31",
+ "short": "Right triangle: similar figures",
+ "book": 6,
+ "number": 31,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop32",
+ "type": "proposition",
+ "label": "Two triangles, two sides proportional, placed with parallel sides",
+ "shortLabel": "Prop. VI.32",
+ "short": "Two triangles: sides parallel",
+ "book": 6,
+ "number": 32,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop33",
+ "type": "proposition",
+ "label": "Angles in equal circles have ratio of circumferences",
+ "shortLabel": "Prop. VI.33",
+ "short": "Angles in circles: ratio of arcs",
+ "book": 6,
+ "number": 33,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "BookI",
+ "to": "Def1"
+ },
+ {
+ "from": "BookV",
+ "to": "Def1"
+ },
+ {
+ "from": "BookI",
+ "to": "Def2"
+ },
+ {
+ "from": "BookV",
+ "to": "Def2"
+ },
+ {
+ "from": "BookI",
+ "to": "Def3"
+ },
+ {
+ "from": "BookV",
+ "to": "Def3"
+ },
+ {
+ "from": "BookI",
+ "to": "Def4"
+ },
+ {
+ "from": "BookV",
+ "to": "Def4"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop2"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop3"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop4"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop5"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop6"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop7"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop8"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop9"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop10"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop11"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop12"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop13"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop14"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop15"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop16"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop17"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop18"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop19"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop19"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop19"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop20"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop20"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop20"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop21"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop21"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop21"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop22"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop22"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop22"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop23"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop23"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop23"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop24"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop24"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop24"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop25"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop25"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop25"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop26"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop26"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop26"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop27"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop27"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop27"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop28"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop28"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop28"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop29"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop29"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop29"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop30"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop30"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop30"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop31"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop31"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop31"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop32"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop32"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop32"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop33"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop33"
+ }
+ ],
+ "colorScheme": {
+ "foundation": {
+ "fill": "#95a5a6",
+ "stroke": "#7f8c8d"
+ },
+ "definition": {
+ "fill": "#3498db",
+ "stroke": "#2980b9"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/euclid-elements-book-vii.json b/data/euclid-elements-book-vii.json
new file mode 100644
index 0000000000000000000000000000000000000000..50d94f40cc7cd95e4e0f8fabd625da005affe6c5
--- /dev/null
+++ b/data/euclid-elements-book-vii.json
@@ -0,0 +1,761 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-vii",
+ "name": "Euclid's Elements, Book VII",
+ "subject": "number_theory",
+ "variant": "classical",
+ "description": "Number theory: GCD (Euclidean algorithm), proportions, primes, LCM. 22 definitions, 39 propositions. Does not depend on previous books. Source: David E. Joyce.",
+ "structure": {
+ "books": 7,
+ "definitions": 22,
+ "propositions": 39,
+ "foundationTypes": [
+ "definition"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-18",
+ "lastUpdated": "2026-03-18",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book VII Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book VII",
+ "number theory",
+ "GCD",
+ "prime",
+ "LCM"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book VII",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookVII/bookVII.html",
+ "notes": "Clark University"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "Def1",
+ "type": "definition",
+ "label": "A unit is that by virtue of which each of the things that exist is called one",
+ "shortLabel": "Def. VII.1",
+ "short": "Unit",
+ "book": 7,
+ "number": 1,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def2",
+ "type": "definition",
+ "label": "A number is a multitude composed of units",
+ "shortLabel": "Def. VII.2",
+ "short": "Number",
+ "book": 7,
+ "number": 2,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def3",
+ "type": "definition",
+ "label": "A number is part of a number when it measures it",
+ "shortLabel": "Def. VII.3",
+ "short": "Part",
+ "book": 7,
+ "number": 3,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def4",
+ "type": "definition",
+ "label": "Parts when it does not measure it",
+ "shortLabel": "Def. VII.4",
+ "short": "Parts",
+ "book": 7,
+ "number": 4,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def5",
+ "type": "definition",
+ "label": "The greater is a multiple of the less when measured by the less",
+ "shortLabel": "Def. VII.5",
+ "short": "Multiple",
+ "book": 7,
+ "number": 5,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def6",
+ "type": "definition",
+ "label": "An even number is that which is divisible into two equal parts",
+ "shortLabel": "Def. VII.6",
+ "short": "Even",
+ "book": 7,
+ "number": 6,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def7",
+ "type": "definition",
+ "label": "An odd number is that which is not divisible into two equal parts",
+ "shortLabel": "Def. VII.7",
+ "short": "Odd",
+ "book": 7,
+ "number": 7,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def8",
+ "type": "definition",
+ "label": "Even-times even: measured by an even number an even number of times",
+ "shortLabel": "Def. VII.8",
+ "short": "Even-times even",
+ "book": 7,
+ "number": 8,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def9",
+ "type": "definition",
+ "label": "Even-times odd: measured by an even number an odd number of times",
+ "shortLabel": "Def. VII.9",
+ "short": "Even-times odd",
+ "book": 7,
+ "number": 9,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def10",
+ "type": "definition",
+ "label": "Odd-times odd: measured by an odd number an odd number of times",
+ "shortLabel": "Def. VII.10",
+ "short": "Odd-times odd",
+ "book": 7,
+ "number": 10,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def11",
+ "type": "definition",
+ "label": "A prime number is that which is measured by a unit alone",
+ "shortLabel": "Def. VII.11",
+ "short": "Prime",
+ "book": 7,
+ "number": 11,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def12",
+ "type": "definition",
+ "label": "Numbers relatively prime when only a unit measures both",
+ "shortLabel": "Def. VII.12",
+ "short": "Relatively prime",
+ "book": 7,
+ "number": 12,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def13",
+ "type": "definition",
+ "label": "A composite number is that measured by some number",
+ "shortLabel": "Def. VII.13",
+ "short": "Composite",
+ "book": 7,
+ "number": 13,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def14",
+ "type": "definition",
+ "label": "Numbers composite to one another when some number measures both",
+ "shortLabel": "Def. VII.14",
+ "short": "Composite to one another",
+ "book": 7,
+ "number": 14,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def15",
+ "type": "definition",
+ "label": "A number multiplies a number when the latter is added as many times as units in the former",
+ "shortLabel": "Def. VII.15",
+ "short": "Multiply",
+ "book": 7,
+ "number": 15,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def16",
+ "type": "definition",
+ "label": "When two numbers multiplied produce a number, the product is plane",
+ "shortLabel": "Def. VII.16",
+ "short": "Product",
+ "book": 7,
+ "number": 16,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def17",
+ "type": "definition",
+ "label": "Sides of the product are the numbers multiplied",
+ "shortLabel": "Def. VII.17",
+ "short": "Side",
+ "book": 7,
+ "number": 17,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def18",
+ "type": "definition",
+ "label": "A plane number is that produced by two numbers",
+ "shortLabel": "Def. VII.18",
+ "short": "Plane number",
+ "book": 7,
+ "number": 18,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def19",
+ "type": "definition",
+ "label": "A solid number is that produced by three numbers",
+ "shortLabel": "Def. VII.19",
+ "short": "Solid number",
+ "book": 7,
+ "number": 19,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def20",
+ "type": "definition",
+ "label": "Similar plane numbers have sides proportional",
+ "shortLabel": "Def. VII.20",
+ "short": "Similar plane",
+ "book": 7,
+ "number": 20,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def21",
+ "type": "definition",
+ "label": "Similar solid numbers have sides proportional",
+ "shortLabel": "Def. VII.21",
+ "short": "Similar solid",
+ "book": 7,
+ "number": 21,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Def22",
+ "type": "definition",
+ "label": "A perfect number is that which equals its own parts",
+ "shortLabel": "Def. VII.22",
+ "short": "Perfect",
+ "book": 7,
+ "number": 22,
+ "colorClass": "definition"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "Unequal numbers: repeated subtraction; if unit left, relatively prime",
+ "shortLabel": "Prop. VII.1",
+ "short": "Antenaresis, relatively prime",
+ "book": 7,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "To find greatest common measure of two numbers not relatively prime",
+ "shortLabel": "Prop. VII.2",
+ "short": "GCD of two numbers",
+ "book": 7,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "To find greatest common measure of three numbers",
+ "shortLabel": "Prop. VII.3",
+ "short": "GCD of three numbers",
+ "book": 7,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "Any number is part or parts of any number, less of greater",
+ "shortLabel": "Prop. VII.4",
+ "short": "Part or parts",
+ "book": 7,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "If a is same part of b as c of d, then a+c same part of b+d",
+ "shortLabel": "Prop. VII.5",
+ "short": "Same part: sum",
+ "book": 7,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "If a is same parts of b as c of d, then a+c same parts of b+d",
+ "shortLabel": "Prop. VII.6",
+ "short": "Same parts: sum",
+ "book": 7,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "If a part of b as c of d, remainder same part of remainder",
+ "shortLabel": "Prop. VII.7",
+ "short": "Same part: remainder",
+ "book": 7,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "If a parts of b as c of d, remainder same parts of remainder",
+ "shortLabel": "Prop. VII.8",
+ "short": "Same parts: remainder",
+ "book": 7,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "If a part of b as c of d, alternately a part/parts of c as b of d",
+ "shortLabel": "Prop. VII.9",
+ "short": "Same part: alternately",
+ "book": 7,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "If a parts of b as c of d, alternately a part/parts of c as b of d",
+ "shortLabel": "Prop. VII.10",
+ "short": "Same parts: alternately",
+ "book": 7,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "If whole:whole as subtracted:subtracted, remainder:remainder as whole:whole",
+ "shortLabel": "Prop. VII.11",
+ "short": "Proportion: remainder",
+ "book": 7,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "Proportional: one antecedent to consequent as sum antecedents to sum consequents",
+ "shortLabel": "Prop. VII.12",
+ "short": "Proportional: sum",
+ "book": 7,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "If four numbers proportional, also proportional alternately",
+ "shortLabel": "Prop. VII.13",
+ "short": "Proportional: alternately",
+ "book": 7,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "If a:b = d:e and b:c = e:f, then a:c = d:f",
+ "shortLabel": "Prop. VII.14",
+ "short": "Ex aequali",
+ "book": 7,
+ "number": 14,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop15",
+ "type": "proposition",
+ "label": "If unit measures a, b measures c same times, alternately unit:c as b:d",
+ "shortLabel": "Prop. VII.15",
+ "short": "Unit measures",
+ "book": 7,
+ "number": 15,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop16",
+ "type": "proposition",
+ "label": "If a×b and c×d, then a×b = c×d (commutativity)",
+ "shortLabel": "Prop. VII.16",
+ "short": "Commutativity of product",
+ "book": 7,
+ "number": 16,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop17",
+ "type": "proposition",
+ "label": "a:b = (a×c):(b×c)",
+ "shortLabel": "Prop. VII.17",
+ "short": "Ratio of products",
+ "book": 7,
+ "number": 17,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop18",
+ "type": "proposition",
+ "label": "a×c : b×c = a:b",
+ "shortLabel": "Prop. VII.18",
+ "short": "Ratio: multipliers",
+ "book": 7,
+ "number": 18,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop19",
+ "type": "proposition",
+ "label": "a:b = c:d iff a×d = b×c",
+ "shortLabel": "Prop. VII.19",
+ "short": "Proportional iff product",
+ "book": 7,
+ "number": 19,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop20",
+ "type": "proposition",
+ "label": "Least numbers in ratio measure others same number of times",
+ "shortLabel": "Prop. VII.20",
+ "short": "Least in ratio",
+ "book": 7,
+ "number": 20,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop21",
+ "type": "proposition",
+ "label": "Relatively prime numbers are least in their ratio",
+ "shortLabel": "Prop. VII.21",
+ "short": "Relatively prime: least",
+ "book": 7,
+ "number": 21,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop22",
+ "type": "proposition",
+ "label": "Least numbers in ratio are relatively prime",
+ "shortLabel": "Prop. VII.22",
+ "short": "Least: relatively prime",
+ "book": 7,
+ "number": 22,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop23",
+ "type": "proposition",
+ "label": "If a,b relatively prime, divisor of a relatively prime to b",
+ "shortLabel": "Prop. VII.23",
+ "short": "Relatively prime: divisor",
+ "book": 7,
+ "number": 23,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop24",
+ "type": "proposition",
+ "label": "If a,b relatively prime to c, then a×b relatively prime to c",
+ "shortLabel": "Prop. VII.24",
+ "short": "Product relatively prime",
+ "book": 7,
+ "number": 24,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop25",
+ "type": "proposition",
+ "label": "If a,b relatively prime, a² relatively prime to b",
+ "shortLabel": "Prop. VII.25",
+ "short": "Square relatively prime",
+ "book": 7,
+ "number": 25,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop26",
+ "type": "proposition",
+ "label": "If a,c and b,d relatively prime, a×b, c×d relatively prime",
+ "shortLabel": "Prop. VII.26",
+ "short": "Products relatively prime",
+ "book": 7,
+ "number": 26,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop27",
+ "type": "proposition",
+ "label": "If a,b relatively prime, a²,b² relatively prime; a×a², b×b²",
+ "shortLabel": "Prop. VII.27",
+ "short": "Squares relatively prime",
+ "book": 7,
+ "number": 27,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop28",
+ "type": "proposition",
+ "label": "If a,b relatively prime, a+b prime to each; converse",
+ "shortLabel": "Prop. VII.28",
+ "short": "Sum relatively prime",
+ "book": 7,
+ "number": 28,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop29",
+ "type": "proposition",
+ "label": "Prime relatively prime to any number it does not measure",
+ "shortLabel": "Prop. VII.29",
+ "short": "Prime to non-multiple",
+ "book": 7,
+ "number": 29,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop30",
+ "type": "proposition",
+ "label": "If prime measures product, it measures one factor",
+ "shortLabel": "Prop. VII.30",
+ "short": "Prime divides product",
+ "book": 7,
+ "number": 30,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop31",
+ "type": "proposition",
+ "label": "Any composite measured by some prime",
+ "shortLabel": "Prop. VII.31",
+ "short": "Composite has prime factor",
+ "book": 7,
+ "number": 31,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop32",
+ "type": "proposition",
+ "label": "Any number is prime or measured by some prime",
+ "shortLabel": "Prop. VII.32",
+ "short": "Prime or has prime factor",
+ "book": 7,
+ "number": 32,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop33",
+ "type": "proposition",
+ "label": "Given numbers, find least in same ratio",
+ "shortLabel": "Prop. VII.33",
+ "short": "Least in ratio",
+ "book": 7,
+ "number": 33,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop34",
+ "type": "proposition",
+ "label": "To find least number that two given numbers measure",
+ "shortLabel": "Prop. VII.34",
+ "short": "LCM of two",
+ "book": 7,
+ "number": 34,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop35",
+ "type": "proposition",
+ "label": "If two numbers measure some number, LCM also measures it",
+ "shortLabel": "Prop. VII.35",
+ "short": "LCM divides common multiple",
+ "book": 7,
+ "number": 35,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop36",
+ "type": "proposition",
+ "label": "To find least number that three given numbers measure",
+ "shortLabel": "Prop. VII.36",
+ "short": "LCM of three",
+ "book": 7,
+ "number": 36,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop37",
+ "type": "proposition",
+ "label": "If a measures b, b has part named by a",
+ "shortLabel": "Prop. VII.37",
+ "short": "Measured has part",
+ "book": 7,
+ "number": 37,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop38",
+ "type": "proposition",
+ "label": "If b has part named by a, a measures b",
+ "shortLabel": "Prop. VII.38",
+ "short": "Part implies measured",
+ "book": 7,
+ "number": 38,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop39",
+ "type": "proposition",
+ "label": "To find least number with given parts",
+ "shortLabel": "Prop. VII.39",
+ "short": "Least with given parts",
+ "book": 7,
+ "number": 39,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "Prop1",
+ "to": "Prop2"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop3"
+ },
+ {
+ "from": "Prop2",
+ "to": "Prop3"
+ },
+ {
+ "from": "Prop19",
+ "to": "Prop20"
+ },
+ {
+ "from": "Prop20",
+ "to": "Prop21"
+ },
+ {
+ "from": "Prop21",
+ "to": "Prop22"
+ },
+ {
+ "from": "Prop22",
+ "to": "Prop23"
+ },
+ {
+ "from": "Prop23",
+ "to": "Prop24"
+ },
+ {
+ "from": "Prop23",
+ "to": "Prop25"
+ },
+ {
+ "from": "Prop24",
+ "to": "Prop26"
+ },
+ {
+ "from": "Prop25",
+ "to": "Prop27"
+ },
+ {
+ "from": "Prop23",
+ "to": "Prop28"
+ },
+ {
+ "from": "Prop23",
+ "to": "Prop29"
+ },
+ {
+ "from": "Prop29",
+ "to": "Prop30"
+ },
+ {
+ "from": "Prop31",
+ "to": "Prop32"
+ },
+ {
+ "from": "Prop20",
+ "to": "Prop33"
+ },
+ {
+ "from": "Prop22",
+ "to": "Prop33"
+ },
+ {
+ "from": "Prop33",
+ "to": "Prop34"
+ },
+ {
+ "from": "Prop34",
+ "to": "Prop35"
+ },
+ {
+ "from": "Prop34",
+ "to": "Prop36"
+ },
+ {
+ "from": "Prop37",
+ "to": "Prop38"
+ },
+ {
+ "from": "Prop38",
+ "to": "Prop39"
+ }
+ ],
+ "colorScheme": {
+ "definition": {
+ "fill": "#3498db",
+ "stroke": "#2980b9"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/euclid-elements-book-viii.json b/data/euclid-elements-book-viii.json
new file mode 100644
index 0000000000000000000000000000000000000000..8bf2f041fd4ad2db9d67e897f5f74f329a80b447
--- /dev/null
+++ b/data/euclid-elements-book-viii.json
@@ -0,0 +1,540 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-viii",
+ "name": "Euclid's Elements, Book VIII",
+ "subject": "number_theory",
+ "variant": "classical",
+ "description": "Continued proportions. 27 propositions. Depends on Book VII. Source: David E. Joyce.",
+ "structure": {
+ "books": 8,
+ "propositions": 27,
+ "foundationTypes": [
+ "foundation"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-18",
+ "lastUpdated": "2026-03-18",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book VIII Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book VIII",
+ "continued proportion",
+ "plane",
+ "solid"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book VIII",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookVIII/bookVIII.html",
+ "notes": "Clark University"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "BookVII",
+ "type": "foundation",
+ "label": "Book VII — Number theory",
+ "shortLabel": "Book VII",
+ "short": "Foundation",
+ "book": 7,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "Continued proportion, extremes relatively prime: least in ratio",
+ "shortLabel": "Prop. VIII.1",
+ "short": "Continued proportion, extremes prime",
+ "book": 8,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "To find numbers in continued proportion, least in given ratio",
+ "shortLabel": "Prop. VIII.2",
+ "short": "Find numbers in continued proportion",
+ "book": 8,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "Least in continued proportion: extremes relatively prime",
+ "shortLabel": "Prop. VIII.3",
+ "short": "Least: extremes prime",
+ "book": 8,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "Given ratios in least numbers, find least in continued proportion",
+ "shortLabel": "Prop. VIII.4",
+ "short": "Find continued proportion from ratios",
+ "book": 8,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "Plane numbers have ratio compounded of ratios of sides",
+ "shortLabel": "Prop. VIII.5",
+ "short": "Plane numbers: compound ratio",
+ "book": 8,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "Continued proportion: if first does not measure second, none measures another",
+ "shortLabel": "Prop. VIII.6",
+ "short": "First does not measure second",
+ "book": 8,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "Continued proportion: if first measures last, it measures second",
+ "shortLabel": "Prop. VIII.7",
+ "short": "First measures last",
+ "book": 8,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "Numbers between two in continued proportion correspond to ratios",
+ "shortLabel": "Prop. VIII.8",
+ "short": "Numbers between in proportion",
+ "book": 8,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "Two relatively prime: numbers between them as between each and unit",
+ "shortLabel": "Prop. VIII.9",
+ "short": "Relatively prime: numbers to unit",
+ "book": 8,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "Numbers between number and unit correspond to between two numbers",
+ "shortLabel": "Prop. VIII.10",
+ "short": "Numbers from unit",
+ "book": 8,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "Between two squares one mean proportional; duplicate ratio",
+ "shortLabel": "Prop. VIII.11",
+ "short": "Mean proportional of squares",
+ "book": 8,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "Between two cubes two mean proportionals; triplicate ratio",
+ "shortLabel": "Prop. VIII.12",
+ "short": "Two means between cubes",
+ "book": 8,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "Continued proportion: products proportional; products of products",
+ "shortLabel": "Prop. VIII.13",
+ "short": "Products proportional",
+ "book": 8,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "Square measures square iff side measures side",
+ "shortLabel": "Prop. VIII.14",
+ "short": "Square measures square",
+ "book": 8,
+ "number": 14,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop15",
+ "type": "proposition",
+ "label": "Cube measures cube iff side measures side",
+ "shortLabel": "Prop. VIII.15",
+ "short": "Cube measures cube",
+ "book": 8,
+ "number": 15,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop16",
+ "type": "proposition",
+ "label": "Square does not measure square iff side does not measure side",
+ "shortLabel": "Prop. VIII.16",
+ "short": "Square does not measure square",
+ "book": 8,
+ "number": 16,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop17",
+ "type": "proposition",
+ "label": "Cube does not measure cube iff side does not measure side",
+ "shortLabel": "Prop. VIII.17",
+ "short": "Cube does not measure cube",
+ "book": 8,
+ "number": 17,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop18",
+ "type": "proposition",
+ "label": "Between similar plane numbers one mean proportional; duplicate ratio",
+ "shortLabel": "Prop. VIII.18",
+ "short": "Similar plane: mean proportional",
+ "book": 8,
+ "number": 18,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop19",
+ "type": "proposition",
+ "label": "Between similar solid numbers two mean proportionals; triplicate ratio",
+ "shortLabel": "Prop. VIII.19",
+ "short": "Similar solid: two means",
+ "book": 8,
+ "number": 19,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop20",
+ "type": "proposition",
+ "label": "If one mean between two numbers, they are similar plane",
+ "shortLabel": "Prop. VIII.20",
+ "short": "One mean: similar plane",
+ "book": 8,
+ "number": 20,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop21",
+ "type": "proposition",
+ "label": "If two means between two numbers, they are similar solid",
+ "shortLabel": "Prop. VIII.21",
+ "short": "Two means: similar solid",
+ "book": 8,
+ "number": 21,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop22",
+ "type": "proposition",
+ "label": "Three in continued proportion, first square: third square",
+ "shortLabel": "Prop. VIII.22",
+ "short": "Three in proportion: first square",
+ "book": 8,
+ "number": 22,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop23",
+ "type": "proposition",
+ "label": "Four in continued proportion, first cube: fourth cube",
+ "shortLabel": "Prop. VIII.23",
+ "short": "Four in proportion: first cube",
+ "book": 8,
+ "number": 23,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop24",
+ "type": "proposition",
+ "label": "If ratio as square to square and first square, second square",
+ "shortLabel": "Prop. VIII.24",
+ "short": "Ratio as square to square",
+ "book": 8,
+ "number": 24,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop25",
+ "type": "proposition",
+ "label": "If ratio as cube to cube and first cube, second cube",
+ "shortLabel": "Prop. VIII.25",
+ "short": "Ratio as cube to cube",
+ "book": 8,
+ "number": 25,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop26",
+ "type": "proposition",
+ "label": "Similar plane numbers have ratio of square to square",
+ "shortLabel": "Prop. VIII.26",
+ "short": "Similar plane: square ratio",
+ "book": 8,
+ "number": 26,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop27",
+ "type": "proposition",
+ "label": "Similar solid numbers have ratio of cube to cube",
+ "shortLabel": "Prop. VIII.27",
+ "short": "Similar solid: cube ratio",
+ "book": 8,
+ "number": 27,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "BookVII",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop2"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop3"
+ },
+ {
+ "from": "Prop2",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop4"
+ },
+ {
+ "from": "Prop2",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop6"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop7"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop9"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop10"
+ },
+ {
+ "from": "Prop9",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop11"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop12"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop14"
+ },
+ {
+ "from": "Prop11",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop15"
+ },
+ {
+ "from": "Prop12",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop16"
+ },
+ {
+ "from": "Prop14",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop17"
+ },
+ {
+ "from": "Prop15",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop18"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop19"
+ },
+ {
+ "from": "Prop8",
+ "to": "Prop19"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop20"
+ },
+ {
+ "from": "Prop18",
+ "to": "Prop20"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop21"
+ },
+ {
+ "from": "Prop19",
+ "to": "Prop21"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop22"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop22"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop23"
+ },
+ {
+ "from": "Prop1",
+ "to": "Prop23"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop24"
+ },
+ {
+ "from": "Prop22",
+ "to": "Prop24"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop25"
+ },
+ {
+ "from": "Prop23",
+ "to": "Prop25"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop26"
+ },
+ {
+ "from": "Prop18",
+ "to": "Prop26"
+ },
+ {
+ "from": "BookVII",
+ "to": "Prop27"
+ },
+ {
+ "from": "Prop19",
+ "to": "Prop27"
+ }
+ ],
+ "colorScheme": {
+ "foundation": {
+ "fill": "#95a5a6",
+ "stroke": "#7f8c8d"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/euclid-elements-book-x.json b/data/euclid-elements-book-x.json
new file mode 100644
index 0000000000000000000000000000000000000000..3ee4513a6245bcb2a8b5445ec40a8269803fd76a
--- /dev/null
+++ b/data/euclid-elements-book-x.json
@@ -0,0 +1,2620 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-x",
+ "name": "Euclid's Elements, Book X",
+ "subject": "incommensurables",
+ "variant": "classical",
+ "description": "Incommensurables, binomials, apotomes. 16 definitions, 115 propositions. Depends on Books I, V, VI. Source: David E. Joyce.",
+ "structure": {
+ "books": 10,
+ "definitions": 16,
+ "propositions": 115,
+ "foundationTypes": [
+ "foundation"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-18",
+ "lastUpdated": "2026-03-18",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book X Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book X",
+ "incommensurable",
+ "binomial",
+ "apotome",
+ "medial"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book X",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookX/bookX.html",
+ "notes": "Clark University"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "BookI",
+ "type": "foundation",
+ "label": "Book I — Plane geometry",
+ "shortLabel": "Book I",
+ "short": "Foundation",
+ "book": 1,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookV",
+ "type": "foundation",
+ "label": "Book V — Proportions",
+ "shortLabel": "Book V",
+ "short": "Foundation",
+ "book": 5,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookVI",
+ "type": "foundation",
+ "label": "Book VI — Similar figures",
+ "shortLabel": "Book VI",
+ "short": "Foundation",
+ "book": 6,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "Two unequal magnitudes: subtract more than half repeatedly; remainder less than lesser",
+ "shortLabel": "Prop. X.1",
+ "short": "Antenaresis for magnitudes",
+ "book": 10,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "If less subtracted from greater repeatedly and remainder never measures prior: incommensurable",
+ "shortLabel": "Prop. X.2",
+ "short": "Incommensurable criterion",
+ "book": 10,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "To find greatest common measure of two commensurable magnitudes",
+ "shortLabel": "Prop. X.3",
+ "short": "GCM of two commensurable",
+ "book": 10,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "To find greatest common measure of three commensurable magnitudes",
+ "shortLabel": "Prop. X.4",
+ "short": "GCM of three commensurable",
+ "book": 10,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "Commensurable magnitudes have ratio which a number has to a number",
+ "shortLabel": "Prop. X.5",
+ "short": "Commensurable: number ratio",
+ "book": 10,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "If two magnitudes have number-to-number ratio, they are commensurable",
+ "shortLabel": "Prop. X.6",
+ "short": "Number ratio: commensurable",
+ "book": 10,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "Incommensurable magnitudes do not have number-to-number ratio",
+ "shortLabel": "Prop. X.7",
+ "short": "Incommensurable: no number ratio",
+ "book": 10,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "If magnitudes lack number-to-number ratio, they are incommensurable",
+ "shortLabel": "Prop. X.8",
+ "short": "No number ratio: incommensurable",
+ "book": 10,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "Squares on commensurable lines have square-to-square ratio; converse",
+ "shortLabel": "Prop. X.9",
+ "short": "Squares: commensurable iff square ratio",
+ "book": 10,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "To find two lines incommensurable with assigned: one in length, one in square",
+ "shortLabel": "Prop. X.10",
+ "short": "Find incommensurable lines",
+ "book": 10,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "Four proportional: first commensurable with second iff third with fourth",
+ "shortLabel": "Prop. X.11",
+ "short": "Proportion preserves commensurability",
+ "book": 10,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "Magnitudes commensurable with same magnitude are commensurable",
+ "shortLabel": "Prop. X.12",
+ "short": "Commensurable with same",
+ "book": 10,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "If two commensurable, one incommensurable with third, so is the other",
+ "shortLabel": "Prop. X.13",
+ "short": "Commensurable: incommensurable partner",
+ "book": 10,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "Given two lines: find square by which greater square exceeds less; find line whose square equals sum",
+ "shortLabel": "Prop. X.14",
+ "short": "Square difference, sum of squares",
+ "book": 10,
+ "number": 14,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop15",
+ "type": "proposition",
+ "label": "Two commensurable added: whole commensurable with each",
+ "shortLabel": "Prop. X.15",
+ "short": "Commensurable sum",
+ "book": 10,
+ "number": 15,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop16",
+ "type": "proposition",
+ "label": "Two incommensurable added: sum incommensurable with each",
+ "shortLabel": "Prop. X.16",
+ "short": "Incommensurable sum",
+ "book": 10,
+ "number": 16,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop17",
+ "type": "proposition",
+ "label": "Applied parallelogram falling short by square: parts commensurable iff square difference commensurable",
+ "shortLabel": "Prop. X.17",
+ "short": "Parallelogram falling short",
+ "book": 10,
+ "number": 17,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop18",
+ "type": "proposition",
+ "label": "Applied parallelogram: parts incommensurable iff square difference incommensurable",
+ "shortLabel": "Prop. X.18",
+ "short": "Parallelogram: incommensurable parts",
+ "book": 10,
+ "number": 18,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop19",
+ "type": "proposition",
+ "label": "Rectangle by rational lines commensurable in length is rational",
+ "shortLabel": "Prop. X.19",
+ "short": "Rational rectangle from rational",
+ "book": 10,
+ "number": 19,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop20",
+ "type": "proposition",
+ "label": "Rational area applied to rational line: breadth rational and commensurable",
+ "shortLabel": "Prop. X.20",
+ "short": "Rational area on rational",
+ "book": 10,
+ "number": 20,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop21",
+ "type": "proposition",
+ "label": "Rectangle by rational in square only is irrational; side called medial",
+ "shortLabel": "Prop. X.21",
+ "short": "Medial: rational in square only",
+ "book": 10,
+ "number": 21,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop22",
+ "type": "proposition",
+ "label": "Square on medial applied to rational: breadth rational, incommensurable in length",
+ "shortLabel": "Prop. X.22",
+ "short": "Medial applied to rational",
+ "book": 10,
+ "number": 22,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop23",
+ "type": "proposition",
+ "label": "Line commensurable with medial is medial",
+ "shortLabel": "Prop. X.23",
+ "short": "Commensurable with medial",
+ "book": 10,
+ "number": 23,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop24",
+ "type": "proposition",
+ "label": "Rectangle by medial lines commensurable in length is medial",
+ "shortLabel": "Prop. X.24",
+ "short": "Medial rectangle: commensurable length",
+ "book": 10,
+ "number": 24,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop25",
+ "type": "proposition",
+ "label": "Rectangle by medial in square only: rational or medial",
+ "shortLabel": "Prop. X.25",
+ "short": "Medial: square only",
+ "book": 10,
+ "number": 25,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop26",
+ "type": "proposition",
+ "label": "Medial area does not exceed medial by rational area",
+ "shortLabel": "Prop. X.26",
+ "short": "Medial minus medial",
+ "book": 10,
+ "number": 26,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop27",
+ "type": "proposition",
+ "label": "To find medial lines commensurable in square only containing rational rectangle",
+ "shortLabel": "Prop. X.27",
+ "short": "Find medial in square only, rational rect",
+ "book": 10,
+ "number": 27,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop28",
+ "type": "proposition",
+ "label": "To find medial lines commensurable in square only containing medial rectangle",
+ "shortLabel": "Prop. X.28",
+ "short": "Find medial in square only, medial rect",
+ "book": 10,
+ "number": 28,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop29",
+ "type": "proposition",
+ "label": "To find two rational in square only: square diff commensurable with greater",
+ "shortLabel": "Prop. X.29",
+ "short": "Find rational in square only, commensurable diff",
+ "book": 10,
+ "number": 29,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop30",
+ "type": "proposition",
+ "label": "To find two rational in square only: square diff incommensurable with greater",
+ "shortLabel": "Prop. X.30",
+ "short": "Find rational in square only, incommensurable diff",
+ "book": 10,
+ "number": 30,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop31",
+ "type": "proposition",
+ "label": "To find two medial in square only, rational rect: diff commensurable with greater",
+ "shortLabel": "Prop. X.31",
+ "short": "Find medial, rational rect, commensurable diff",
+ "book": 10,
+ "number": 31,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop32",
+ "type": "proposition",
+ "label": "To find two medial in square only, medial rect: diff commensurable with greater",
+ "shortLabel": "Prop. X.32",
+ "short": "Find medial, medial rect, commensurable diff",
+ "book": 10,
+ "number": 32,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop33",
+ "type": "proposition",
+ "label": "To find two incommensurable in square: sum of squares rational, rectangle medial",
+ "shortLabel": "Prop. X.33",
+ "short": "Find incommensurable: rational sum, medial rect",
+ "book": 10,
+ "number": 33,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop34",
+ "type": "proposition",
+ "label": "To find two incommensurable in square: sum medial, rectangle rational",
+ "shortLabel": "Prop. X.34",
+ "short": "Find incommensurable: medial sum, rational rect",
+ "book": 10,
+ "number": 34,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop35",
+ "type": "proposition",
+ "label": "To find two incommensurable in square: sum and rect medial, incommensurable",
+ "shortLabel": "Prop. X.35",
+ "short": "Find incommensurable: both medial",
+ "book": 10,
+ "number": 35,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop36",
+ "type": "proposition",
+ "label": "Two rational in square only added: whole irrational, called binomial",
+ "shortLabel": "Prop. X.36",
+ "short": "Binomial defined",
+ "book": 10,
+ "number": 36,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop37",
+ "type": "proposition",
+ "label": "Two medial in square only, rational rect added: first bimedial",
+ "shortLabel": "Prop. X.37",
+ "short": "First bimedial defined",
+ "book": 10,
+ "number": 37,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop38",
+ "type": "proposition",
+ "label": "Two medial in square only, medial rect added: second bimedial",
+ "shortLabel": "Prop. X.38",
+ "short": "Second bimedial defined",
+ "book": 10,
+ "number": 38,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop39",
+ "type": "proposition",
+ "label": "Two incommensurable in square, rational sum, medial rect added: major",
+ "shortLabel": "Prop. X.39",
+ "short": "Major defined",
+ "book": 10,
+ "number": 39,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop40",
+ "type": "proposition",
+ "label": "Two incommensurable in square, medial sum, rational rect added: side of rational plus medial",
+ "shortLabel": "Prop. X.40",
+ "short": "Side of rational plus medial",
+ "book": 10,
+ "number": 40,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop41",
+ "type": "proposition",
+ "label": "Two incommensurable in square, both medial added: side of sum of two medial",
+ "shortLabel": "Prop. X.41",
+ "short": "Side of sum of two medial",
+ "book": 10,
+ "number": 41,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop42",
+ "type": "proposition",
+ "label": "Binomial divided into terms at one point only",
+ "shortLabel": "Prop. X.42",
+ "short": "Binomial: unique division",
+ "book": 10,
+ "number": 42,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop43",
+ "type": "proposition",
+ "label": "First bimedial divided at one point only",
+ "shortLabel": "Prop. X.43",
+ "short": "First bimedial: unique division",
+ "book": 10,
+ "number": 43,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop44",
+ "type": "proposition",
+ "label": "Second bimedial divided at one point only",
+ "shortLabel": "Prop. X.44",
+ "short": "Second bimedial: unique division",
+ "book": 10,
+ "number": 44,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop45",
+ "type": "proposition",
+ "label": "Major divided at one point only",
+ "shortLabel": "Prop. X.45",
+ "short": "Major: unique division",
+ "book": 10,
+ "number": 45,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop46",
+ "type": "proposition",
+ "label": "Side of rational plus medial divided at one point only",
+ "shortLabel": "Prop. X.46",
+ "short": "Side rational+medial: unique division",
+ "book": 10,
+ "number": 46,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop47",
+ "type": "proposition",
+ "label": "Side of sum of two medial divided at one point only",
+ "shortLabel": "Prop. X.47",
+ "short": "Side two medial: unique division",
+ "book": 10,
+ "number": 47,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop48",
+ "type": "proposition",
+ "label": "To find the first binomial line",
+ "shortLabel": "Prop. X.48",
+ "short": "Find first binomial",
+ "book": 10,
+ "number": 48,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop49",
+ "type": "proposition",
+ "label": "To find the second binomial line",
+ "shortLabel": "Prop. X.49",
+ "short": "Find second binomial",
+ "book": 10,
+ "number": 49,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop50",
+ "type": "proposition",
+ "label": "To find the third binomial line",
+ "shortLabel": "Prop. X.50",
+ "short": "Find third binomial",
+ "book": 10,
+ "number": 50,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop51",
+ "type": "proposition",
+ "label": "To find the fourth binomial line",
+ "shortLabel": "Prop. X.51",
+ "short": "Find fourth binomial",
+ "book": 10,
+ "number": 51,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop52",
+ "type": "proposition",
+ "label": "To find the fifth binomial line",
+ "shortLabel": "Prop. X.52",
+ "short": "Find fifth binomial",
+ "book": 10,
+ "number": 52,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop53",
+ "type": "proposition",
+ "label": "To find the sixth binomial line",
+ "shortLabel": "Prop. X.53",
+ "short": "Find sixth binomial",
+ "book": 10,
+ "number": 53,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop54",
+ "type": "proposition",
+ "label": "Area by rational and first binomial: side is binomial",
+ "shortLabel": "Prop. X.54",
+ "short": "Rational × first binomial",
+ "book": 10,
+ "number": 54,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop55",
+ "type": "proposition",
+ "label": "Area by rational and second binomial: side is first bimedial",
+ "shortLabel": "Prop. X.55",
+ "short": "Rational × second binomial",
+ "book": 10,
+ "number": 55,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop56",
+ "type": "proposition",
+ "label": "Area by rational and third binomial: side is second bimedial",
+ "shortLabel": "Prop. X.56",
+ "short": "Rational × third binomial",
+ "book": 10,
+ "number": 56,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop57",
+ "type": "proposition",
+ "label": "Area by rational and fourth binomial: side is major",
+ "shortLabel": "Prop. X.57",
+ "short": "Rational × fourth binomial",
+ "book": 10,
+ "number": 57,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop58",
+ "type": "proposition",
+ "label": "Area by rational and fifth binomial: side is rational plus medial",
+ "shortLabel": "Prop. X.58",
+ "short": "Rational × fifth binomial",
+ "book": 10,
+ "number": 58,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop59",
+ "type": "proposition",
+ "label": "Area by rational and sixth binomial: side is sum of two medial",
+ "shortLabel": "Prop. X.59",
+ "short": "Rational × sixth binomial",
+ "book": 10,
+ "number": 59,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop60",
+ "type": "proposition",
+ "label": "Square on binomial applied to rational: breadth first binomial",
+ "shortLabel": "Prop. X.60",
+ "short": "Square on binomial",
+ "book": 10,
+ "number": 60,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop61",
+ "type": "proposition",
+ "label": "Square on first bimedial applied to rational: breadth second binomial",
+ "shortLabel": "Prop. X.61",
+ "short": "Square on first bimedial",
+ "book": 10,
+ "number": 61,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop62",
+ "type": "proposition",
+ "label": "Square on second bimedial applied to rational: breadth third binomial",
+ "shortLabel": "Prop. X.62",
+ "short": "Square on second bimedial",
+ "book": 10,
+ "number": 62,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop63",
+ "type": "proposition",
+ "label": "Square on major applied to rational: breadth fourth binomial",
+ "shortLabel": "Prop. X.63",
+ "short": "Square on major",
+ "book": 10,
+ "number": 63,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop64",
+ "type": "proposition",
+ "label": "Square on rational+medial applied to rational: breadth fifth binomial",
+ "shortLabel": "Prop. X.64",
+ "short": "Square on rational+medial",
+ "book": 10,
+ "number": 64,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop65",
+ "type": "proposition",
+ "label": "Square on sum of two medial applied to rational: breadth sixth binomial",
+ "shortLabel": "Prop. X.65",
+ "short": "Square on two medial",
+ "book": 10,
+ "number": 65,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop66",
+ "type": "proposition",
+ "label": "Line commensurable with binomial is binomial, same order",
+ "shortLabel": "Prop. X.66",
+ "short": "Commensurable with binomial",
+ "book": 10,
+ "number": 66,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop67",
+ "type": "proposition",
+ "label": "Line commensurable with bimedial is bimedial, same order",
+ "shortLabel": "Prop. X.67",
+ "short": "Commensurable with bimedial",
+ "book": 10,
+ "number": 67,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop68",
+ "type": "proposition",
+ "label": "Line commensurable with major is major",
+ "shortLabel": "Prop. X.68",
+ "short": "Commensurable with major",
+ "book": 10,
+ "number": 68,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop69",
+ "type": "proposition",
+ "label": "Line commensurable with rational+medial is rational+medial",
+ "shortLabel": "Prop. X.69",
+ "short": "Commensurable with rational+medial",
+ "book": 10,
+ "number": 69,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop70",
+ "type": "proposition",
+ "label": "Line commensurable with sum of two medial is sum of two medial",
+ "shortLabel": "Prop. X.70",
+ "short": "Commensurable with two medial",
+ "book": 10,
+ "number": 70,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop71",
+ "type": "proposition",
+ "label": "Rational and medial added: four irrationals arise",
+ "shortLabel": "Prop. X.71",
+ "short": "Rational + medial: four irrationals",
+ "book": 10,
+ "number": 71,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop72",
+ "type": "proposition",
+ "label": "Two medial incommensurable added: two irrationals arise",
+ "shortLabel": "Prop. X.72",
+ "short": "Two medial: two irrationals",
+ "book": 10,
+ "number": 72,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop73",
+ "type": "proposition",
+ "label": "Rational minus rational in square only: remainder irrational, apotome",
+ "shortLabel": "Prop. X.73",
+ "short": "Apotome defined",
+ "book": 10,
+ "number": 73,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop74",
+ "type": "proposition",
+ "label": "Medial minus medial, rational rect: first apotome of medial",
+ "shortLabel": "Prop. X.74",
+ "short": "First apotome of medial",
+ "book": 10,
+ "number": 74,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop75",
+ "type": "proposition",
+ "label": "Medial minus medial, medial rect: second apotome of medial",
+ "shortLabel": "Prop. X.75",
+ "short": "Second apotome of medial",
+ "book": 10,
+ "number": 75,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop76",
+ "type": "proposition",
+ "label": "Line minus incommensurable: sum rational, rect medial: remainder minor",
+ "shortLabel": "Prop. X.76",
+ "short": "Minor defined",
+ "book": 10,
+ "number": 76,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop77",
+ "type": "proposition",
+ "label": "Line minus incommensurable: sum medial, rect rational: produces rational+medial",
+ "shortLabel": "Prop. X.77",
+ "short": "Produces rational+medial",
+ "book": 10,
+ "number": 77,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop78",
+ "type": "proposition",
+ "label": "Line minus incommensurable: both medial, incommensurable: produces medial+medial",
+ "shortLabel": "Prop. X.78",
+ "short": "Produces medial+medial",
+ "book": 10,
+ "number": 78,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop79",
+ "type": "proposition",
+ "label": "To apotome only one rational can be annexed in square only",
+ "shortLabel": "Prop. X.79",
+ "short": "Apotome: unique annex",
+ "book": 10,
+ "number": 79,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop80",
+ "type": "proposition",
+ "label": "To first apotome of medial: unique medial annex, rational rect",
+ "shortLabel": "Prop. X.80",
+ "short": "First apotome medial: unique annex",
+ "book": 10,
+ "number": 80,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop81",
+ "type": "proposition",
+ "label": "To second apotome of medial: unique medial annex, medial rect",
+ "shortLabel": "Prop. X.81",
+ "short": "Second apotome medial: unique annex",
+ "book": 10,
+ "number": 81,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop82",
+ "type": "proposition",
+ "label": "To minor: unique annex incommensurable in square",
+ "shortLabel": "Prop. X.82",
+ "short": "Minor: unique annex",
+ "book": 10,
+ "number": 82,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop83",
+ "type": "proposition",
+ "label": "To produces rational+medial: unique annex",
+ "shortLabel": "Prop. X.83",
+ "short": "Rational+medial: unique annex",
+ "book": 10,
+ "number": 83,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop84",
+ "type": "proposition",
+ "label": "To produces medial+medial: unique annex",
+ "shortLabel": "Prop. X.84",
+ "short": "Medial+medial: unique annex",
+ "book": 10,
+ "number": 84,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop85",
+ "type": "proposition",
+ "label": "To find the first apotome",
+ "shortLabel": "Prop. X.85",
+ "short": "Find first apotome",
+ "book": 10,
+ "number": 85,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop86",
+ "type": "proposition",
+ "label": "To find the second apotome",
+ "shortLabel": "Prop. X.86",
+ "short": "Find second apotome",
+ "book": 10,
+ "number": 86,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop87",
+ "type": "proposition",
+ "label": "To find the third apotome",
+ "shortLabel": "Prop. X.87",
+ "short": "Find third apotome",
+ "book": 10,
+ "number": 87,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop88",
+ "type": "proposition",
+ "label": "To find the fourth apotome",
+ "shortLabel": "Prop. X.88",
+ "short": "Find fourth apotome",
+ "book": 10,
+ "number": 88,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop89",
+ "type": "proposition",
+ "label": "To find the fifth apotome",
+ "shortLabel": "Prop. X.89",
+ "short": "Find fifth apotome",
+ "book": 10,
+ "number": 89,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop90",
+ "type": "proposition",
+ "label": "To find the sixth apotome",
+ "shortLabel": "Prop. X.90",
+ "short": "Find sixth apotome",
+ "book": 10,
+ "number": 90,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop91",
+ "type": "proposition",
+ "label": "Area by rational and first apotome: side is apotome",
+ "shortLabel": "Prop. X.91",
+ "short": "Rational × first apotome",
+ "book": 10,
+ "number": 91,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop92",
+ "type": "proposition",
+ "label": "Area by rational and second apotome: side is first apotome of medial",
+ "shortLabel": "Prop. X.92",
+ "short": "Rational × second apotome",
+ "book": 10,
+ "number": 92,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop93",
+ "type": "proposition",
+ "label": "Area by rational and third apotome: side is second apotome of medial",
+ "shortLabel": "Prop. X.93",
+ "short": "Rational × third apotome",
+ "book": 10,
+ "number": 93,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop94",
+ "type": "proposition",
+ "label": "Area by rational and fourth apotome: side is minor",
+ "shortLabel": "Prop. X.94",
+ "short": "Rational × fourth apotome",
+ "book": 10,
+ "number": 94,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop95",
+ "type": "proposition",
+ "label": "Area by rational and fifth apotome: side produces rational+medial",
+ "shortLabel": "Prop. X.95",
+ "short": "Rational × fifth apotome",
+ "book": 10,
+ "number": 95,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop96",
+ "type": "proposition",
+ "label": "Area by rational and sixth apotome: side produces medial+medial",
+ "shortLabel": "Prop. X.96",
+ "short": "Rational × sixth apotome",
+ "book": 10,
+ "number": 96,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop97",
+ "type": "proposition",
+ "label": "Square on apotome of medial applied to rational: breadth first apotome",
+ "shortLabel": "Prop. X.97",
+ "short": "Square on apotome medial",
+ "book": 10,
+ "number": 97,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop98",
+ "type": "proposition",
+ "label": "Square on first apotome of medial: breadth second apotome",
+ "shortLabel": "Prop. X.98",
+ "short": "Square on first apotome medial",
+ "book": 10,
+ "number": 98,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop99",
+ "type": "proposition",
+ "label": "Square on second apotome of medial: breadth third apotome",
+ "shortLabel": "Prop. X.99",
+ "short": "Square on second apotome medial",
+ "book": 10,
+ "number": 99,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop100",
+ "type": "proposition",
+ "label": "Square on minor applied to rational: breadth fourth apotome",
+ "shortLabel": "Prop. X.100",
+ "short": "Square on minor",
+ "book": 10,
+ "number": 100,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop101",
+ "type": "proposition",
+ "label": "Square on produces rational+medial: breadth fifth apotome",
+ "shortLabel": "Prop. X.101",
+ "short": "Square on rational+medial",
+ "book": 10,
+ "number": 101,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop102",
+ "type": "proposition",
+ "label": "Square on produces medial+medial: breadth sixth apotome",
+ "shortLabel": "Prop. X.102",
+ "short": "Square on medial+medial",
+ "book": 10,
+ "number": 102,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop103",
+ "type": "proposition",
+ "label": "Line commensurable with apotome is apotome, same order",
+ "shortLabel": "Prop. X.103",
+ "short": "Commensurable with apotome",
+ "book": 10,
+ "number": 103,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop104",
+ "type": "proposition",
+ "label": "Line commensurable with apotome of medial is apotome of medial",
+ "shortLabel": "Prop. X.104",
+ "short": "Commensurable with apotome medial",
+ "book": 10,
+ "number": 104,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop105",
+ "type": "proposition",
+ "label": "Line commensurable with minor is minor",
+ "shortLabel": "Prop. X.105",
+ "short": "Commensurable with minor",
+ "book": 10,
+ "number": 105,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop106",
+ "type": "proposition",
+ "label": "Line commensurable with produces rational+medial is same",
+ "shortLabel": "Prop. X.106",
+ "short": "Commensurable with rational+medial",
+ "book": 10,
+ "number": 106,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop107",
+ "type": "proposition",
+ "label": "Line commensurable with produces medial+medial is same",
+ "shortLabel": "Prop. X.107",
+ "short": "Commensurable with medial+medial",
+ "book": 10,
+ "number": 107,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop108",
+ "type": "proposition",
+ "label": "From rational area subtract medial: side is apotome or minor",
+ "shortLabel": "Prop. X.108",
+ "short": "Rational minus medial",
+ "book": 10,
+ "number": 108,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop109",
+ "type": "proposition",
+ "label": "From medial subtract rational: first apotome of medial or rational+medial",
+ "shortLabel": "Prop. X.109",
+ "short": "Medial minus rational",
+ "book": 10,
+ "number": 109,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop110",
+ "type": "proposition",
+ "label": "From medial subtract medial incommensurable: second apotome or medial+medial",
+ "shortLabel": "Prop. X.110",
+ "short": "Medial minus medial",
+ "book": 10,
+ "number": 110,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop111",
+ "type": "proposition",
+ "label": "Apotome is not the same as binomial",
+ "shortLabel": "Prop. X.111",
+ "short": "Apotome ≠ binomial",
+ "book": 10,
+ "number": 111,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop112",
+ "type": "proposition",
+ "label": "Square on rational applied to binomial: breadth apotome, same order",
+ "shortLabel": "Prop. X.112",
+ "short": "Rational on binomial",
+ "book": 10,
+ "number": 112,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop113",
+ "type": "proposition",
+ "label": "Square on rational applied to apotome: breadth binomial, same order",
+ "shortLabel": "Prop. X.113",
+ "short": "Rational on apotome",
+ "book": 10,
+ "number": 113,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop114",
+ "type": "proposition",
+ "label": "Area by apotome and binomial (commensurable terms): side is rational",
+ "shortLabel": "Prop. X.114",
+ "short": "Apotome × binomial: rational",
+ "book": 10,
+ "number": 114,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop115",
+ "type": "proposition",
+ "label": "From medial arise irrationals infinite in number, none same as preceding",
+ "shortLabel": "Prop. X.115",
+ "short": "Medial: infinite irrationals",
+ "book": 10,
+ "number": 115,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "BookI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop19"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop19"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop19"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop20"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop20"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop20"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop21"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop21"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop21"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop22"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop22"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop22"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop23"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop23"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop23"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop24"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop24"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop24"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop25"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop25"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop25"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop26"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop26"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop26"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop27"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop27"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop27"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop28"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop28"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop28"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop29"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop29"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop29"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop30"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop30"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop30"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop31"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop31"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop31"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop32"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop32"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop32"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop33"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop33"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop33"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop34"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop34"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop34"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop35"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop35"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop35"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop36"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop36"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop36"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop37"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop37"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop37"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop38"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop38"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop38"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop39"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop39"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop39"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop40"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop40"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop40"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop41"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop41"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop41"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop42"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop42"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop42"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop43"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop43"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop43"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop44"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop44"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop44"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop45"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop45"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop45"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop46"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop46"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop46"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop47"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop47"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop47"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop48"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop48"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop48"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop49"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop49"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop49"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop50"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop50"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop50"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop51"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop51"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop51"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop52"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop52"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop52"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop53"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop53"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop53"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop54"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop54"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop54"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop55"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop55"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop55"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop56"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop56"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop56"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop57"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop57"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop57"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop58"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop58"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop58"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop59"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop59"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop59"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop60"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop60"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop60"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop61"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop61"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop61"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop62"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop62"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop62"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop63"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop63"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop63"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop64"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop64"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop64"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop65"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop65"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop65"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop66"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop66"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop66"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop67"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop67"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop67"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop68"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop68"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop68"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop69"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop69"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop69"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop70"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop70"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop70"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop71"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop71"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop71"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop72"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop72"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop72"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop73"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop73"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop73"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop74"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop74"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop74"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop75"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop75"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop75"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop76"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop76"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop76"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop77"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop77"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop77"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop78"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop78"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop78"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop79"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop79"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop79"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop80"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop80"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop80"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop81"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop81"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop81"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop82"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop82"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop82"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop83"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop83"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop83"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop84"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop84"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop84"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop85"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop85"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop85"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop86"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop86"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop86"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop87"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop87"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop87"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop88"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop88"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop88"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop89"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop89"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop89"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop90"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop90"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop90"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop91"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop91"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop91"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop92"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop92"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop92"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop93"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop93"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop93"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop94"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop94"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop94"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop95"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop95"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop95"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop96"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop96"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop96"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop97"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop97"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop97"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop98"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop98"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop98"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop99"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop99"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop99"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop100"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop100"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop100"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop101"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop101"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop101"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop102"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop102"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop102"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop103"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop103"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop103"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop104"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop104"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop104"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop105"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop105"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop105"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop106"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop106"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop106"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop107"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop107"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop107"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop108"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop108"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop108"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop109"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop109"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop109"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop110"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop110"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop110"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop111"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop111"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop111"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop112"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop112"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop112"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop113"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop113"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop113"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop114"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop114"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop114"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop115"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop115"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop115"
+ }
+ ],
+ "colorScheme": {
+ "foundation": {
+ "fill": "#95a5a6",
+ "stroke": "#7f8c8d"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/euclid-elements-book-xi.json b/data/euclid-elements-book-xi.json
new file mode 100644
index 0000000000000000000000000000000000000000..44ad6c94b87a26b702b12bcb5f07f9081205ed43
--- /dev/null
+++ b/data/euclid-elements-book-xi.json
@@ -0,0 +1,783 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-xi",
+ "name": "Euclid's Elements, Book XI",
+ "subject": "solid_geometry",
+ "variant": "classical",
+ "description": "Solid geometry: planes, perpendiculars, parallelepipeds, prisms. 28 definitions, 39 propositions. Depends on Books I and VI. Source: David E. Joyce.",
+ "structure": {
+ "books": 11,
+ "definitions": 28,
+ "propositions": 39,
+ "foundationTypes": [
+ "foundation"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-18",
+ "lastUpdated": "2026-03-18",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book XI Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book XI",
+ "solid geometry",
+ "plane",
+ "parallelepiped",
+ "prism"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book XI",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookXI/bookXI.html",
+ "notes": "Clark University"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "BookI",
+ "type": "foundation",
+ "label": "Book I — Plane geometry",
+ "shortLabel": "Book I",
+ "short": "Foundation",
+ "book": 1,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookVI",
+ "type": "foundation",
+ "label": "Book VI — Similar figures",
+ "shortLabel": "Book VI",
+ "short": "Foundation",
+ "book": 6,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "A part of a straight line cannot be in one plane and part in another elevated",
+ "shortLabel": "Prop. XI.1",
+ "short": "Line part in plane",
+ "book": 11,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "If two straight lines cut one another, they lie in one plane; every triangle in one plane",
+ "shortLabel": "Prop. XI.2",
+ "short": "Two lines cut: one plane",
+ "book": 11,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "If two planes cut one another, their intersection is a straight line",
+ "shortLabel": "Prop. XI.3",
+ "short": "Planes cut: line",
+ "book": 11,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "If line at right angles to two lines cutting at point, also perpendicular to plane through them",
+ "shortLabel": "Prop. XI.4",
+ "short": "Line perpendicular to plane",
+ "book": 11,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "If line at right angles to three lines meeting at point, the three lie in one plane",
+ "shortLabel": "Prop. XI.5",
+ "short": "Three lines from point",
+ "book": 11,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "If two lines at right angles to same plane, they are parallel",
+ "shortLabel": "Prop. XI.6",
+ "short": "Perpendicular to same plane: parallel",
+ "book": 11,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "If two lines parallel, line joining points on each is in same plane",
+ "shortLabel": "Prop. XI.7",
+ "short": "Parallel lines: join in plane",
+ "book": 11,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "If two lines parallel, one perpendicular to plane, so is the other",
+ "shortLabel": "Prop. XI.8",
+ "short": "Parallel: one perpendicular",
+ "book": 11,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "Lines parallel to same line but not in same plane are parallel to each other",
+ "shortLabel": "Prop. XI.9",
+ "short": "Parallel to same: parallel",
+ "book": 11,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "Two lines meeting parallel to two meeting not in same plane: contain equal angles",
+ "shortLabel": "Prop. XI.10",
+ "short": "Skew lines: equal angles",
+ "book": 11,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "To draw line perpendicular to given plane from given elevated point",
+ "shortLabel": "Prop. XI.11",
+ "short": "Perpendicular from point to plane",
+ "book": 11,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "To set up line at right angles to plane from given point in it",
+ "shortLabel": "Prop. XI.12",
+ "short": "Perpendicular from point in plane",
+ "book": 11,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "From same point two lines cannot be perpendicular to same plane on same side",
+ "shortLabel": "Prop. XI.13",
+ "short": "One perpendicular only",
+ "book": 11,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "Planes to which same line is perpendicular are parallel",
+ "shortLabel": "Prop. XI.14",
+ "short": "Planes perpendicular to line: parallel",
+ "book": 11,
+ "number": 14,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop15",
+ "type": "proposition",
+ "label": "Two lines meeting parallel to two meeting not in same plane: planes through them parallel",
+ "shortLabel": "Prop. XI.15",
+ "short": "Skew lines: planes parallel",
+ "book": 11,
+ "number": 15,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop16",
+ "type": "proposition",
+ "label": "If two parallel planes cut by any plane, intersections are parallel",
+ "shortLabel": "Prop. XI.16",
+ "short": "Parallel planes cut: parallel",
+ "book": 11,
+ "number": 16,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop17",
+ "type": "proposition",
+ "label": "If two lines cut by parallel planes, they are cut in same ratios",
+ "shortLabel": "Prop. XI.17",
+ "short": "Parallel planes: same ratio",
+ "book": 11,
+ "number": 17,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop18",
+ "type": "proposition",
+ "label": "If line perpendicular to plane, all planes through it perpendicular to that plane",
+ "shortLabel": "Prop. XI.18",
+ "short": "Line perpendicular: planes through it",
+ "book": 11,
+ "number": 18,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop19",
+ "type": "proposition",
+ "label": "If two planes cutting one another perpendicular to plane, intersection perpendicular",
+ "shortLabel": "Prop. XI.19",
+ "short": "Planes perpendicular: intersection",
+ "book": 11,
+ "number": 19,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop20",
+ "type": "proposition",
+ "label": "Solid angle by three plane angles: sum of any two greater than third",
+ "shortLabel": "Prop. XI.20",
+ "short": "Solid angle: plane angles",
+ "book": 11,
+ "number": 20,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop21",
+ "type": "proposition",
+ "label": "Any solid angle contained by plane angles summing to less than four right angles",
+ "shortLabel": "Prop. XI.21",
+ "short": "Solid angle: less than four right",
+ "book": 11,
+ "number": 21,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop22",
+ "type": "proposition",
+ "label": "Three plane angles with sum of any two greater than third, equal sides: construct triangle",
+ "shortLabel": "Prop. XI.22",
+ "short": "Three plane angles: construct triangle",
+ "book": 11,
+ "number": 22,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop23",
+ "type": "proposition",
+ "label": "To construct solid angle from three plane angles (sum of any two greater than third)",
+ "shortLabel": "Prop. XI.23",
+ "short": "Construct solid angle",
+ "book": 11,
+ "number": 23,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop24",
+ "type": "proposition",
+ "label": "If solid contained by parallel planes, opposite planes equal and parallelogrammic",
+ "shortLabel": "Prop. XI.24",
+ "short": "Solid by parallel planes",
+ "book": 11,
+ "number": 24,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop25",
+ "type": "proposition",
+ "label": "Parallelepiped cut by plane parallel to opposite: base to base as solid to solid",
+ "shortLabel": "Prop. XI.25",
+ "short": "Parallelepiped cut: base ratio",
+ "book": 11,
+ "number": 25,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop26",
+ "type": "proposition",
+ "label": "To construct solid angle equal to given on given line at given point",
+ "shortLabel": "Prop. XI.26",
+ "short": "Construct equal solid angle",
+ "book": 11,
+ "number": 26,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop27",
+ "type": "proposition",
+ "label": "To describe parallelepiped similar to given on given straight line",
+ "shortLabel": "Prop. XI.27",
+ "short": "Similar parallelepiped on line",
+ "book": 11,
+ "number": 27,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop28",
+ "type": "proposition",
+ "label": "Parallelepiped cut by plane through diagonals of opposite planes: bisected",
+ "shortLabel": "Prop. XI.28",
+ "short": "Parallelepiped: diagonal plane bisects",
+ "book": 11,
+ "number": 28,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop29",
+ "type": "proposition",
+ "label": "Parallelepipeds same base, height, ends on same lines: equal",
+ "shortLabel": "Prop. XI.29",
+ "short": "Same base, height, same lines: equal",
+ "book": 11,
+ "number": 29,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop30",
+ "type": "proposition",
+ "label": "Parallelepipeds same base, height, ends not on same lines: equal",
+ "shortLabel": "Prop. XI.30",
+ "short": "Same base, height, different lines: equal",
+ "book": 11,
+ "number": 30,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop31",
+ "type": "proposition",
+ "label": "Parallelepipeds on equal bases, same height: equal",
+ "shortLabel": "Prop. XI.31",
+ "short": "Equal bases, same height: equal",
+ "book": 11,
+ "number": 31,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop32",
+ "type": "proposition",
+ "label": "Parallelepipeds same height: to one another as bases",
+ "shortLabel": "Prop. XI.32",
+ "short": "Same height: as bases",
+ "book": 11,
+ "number": 32,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop33",
+ "type": "proposition",
+ "label": "Similar parallelepipeds: to one another in triplicate ratio of corresponding sides",
+ "shortLabel": "Prop. XI.33",
+ "short": "Similar: triplicate ratio",
+ "book": 11,
+ "number": 33,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop34",
+ "type": "proposition",
+ "label": "Equal parallelepipeds: bases reciprocally proportional to heights",
+ "shortLabel": "Prop. XI.34",
+ "short": "Equal: bases reciprocally proportional",
+ "book": 11,
+ "number": 34,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop35",
+ "type": "proposition",
+ "label": "Equal plane angles, elevated lines with equal angles: perpendiculars, joins",
+ "shortLabel": "Prop. XI.35",
+ "short": "Equal plane angles: elevated lines",
+ "book": 11,
+ "number": 35,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop36",
+ "type": "proposition",
+ "label": "Three proportional lines: parallelepiped from three equals that on mean equilateral",
+ "shortLabel": "Prop. XI.36",
+ "short": "Three proportional: parallelepiped",
+ "book": 11,
+ "number": 36,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop37",
+ "type": "proposition",
+ "label": "Four proportional: similar parallelepipeds proportional; converse",
+ "shortLabel": "Prop. XI.37",
+ "short": "Four proportional: parallelepipeds",
+ "book": 11,
+ "number": 37,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop38",
+ "type": "proposition",
+ "label": "Cube opposite sides bisected, planes through: intersection and diameter bisect each other",
+ "shortLabel": "Prop. XI.38",
+ "short": "Cube: bisected by planes",
+ "book": 11,
+ "number": 38,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop39",
+ "type": "proposition",
+ "label": "Two prisms equal height, parallelogram and triangle bases, parallelogram double: equal",
+ "shortLabel": "Prop. XI.39",
+ "short": "Prisms: parallelogram, triangle",
+ "book": 11,
+ "number": 39,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "BookI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop19"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop19"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop20"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop20"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop21"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop21"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop22"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop22"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop23"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop23"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop24"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop24"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop25"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop25"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop26"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop26"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop27"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop27"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop28"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop28"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop29"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop29"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop30"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop30"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop31"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop31"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop32"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop32"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop33"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop33"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop34"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop34"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop35"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop35"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop36"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop36"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop37"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop37"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop38"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop38"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop39"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop39"
+ }
+ ],
+ "colorScheme": {
+ "foundation": {
+ "fill": "#95a5a6",
+ "stroke": "#7f8c8d"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/euclid-elements-book-xii.json b/data/euclid-elements-book-xii.json
new file mode 100644
index 0000000000000000000000000000000000000000..c5ddc1e9f71196671578522ce792df9b9b42ee10
--- /dev/null
+++ b/data/euclid-elements-book-xii.json
@@ -0,0 +1,567 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-xii",
+ "name": "Euclid's Elements, Book XII",
+ "subject": "measurement",
+ "variant": "classical",
+ "description": "Measurement of figures: circles, pyramids, cones, cylinders, spheres. 18 propositions. Depends on Books I, V, VI, XI. Source: David E. Joyce.",
+ "structure": {
+ "books": 12,
+ "propositions": 18,
+ "foundationTypes": [
+ "foundation"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-18",
+ "lastUpdated": "2026-03-18",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book XII Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book XII",
+ "measurement",
+ "pyramid",
+ "cone",
+ "cylinder",
+ "sphere"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book XII",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookXII/bookXII.html",
+ "notes": "Clark University"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "BookI",
+ "type": "foundation",
+ "label": "Book I — Plane geometry",
+ "shortLabel": "Book I",
+ "short": "Foundation",
+ "book": 1,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookV",
+ "type": "foundation",
+ "label": "Book V — Proportions",
+ "shortLabel": "Book V",
+ "short": "Foundation",
+ "book": 5,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookVI",
+ "type": "foundation",
+ "label": "Book VI — Similar figures",
+ "shortLabel": "Book VI",
+ "short": "Foundation",
+ "book": 6,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookXI",
+ "type": "foundation",
+ "label": "Book XI — Solid geometry",
+ "shortLabel": "Book XI",
+ "short": "Foundation",
+ "book": 11,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "Similar polygons in circles: to one another as squares on diameters",
+ "shortLabel": "Prop. XII.1",
+ "short": "Similar polygons: as squares on diameters",
+ "book": 12,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "Circles are to one another as the squares on their diameters",
+ "shortLabel": "Prop. XII.2",
+ "short": "Circles: as squares on diameters",
+ "book": 12,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "Pyramid with triangular base: divided into two pyramids, two prisms; prisms greater than half",
+ "shortLabel": "Prop. XII.3",
+ "short": "Pyramid divided",
+ "book": 12,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "Two pyramids same height, triangular bases, divided: base to base as all prisms",
+ "shortLabel": "Prop. XII.4",
+ "short": "Pyramids: base as prisms",
+ "book": 12,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "Pyramids same height, triangular bases: to one another as bases",
+ "shortLabel": "Prop. XII.5",
+ "short": "Pyramids: as bases",
+ "book": 12,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "Pyramids same height, polygonal bases: to one another as bases",
+ "shortLabel": "Prop. XII.6",
+ "short": "Pyramids polygonal: as bases",
+ "book": 12,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "Prism with triangular base: divided into three equal pyramids",
+ "shortLabel": "Prop. XII.7",
+ "short": "Prism into three pyramids",
+ "book": 12,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "Similar pyramids triangular bases: in triplicate ratio of corresponding sides",
+ "shortLabel": "Prop. XII.8",
+ "short": "Similar pyramids: triplicate ratio",
+ "book": 12,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "Equal pyramids triangular bases: bases reciprocally proportional to heights",
+ "shortLabel": "Prop. XII.9",
+ "short": "Equal pyramids: bases reciprocally proportional",
+ "book": 12,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "Any cone is third part of cylinder same base and equal height",
+ "shortLabel": "Prop. XII.10",
+ "short": "Cone third of cylinder",
+ "book": 12,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "Cones and cylinders same height: to one another as bases",
+ "shortLabel": "Prop. XII.11",
+ "short": "Cones, cylinders: as bases",
+ "book": 12,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "Similar cones and cylinders: in triplicate ratio of diameters of bases",
+ "shortLabel": "Prop. XII.12",
+ "short": "Similar cones, cylinders: triplicate",
+ "book": 12,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "Cylinder cut by plane parallel to opposite: cylinder to cylinder as axis to axis",
+ "shortLabel": "Prop. XII.13",
+ "short": "Cylinder cut: as axes",
+ "book": 12,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "Cones and cylinders on equal bases: to one another as heights",
+ "shortLabel": "Prop. XII.14",
+ "short": "Cones, cylinders equal bases: as heights",
+ "book": 12,
+ "number": 14,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop15",
+ "type": "proposition",
+ "label": "Equal cones and cylinders: bases reciprocally proportional to heights",
+ "shortLabel": "Prop. XII.15",
+ "short": "Equal cones, cylinders: reciprocally proportional",
+ "book": 12,
+ "number": 15,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop16",
+ "type": "proposition",
+ "label": "Given two circles same center: inscribe in greater equilateral polygon even sides not touching lesser",
+ "shortLabel": "Prop. XII.16",
+ "short": "Inscribe polygon in greater circle",
+ "book": 12,
+ "number": 16,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop17",
+ "type": "proposition",
+ "label": "Given two spheres same center: inscribe in greater polyhedral solid not touching lesser",
+ "shortLabel": "Prop. XII.17",
+ "short": "Inscribe polyhedron in greater sphere",
+ "book": 12,
+ "number": 17,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop18",
+ "type": "proposition",
+ "label": "Spheres are to one another in triplicate ratio of their diameters",
+ "shortLabel": "Prop. XII.18",
+ "short": "Spheres: triplicate ratio",
+ "book": 12,
+ "number": 18,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "BookI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookV",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop18"
+ }
+ ],
+ "colorScheme": {
+ "foundation": {
+ "fill": "#95a5a6",
+ "stroke": "#7f8c8d"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/euclid-elements-book-xiii.json b/data/euclid-elements-book-xiii.json
new file mode 100644
index 0000000000000000000000000000000000000000..2edb8f726cdee55362f6d0b8356b51606fba7d5b
--- /dev/null
+++ b/data/euclid-elements-book-xiii.json
@@ -0,0 +1,650 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "euclid-elements-book-xiii",
+ "name": "Euclid's Elements, Book XIII",
+ "subject": "regular_solids",
+ "variant": "classical",
+ "description": "Regular solids: tetrahedron, octahedron, cube, icosahedron, dodecahedron. 18 propositions. Depends on Books I, IV, VI, X, XI. Source: David E. Joyce.",
+ "structure": {
+ "books": 13,
+ "propositions": 18,
+ "foundationTypes": [
+ "foundation"
+ ]
+ }
+ },
+ "metadata": {
+ "created": "2026-03-18",
+ "lastUpdated": "2026-03-18",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Euclid's Elements Book XIII Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Euclid",
+ "Elements",
+ "Book XIII",
+ "regular solids",
+ "Platonic",
+ "tetrahedron",
+ "octahedron",
+ "cube",
+ "icosahedron",
+ "dodecahedron"
+ ]
+ },
+ "sources": [
+ {
+ "id": "joyce",
+ "type": "digital",
+ "authors": "Joyce, David E.",
+ "title": "Euclid's Elements, Book XIII",
+ "year": "1996",
+ "url": "https://mathcs.clarku.edu/~djoyce/java/elements/bookXIII/bookXIII.html",
+ "notes": "Clark University"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "BookI",
+ "type": "foundation",
+ "label": "Book I — Plane geometry",
+ "shortLabel": "Book I",
+ "short": "Foundation",
+ "book": 1,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookIV",
+ "type": "foundation",
+ "label": "Book IV — Inscribed figures",
+ "shortLabel": "Book IV",
+ "short": "Foundation",
+ "book": 4,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookVI",
+ "type": "foundation",
+ "label": "Book VI — Similar figures",
+ "shortLabel": "Book VI",
+ "short": "Foundation",
+ "book": 6,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookX",
+ "type": "foundation",
+ "label": "Book X — Incommensurables",
+ "shortLabel": "Book X",
+ "short": "Foundation",
+ "book": 10,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "BookXI",
+ "type": "foundation",
+ "label": "Book XI — Solid geometry",
+ "shortLabel": "Book XI",
+ "short": "Foundation",
+ "book": 11,
+ "colorClass": "foundation"
+ },
+ {
+ "id": "Prop1",
+ "type": "proposition",
+ "label": "Line cut in extreme and mean ratio: square on greater plus half whole equals five times square on half",
+ "shortLabel": "Prop. XIII.1",
+ "short": "Extreme and mean: square on greater",
+ "book": 13,
+ "number": 1,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop2",
+ "type": "proposition",
+ "label": "If square on line five times square on segment: double segment cut in extreme and mean, greater is remainder",
+ "shortLabel": "Prop. XIII.2",
+ "short": "Square five times: extreme and mean",
+ "book": 13,
+ "number": 2,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop3",
+ "type": "proposition",
+ "label": "Line cut in extreme and mean: square on lesser + half greater equals five times square on half",
+ "shortLabel": "Prop. XIII.3",
+ "short": "Extreme and mean: sum of segments",
+ "book": 13,
+ "number": 3,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop4",
+ "type": "proposition",
+ "label": "Line cut in extreme and mean: sum of squares on whole and lesser triple square on greater",
+ "shortLabel": "Prop. XIII.4",
+ "short": "Extreme and mean: sum of squares",
+ "book": 13,
+ "number": 4,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop5",
+ "type": "proposition",
+ "label": "Line cut in extreme and mean, add greater: whole cut in extreme and mean, original is greater",
+ "shortLabel": "Prop. XIII.5",
+ "short": "Extreme and mean: add greater",
+ "book": 13,
+ "number": 5,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop6",
+ "type": "proposition",
+ "label": "Rational line cut in extreme and mean ratio: each segment is apotome",
+ "shortLabel": "Prop. XIII.6",
+ "short": "Rational cut: apotome",
+ "book": 13,
+ "number": 6,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop7",
+ "type": "proposition",
+ "label": "Equilateral pentagon: if three angles equal (order or not), pentagon equiangular",
+ "shortLabel": "Prop. XIII.7",
+ "short": "Equilateral pentagon: three angles",
+ "book": 13,
+ "number": 7,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop8",
+ "type": "proposition",
+ "label": "Equilateral equiangular pentagon: diagonals subtending two angles cut in extreme and mean ratio",
+ "shortLabel": "Prop. XIII.8",
+ "short": "Pentagon: diagonals in extreme and mean",
+ "book": 13,
+ "number": 8,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop9",
+ "type": "proposition",
+ "label": "Side of hexagon + decagon in same circle: cut in extreme and mean, greater is hexagon",
+ "shortLabel": "Prop. XIII.9",
+ "short": "Hexagon + decagon: extreme and mean",
+ "book": 13,
+ "number": 9,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop10",
+ "type": "proposition",
+ "label": "Equilateral pentagon in circle: square on side equals sum of squares on hexagon and decagon",
+ "shortLabel": "Prop. XIII.10",
+ "short": "Pentagon: square equals hexagon + decagon",
+ "book": 13,
+ "number": 10,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop11",
+ "type": "proposition",
+ "label": "Equilateral pentagon in circle with rational diameter: side is minor",
+ "shortLabel": "Prop. XIII.11",
+ "short": "Pentagon in rational circle: minor",
+ "book": 13,
+ "number": 11,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop12",
+ "type": "proposition",
+ "label": "Equilateral triangle in circle: square on side triple square on radius",
+ "shortLabel": "Prop. XIII.12",
+ "short": "Equilateral triangle: side triple radius",
+ "book": 13,
+ "number": 12,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop13",
+ "type": "proposition",
+ "label": "To construct pyramid (tetrahedron) in given sphere; diameter squared 1.5 times side squared",
+ "shortLabel": "Prop. XIII.13",
+ "short": "Construct tetrahedron in sphere",
+ "book": 13,
+ "number": 13,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop14",
+ "type": "proposition",
+ "label": "To construct octahedron in sphere; diameter squared double side squared",
+ "shortLabel": "Prop. XIII.14",
+ "short": "Construct octahedron in sphere",
+ "book": 13,
+ "number": 14,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop15",
+ "type": "proposition",
+ "label": "To construct cube in sphere; diameter squared triple side squared",
+ "shortLabel": "Prop. XIII.15",
+ "short": "Construct cube in sphere",
+ "book": 13,
+ "number": 15,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop16",
+ "type": "proposition",
+ "label": "To construct icosahedron in sphere; side is minor",
+ "shortLabel": "Prop. XIII.16",
+ "short": "Construct icosahedron in sphere",
+ "book": 13,
+ "number": 16,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop17",
+ "type": "proposition",
+ "label": "To construct dodecahedron in sphere; side is apotome",
+ "shortLabel": "Prop. XIII.17",
+ "short": "Construct dodecahedron in sphere",
+ "book": 13,
+ "number": 17,
+ "colorClass": "proposition"
+ },
+ {
+ "id": "Prop18",
+ "type": "proposition",
+ "label": "To set out sides of five figures and compare them; no other such figure exists",
+ "shortLabel": "Prop. XIII.18",
+ "short": "Compare five regular solids",
+ "book": 13,
+ "number": 18,
+ "colorClass": "proposition"
+ }
+ ],
+ "edges": [
+ {
+ "from": "BookI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop1"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop2"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop3"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop4"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop5"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop6"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop7"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop8"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop9"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop10"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop11"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop12"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop13"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop14"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop15"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop16"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop17"
+ },
+ {
+ "from": "BookI",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookIV",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookVI",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookX",
+ "to": "Prop18"
+ },
+ {
+ "from": "BookXI",
+ "to": "Prop18"
+ }
+ ],
+ "colorScheme": {
+ "foundation": {
+ "fill": "#95a5a6",
+ "stroke": "#7f8c8d"
+ },
+ "proposition": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/peano-arithmetic-addition-multiplication.mmd b/data/peano-arithmetic-addition-multiplication.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..c7b15fc92f1b439abafe644294b8ed8d41d02c27
--- /dev/null
+++ b/data/peano-arithmetic-addition-multiplication.mmd
@@ -0,0 +1,65 @@
+graph TD
+ A5["A5\nInduction"]
+ DefAdd["DefAdd\nDefinition of +"]
+ T5["T5\nAssociativity of +"]
+ T6["T6\nLeft identity"]
+ T7["T7\nSuccessor and add"]
+ T8["T8\nCommutativity of +"]
+ T9["T9\nCancellation for +"]
+ DefMul["DefMul\nDefinition of ·"]
+ T10["T10\nMul well-defined"]
+ T11["T11\nZero times"]
+ T12["T12\nZero from left"]
+ T13["T13\nSuccessor and mul"]
+ T14["T14\nCommutativity of ·"]
+ T15["T15\nAssociativity of ·"]
+ T16["T16\nDistributivity"]
+ T17["T17\nDistributivity (right)"]
+ A5 --> DefAdd
+ DefAdd --> T5
+ A5 --> T5
+ DefAdd --> T6
+ A5 --> T6
+ DefAdd --> T7
+ T6 --> T7
+ A5 --> T7
+ DefAdd --> T8
+ T5 --> T8
+ T6 --> T8
+ T7 --> T8
+ A5 --> T8
+ DefAdd --> T9
+ T8 --> T9
+ A5 --> T9
+ DefAdd --> DefMul
+ A5 --> DefMul
+ DefMul --> T10
+ A5 --> T10
+ DefMul --> T11
+ DefMul --> T12
+ T6 --> T12
+ A5 --> T12
+ DefMul --> T13
+ T8 --> T13
+ A5 --> T13
+ DefMul --> T14
+ T12 --> T14
+ T13 --> T14
+ A5 --> T14
+ DefMul --> T15
+ T5 --> T15
+ T8 --> T15
+ A5 --> T15
+ DefMul --> T16
+ T5 --> T16
+ T8 --> T16
+ T15 --> T16
+ A5 --> T16
+ T16 --> T17
+ T8 --> T17
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class A5 axiom
+ class DefAdd,DefMul definition
+ class T5,T6,T7,T8,T9,T10,T11,T12,T13,T14,T15,T16,T17 theorem
\ No newline at end of file
diff --git a/data/peano-arithmetic-axioms-foundations.mmd b/data/peano-arithmetic-axioms-foundations.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..a7c75d037fbfe82ae9fe1047012f88c611bf07f2
--- /dev/null
+++ b/data/peano-arithmetic-axioms-foundations.mmd
@@ -0,0 +1,33 @@
+graph TD
+ A1["A1\n0 ∈ N"]
+ A2["A2\n0 not a successor"]
+ A3["A3\nS injective"]
+ A4["A4\nN closed under S"]
+ A5["A5\nInduction"]
+ T1["T1\nContrapositive of A3"]
+ T2["T2\nSuccessor ≠ identity"]
+ T3["T3\nEvery nonzero is successor"]
+ DefAdd["DefAdd\nDefinition of +"]
+ T4["T4\nAdd well-defined"]
+ T5["T5\nAssociativity of +"]
+ T6["T6\nLeft identity"]
+ A3 --> T1
+ A1 --> T2
+ A2 --> T2
+ A3 --> T2
+ T1 --> T2
+ A5 --> T2
+ A5 --> T3
+ A5 --> DefAdd
+ DefAdd --> T4
+ A5 --> T4
+ DefAdd --> T5
+ A5 --> T5
+ DefAdd --> T6
+ A5 --> T6
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class A1,A2,A3,A4,A5 axiom
+ class DefAdd definition
+ class T1,T2,T3,T4,T5,T6 theorem
\ No newline at end of file
diff --git a/data/peano-arithmetic-order-induction.mmd b/data/peano-arithmetic-order-induction.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..cb69bea46270225b431c587eb2e7252813ae5ca4
--- /dev/null
+++ b/data/peano-arithmetic-order-induction.mmd
@@ -0,0 +1,86 @@
+graph TD
+ A5["A5\nInduction"]
+ DefAdd["DefAdd\nDefinition of +"]
+ T5["T5\nAssociativity of +"]
+ T6["T6\nLeft identity"]
+ T7["T7\nSuccessor and add"]
+ T8["T8\nCommutativity of +"]
+ T9["T9\nCancellation for +"]
+ DefMul["DefMul\nDefinition of ·"]
+ T12["T12\nZero from left"]
+ T13["T13\nSuccessor and mul"]
+ T14["T14\nCommutativity of ·"]
+ T15["T15\nAssociativity of ·"]
+ T16["T16\nDistributivity"]
+ T18["T18\nOrder definition"]
+ T19["T19\nTrichotomy"]
+ T20["T20\nOrder + add"]
+ T21["T21\nOrder + mul"]
+ T22["T22\nMultiplicative identity"]
+ T23["T23\nRight identity"]
+ T24["T24\nWell-ordering"]
+ T25["T25\nStrong induction"]
+ A5 --> DefAdd
+ DefAdd --> T5
+ A5 --> T5
+ DefAdd --> T6
+ A5 --> T6
+ DefAdd --> T7
+ T6 --> T7
+ A5 --> T7
+ DefAdd --> T8
+ T5 --> T8
+ T6 --> T8
+ T7 --> T8
+ A5 --> T8
+ DefAdd --> T9
+ T8 --> T9
+ A5 --> T9
+ DefAdd --> DefMul
+ A5 --> DefMul
+ DefMul --> T12
+ T6 --> T12
+ A5 --> T12
+ DefMul --> T13
+ T8 --> T13
+ A5 --> T13
+ DefMul --> T14
+ T12 --> T14
+ T13 --> T14
+ A5 --> T14
+ DefMul --> T15
+ T5 --> T15
+ T8 --> T15
+ A5 --> T15
+ DefMul --> T16
+ T5 --> T16
+ T8 --> T16
+ T15 --> T16
+ A5 --> T16
+ DefAdd --> T18
+ T8 --> T18
+ T9 --> T18
+ T18 --> T19
+ T9 --> T19
+ T18 --> T20
+ T8 --> T20
+ T18 --> T21
+ T16 --> T21
+ T14 --> T21
+ DefMul --> T22
+ T6 --> T22
+ A5 --> T22
+ T14 --> T23
+ T22 --> T23
+ T18 --> T24
+ T19 --> T24
+ A5 --> T24
+ T18 --> T25
+ T24 --> T25
+ A5 --> T25
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class A5 axiom
+ class DefAdd,DefMul definition
+ class T5,T6,T7,T8,T9,T12,T13,T14,T15,T16,T18,T19,T20,T21,T22,T23,T24,T25 theorem
\ No newline at end of file
diff --git a/data/peano-arithmetic.json b/data/peano-arithmetic.json
new file mode 100644
index 0000000000000000000000000000000000000000..47b74516536d3eecf7a0339d1842bb541217fbd3
--- /dev/null
+++ b/data/peano-arithmetic.json
@@ -0,0 +1,614 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "peano-arithmetic",
+ "name": "Peano Arithmetic",
+ "subject": "arithmetic",
+ "variant": "classical",
+ "description": "Axiomatic development of natural number arithmetic. Five axioms, definitions of addition and multiplication, and key theorems (associativity, commutativity, distributivity, order). Based on Landau, Foundations of Analysis.",
+ "structure": {
+ "axioms": 5,
+ "definitions": 2,
+ "theorems": 25
+ }
+ },
+ "metadata": {
+ "created": "2026-03-15",
+ "lastUpdated": "2026-03-15",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Peano Arithmetic Dependency Graph. Programming Framework.",
+ "keywords": [
+ "Peano",
+ "arithmetic",
+ "natural numbers",
+ "induction",
+ "foundations"
+ ]
+ },
+ "sources": [
+ {
+ "id": "landau",
+ "type": "primary",
+ "authors": "Landau, E.",
+ "title": "Foundations of Analysis",
+ "year": "1930",
+ "publisher": "Chelsea",
+ "edition": "1951",
+ "notes": "Canonical development"
+ },
+ {
+ "id": "wikipedia",
+ "type": "digital",
+ "title": "Peano axioms",
+ "url": "https://en.wikipedia.org/wiki/Peano_axioms",
+ "notes": "Overview and definitions"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "A1",
+ "type": "axiom",
+ "label": "0 is a natural number",
+ "shortLabel": "A1",
+ "short": "0 ∈ N",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "A2",
+ "type": "axiom",
+ "label": "No predecessor of 0: S(x) ≠ 0",
+ "shortLabel": "A2",
+ "short": "0 not a successor",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "A3",
+ "type": "axiom",
+ "label": "Successor injective: S(x)=S(y) ⇒ x=y",
+ "shortLabel": "A3",
+ "short": "S injective",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "A4",
+ "type": "axiom",
+ "label": "Closure: S(x) ∈ N for all x ∈ N",
+ "shortLabel": "A4",
+ "short": "N closed under S",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "A5",
+ "type": "axiom",
+ "label": "Induction: if 0∈K and (x∈K⇒S(x)∈K) then K=N",
+ "shortLabel": "A5",
+ "short": "Induction",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "T1",
+ "type": "theorem",
+ "label": "x≠y ⇒ S(x)≠S(y)",
+ "shortLabel": "T1",
+ "short": "Contrapositive of A3",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T2",
+ "type": "theorem",
+ "label": "S(x)≠x for all x",
+ "shortLabel": "T2",
+ "short": "Successor ≠ identity",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T3",
+ "type": "theorem",
+ "label": "If x≠0 then x=S(u) for some u",
+ "shortLabel": "T3",
+ "short": "Every nonzero is successor",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "DefAdd",
+ "type": "definition",
+ "label": "Addition: x+0=x, x+S(y)=S(x+y)",
+ "shortLabel": "DefAdd",
+ "short": "Definition of +",
+ "colorClass": "definition"
+ },
+ {
+ "id": "T4",
+ "type": "theorem",
+ "label": "Addition is well-defined for all x,y",
+ "shortLabel": "T4",
+ "short": "Add well-defined",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T5",
+ "type": "theorem",
+ "label": "(x+y)+z = x+(y+z)",
+ "shortLabel": "T5",
+ "short": "Associativity of +",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T6",
+ "type": "theorem",
+ "label": "0+x = x",
+ "shortLabel": "T6",
+ "short": "Left identity",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T7",
+ "type": "theorem",
+ "label": "S(x)+y = S(x+y)",
+ "shortLabel": "T7",
+ "short": "Successor and add",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T8",
+ "type": "theorem",
+ "label": "x+y = y+x",
+ "shortLabel": "T8",
+ "short": "Commutativity of +",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T9",
+ "type": "theorem",
+ "label": "x+y=x+z ⇒ y=z",
+ "shortLabel": "T9",
+ "short": "Cancellation for +",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "DefMul",
+ "type": "definition",
+ "label": "Multiplication: x·0=0, x·S(y)=x·y+x",
+ "shortLabel": "DefMul",
+ "short": "Definition of ·",
+ "colorClass": "definition"
+ },
+ {
+ "id": "T10",
+ "type": "theorem",
+ "label": "Multiplication is well-defined for all x,y",
+ "shortLabel": "T10",
+ "short": "Mul well-defined",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T11",
+ "type": "theorem",
+ "label": "x·0 = 0",
+ "shortLabel": "T11",
+ "short": "Zero times",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T12",
+ "type": "theorem",
+ "label": "0·x = 0",
+ "shortLabel": "T12",
+ "short": "Zero from left",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T13",
+ "type": "theorem",
+ "label": "S(x)·y = x·y + y",
+ "shortLabel": "T13",
+ "short": "Successor and mul",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T14",
+ "type": "theorem",
+ "label": "x·y = y·x",
+ "shortLabel": "T14",
+ "short": "Commutativity of ·",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T15",
+ "type": "theorem",
+ "label": "(x·y)·z = x·(y·z)",
+ "shortLabel": "T15",
+ "short": "Associativity of ·",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T16",
+ "type": "theorem",
+ "label": "x·(y+z) = x·y + x·z",
+ "shortLabel": "T16",
+ "short": "Distributivity",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T17",
+ "type": "theorem",
+ "label": "(x+y)·z = x·z + y·z",
+ "shortLabel": "T17",
+ "short": "Distributivity (right)",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T18",
+ "type": "theorem",
+ "label": "x≤y iff ∃z x+z=y",
+ "shortLabel": "T18",
+ "short": "Order definition",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T19",
+ "type": "theorem",
+ "label": "Trichotomy: exactly one of x
0 ⇒ x·z≤y·z",
+ "shortLabel": "T21",
+ "short": "Order + mul",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T22",
+ "type": "theorem",
+ "label": "1·x = x (where 1=S(0))",
+ "shortLabel": "T22",
+ "short": "Multiplicative identity",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T23",
+ "type": "theorem",
+ "label": "x·1 = x",
+ "shortLabel": "T23",
+ "short": "Right identity",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T24",
+ "type": "theorem",
+ "label": "Well-ordering: every nonempty subset has least element",
+ "shortLabel": "T24",
+ "short": "Well-ordering",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T25",
+ "type": "theorem",
+ "label": "Strong induction principle",
+ "shortLabel": "T25",
+ "short": "Strong induction",
+ "colorClass": "theorem"
+ }
+ ],
+ "edges": [
+ {
+ "from": "A3",
+ "to": "T1"
+ },
+ {
+ "from": "A1",
+ "to": "T2"
+ },
+ {
+ "from": "A2",
+ "to": "T2"
+ },
+ {
+ "from": "A3",
+ "to": "T2"
+ },
+ {
+ "from": "T1",
+ "to": "T2"
+ },
+ {
+ "from": "A5",
+ "to": "T2"
+ },
+ {
+ "from": "A5",
+ "to": "T3"
+ },
+ {
+ "from": "A5",
+ "to": "DefAdd"
+ },
+ {
+ "from": "DefAdd",
+ "to": "T4"
+ },
+ {
+ "from": "A5",
+ "to": "T4"
+ },
+ {
+ "from": "DefAdd",
+ "to": "T5"
+ },
+ {
+ "from": "A5",
+ "to": "T5"
+ },
+ {
+ "from": "DefAdd",
+ "to": "T6"
+ },
+ {
+ "from": "A5",
+ "to": "T6"
+ },
+ {
+ "from": "DefAdd",
+ "to": "T7"
+ },
+ {
+ "from": "T6",
+ "to": "T7"
+ },
+ {
+ "from": "A5",
+ "to": "T7"
+ },
+ {
+ "from": "DefAdd",
+ "to": "T8"
+ },
+ {
+ "from": "T5",
+ "to": "T8"
+ },
+ {
+ "from": "T6",
+ "to": "T8"
+ },
+ {
+ "from": "T7",
+ "to": "T8"
+ },
+ {
+ "from": "A5",
+ "to": "T8"
+ },
+ {
+ "from": "DefAdd",
+ "to": "T9"
+ },
+ {
+ "from": "T8",
+ "to": "T9"
+ },
+ {
+ "from": "A5",
+ "to": "T9"
+ },
+ {
+ "from": "DefAdd",
+ "to": "DefMul"
+ },
+ {
+ "from": "A5",
+ "to": "DefMul"
+ },
+ {
+ "from": "DefMul",
+ "to": "T10"
+ },
+ {
+ "from": "A5",
+ "to": "T10"
+ },
+ {
+ "from": "DefMul",
+ "to": "T11"
+ },
+ {
+ "from": "DefMul",
+ "to": "T12"
+ },
+ {
+ "from": "T6",
+ "to": "T12"
+ },
+ {
+ "from": "A5",
+ "to": "T12"
+ },
+ {
+ "from": "DefMul",
+ "to": "T13"
+ },
+ {
+ "from": "T8",
+ "to": "T13"
+ },
+ {
+ "from": "A5",
+ "to": "T13"
+ },
+ {
+ "from": "DefMul",
+ "to": "T14"
+ },
+ {
+ "from": "T12",
+ "to": "T14"
+ },
+ {
+ "from": "T13",
+ "to": "T14"
+ },
+ {
+ "from": "A5",
+ "to": "T14"
+ },
+ {
+ "from": "DefMul",
+ "to": "T15"
+ },
+ {
+ "from": "T5",
+ "to": "T15"
+ },
+ {
+ "from": "T8",
+ "to": "T15"
+ },
+ {
+ "from": "A5",
+ "to": "T15"
+ },
+ {
+ "from": "DefMul",
+ "to": "T16"
+ },
+ {
+ "from": "T5",
+ "to": "T16"
+ },
+ {
+ "from": "T8",
+ "to": "T16"
+ },
+ {
+ "from": "T15",
+ "to": "T16"
+ },
+ {
+ "from": "A5",
+ "to": "T16"
+ },
+ {
+ "from": "T16",
+ "to": "T17"
+ },
+ {
+ "from": "T8",
+ "to": "T17"
+ },
+ {
+ "from": "DefAdd",
+ "to": "T18"
+ },
+ {
+ "from": "T8",
+ "to": "T18"
+ },
+ {
+ "from": "T9",
+ "to": "T18"
+ },
+ {
+ "from": "T18",
+ "to": "T19"
+ },
+ {
+ "from": "T9",
+ "to": "T19"
+ },
+ {
+ "from": "T18",
+ "to": "T20"
+ },
+ {
+ "from": "T8",
+ "to": "T20"
+ },
+ {
+ "from": "T18",
+ "to": "T21"
+ },
+ {
+ "from": "T16",
+ "to": "T21"
+ },
+ {
+ "from": "T14",
+ "to": "T21"
+ },
+ {
+ "from": "DefMul",
+ "to": "T22"
+ },
+ {
+ "from": "T6",
+ "to": "T22"
+ },
+ {
+ "from": "A5",
+ "to": "T22"
+ },
+ {
+ "from": "T14",
+ "to": "T23"
+ },
+ {
+ "from": "T22",
+ "to": "T23"
+ },
+ {
+ "from": "T18",
+ "to": "T24"
+ },
+ {
+ "from": "T19",
+ "to": "T24"
+ },
+ {
+ "from": "A5",
+ "to": "T24"
+ },
+ {
+ "from": "T18",
+ "to": "T25"
+ },
+ {
+ "from": "T24",
+ "to": "T25"
+ },
+ {
+ "from": "A5",
+ "to": "T25"
+ }
+ ],
+ "colorScheme": {
+ "axiom": {
+ "fill": "#e74c3c",
+ "stroke": "#c0392b"
+ },
+ "definition": {
+ "fill": "#3498db",
+ "stroke": "#2980b9"
+ },
+ "theorem": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/peano-arithmetic.mmd b/data/peano-arithmetic.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..89105eff1d6888d1ff9f85c5c220434dbf6ba401
--- /dev/null
+++ b/data/peano-arithmetic.mmd
@@ -0,0 +1,111 @@
+graph TD
+ A1["A1\n0 ∈ N"]
+ A2["A2\n0 not a successor"]
+ A3["A3\nS injective"]
+ A4["A4\nN closed under S"]
+ A5["A5\nInduction"]
+ T1["T1\nContrapositive of A3"]
+ T2["T2\nSuccessor ≠ identity"]
+ T3["T3\nEvery nonzero is successor"]
+ DefAdd["DefAdd\nDefinition of +"]
+ T4["T4\nAdd well-defined"]
+ T5["T5\nAssociativity of +"]
+ T6["T6\nLeft identity"]
+ T7["T7\nSuccessor and add"]
+ T8["T8\nCommutativity of +"]
+ T9["T9\nCancellation for +"]
+ DefMul["DefMul\nDefinition of ·"]
+ T10["T10\nMul well-defined"]
+ T11["T11\nZero times"]
+ T12["T12\nZero from left"]
+ T13["T13\nSuccessor and mul"]
+ T14["T14\nCommutativity of ·"]
+ T15["T15\nAssociativity of ·"]
+ T16["T16\nDistributivity"]
+ T17["T17\nDistributivity (right)"]
+ T18["T18\nOrder definition"]
+ T19["T19\nTrichotomy"]
+ T20["T20\nOrder + add"]
+ T21["T21\nOrder + mul"]
+ T22["T22\nMultiplicative identity"]
+ T23["T23\nRight identity"]
+ T24["T24\nWell-ordering"]
+ T25["T25\nStrong induction"]
+ A3 --> T1
+ A1 --> T2
+ A2 --> T2
+ A3 --> T2
+ T1 --> T2
+ A5 --> T2
+ A5 --> T3
+ A5 --> DefAdd
+ DefAdd --> T4
+ A5 --> T4
+ DefAdd --> T5
+ A5 --> T5
+ DefAdd --> T6
+ A5 --> T6
+ DefAdd --> T7
+ T6 --> T7
+ A5 --> T7
+ DefAdd --> T8
+ T5 --> T8
+ T6 --> T8
+ T7 --> T8
+ A5 --> T8
+ DefAdd --> T9
+ T8 --> T9
+ A5 --> T9
+ DefAdd --> DefMul
+ A5 --> DefMul
+ DefMul --> T10
+ A5 --> T10
+ DefMul --> T11
+ DefMul --> T12
+ T6 --> T12
+ A5 --> T12
+ DefMul --> T13
+ T8 --> T13
+ A5 --> T13
+ DefMul --> T14
+ T12 --> T14
+ T13 --> T14
+ A5 --> T14
+ DefMul --> T15
+ T5 --> T15
+ T8 --> T15
+ A5 --> T15
+ DefMul --> T16
+ T5 --> T16
+ T8 --> T16
+ T15 --> T16
+ A5 --> T16
+ T16 --> T17
+ T8 --> T17
+ DefAdd --> T18
+ T8 --> T18
+ T9 --> T18
+ T18 --> T19
+ T9 --> T19
+ T18 --> T20
+ T8 --> T20
+ T18 --> T21
+ T16 --> T21
+ T14 --> T21
+ DefMul --> T22
+ T6 --> T22
+ A5 --> T22
+ T14 --> T23
+ T22 --> T23
+ T18 --> T24
+ T19 --> T24
+ A5 --> T24
+ T18 --> T25
+ T24 --> T25
+ A5 --> T25
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class A1,A2,A3,A4,A5 axiom
+ class DefAdd,DefMul definition
+ class T1,T2,T3,T4,T5,T6,T7,T8,T9,T10,T11,T12,T13,T14,T15,T16,T17,T18,T19,T20,T21,T22,T23,T24,T25 theorem
\ No newline at end of file
diff --git a/data/propositional-logic-axioms-implication.mmd b/data/propositional-logic-axioms-implication.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..3bdf2d938eefabc8edfb07a47df7659c130f4034
--- /dev/null
+++ b/data/propositional-logic-axioms-implication.mmd
@@ -0,0 +1,31 @@
+graph TD
+ A1("A1 Weakening")
+ A2("A2 Distrib. of impl.")
+ A3("A3 Contraposition")
+ MP("MP MP")
+ T1("T1 Self-implication")
+ T2("T2 Double neg. elim")
+ T3("T3 Double neg. intro")
+ T4("T4 Transposition")
+ T5("T5 Hyp. syllogism")
+ A1 --> T1
+ A2 --> T1
+ MP --> T1
+ A3 --> T2
+ T1 --> T2
+ MP --> T2
+ A1 --> T3
+ A3 --> T3
+ MP --> T3
+ A3 --> T4
+ T2 --> T4
+ T3 --> T4
+ MP --> T4
+ A2 --> T5
+ T1 --> T5
+ MP --> T5
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class A1,A2,A3,MP axiom
+ class T1,T2,T3,T4,T5 theorem
\ No newline at end of file
diff --git a/data/propositional-logic-definitions-connectives.mmd b/data/propositional-logic-definitions-connectives.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..f05e7115c76575c70e47523c1467294eb1ef204f
--- /dev/null
+++ b/data/propositional-logic-definitions-connectives.mmd
@@ -0,0 +1,74 @@
+graph TD
+ A1("A1 Weakening")
+ A2("A2 Distrib. of impl.")
+ A3("A3 Contraposition")
+ MP("MP MP")
+ T1("T1 Self-implication")
+ T2("T2 Double neg. elim")
+ T3("T3 Double neg. intro")
+ T4("T4 Transposition")
+ DefOr("DefOr Def. disjunction")
+ DefAnd("DefAnd Def. conjunction")
+ DefIff("DefIff Def. biconditional")
+ T6("T6 Addition (orI)")
+ T7("T7 Simplification (andE)")
+ T8("T8 Simplification (andE)")
+ T9("T9 Conjunction (andI)")
+ T10("T10 Material impl.")
+ T11("T11 De Morgan (1)")
+ T12("T12 De Morgan (2)")
+ A1 --> T1
+ A2 --> T1
+ MP --> T1
+ A3 --> T2
+ T1 --> T2
+ MP --> T2
+ A1 --> T3
+ A3 --> T3
+ MP --> T3
+ A3 --> T4
+ T2 --> T4
+ T3 --> T4
+ MP --> T4
+ A1 --> DefOr
+ A2 --> DefOr
+ A3 --> DefOr
+ MP --> DefOr
+ DefOr --> DefAnd
+ A3 --> DefAnd
+ MP --> DefAnd
+ DefAnd --> DefIff
+ T9 --> DefIff
+ MP --> DefIff
+ DefOr --> T6
+ A1 --> T6
+ MP --> T6
+ DefAnd --> T7
+ A1 --> T7
+ A3 --> T7
+ MP --> T7
+ DefAnd --> T8
+ A1 --> T8
+ A3 --> T8
+ MP --> T8
+ A1 --> T9
+ A2 --> T9
+ MP --> T9
+ DefOr --> T10
+ T4 --> T10
+ MP --> T10
+ DefAnd --> T11
+ DefOr --> T11
+ T4 --> T11
+ T6 --> T11
+ MP --> T11
+ DefAnd --> T12
+ DefOr --> T12
+ T4 --> T12
+ MP --> T12
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class A1,A2,A3,MP axiom
+ class DefOr,DefAnd,DefIff definition
+ class T1,T2,T3,T4,T6,T7,T8,T9,T10,T11,T12 theorem
\ No newline at end of file
diff --git a/data/propositional-logic-tautologies-metalogic.mmd b/data/propositional-logic-tautologies-metalogic.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..8149a976d6d0efe07491f230d42406370794dffd
--- /dev/null
+++ b/data/propositional-logic-tautologies-metalogic.mmd
@@ -0,0 +1,89 @@
+graph TD
+ A1("A1 Weakening")
+ A2("A2 Distrib. of impl.")
+ A3("A3 Contraposition")
+ MP("MP MP")
+ T1("T1 Self-implication")
+ T2("T2 Double neg. elim")
+ T3("T3 Double neg. intro")
+ T4("T4 Transposition")
+ DefOr("DefOr Def. disjunction")
+ DefAnd("DefAnd Def. conjunction")
+ T6("T6 Addition (orI)")
+ T7("T7 Simplification (andE)")
+ T8("T8 Simplification (andE)")
+ T9("T9 Conjunction (andI)")
+ T13("T13 Excluded middle")
+ T14("T14 Non-contradiction")
+ T15("T15 Explosion")
+ T16("T16 Commutation (or)")
+ T17("T17 Commutation (and)")
+ T18("T18 Distribution")
+ T19("T19 Deduction thm")
+ A1 --> T1
+ A2 --> T1
+ MP --> T1
+ A3 --> T2
+ T1 --> T2
+ MP --> T2
+ A1 --> T3
+ A3 --> T3
+ MP --> T3
+ A3 --> T4
+ T2 --> T4
+ T3 --> T4
+ MP --> T4
+ A1 --> DefOr
+ A2 --> DefOr
+ A3 --> DefOr
+ MP --> DefOr
+ DefOr --> DefAnd
+ A3 --> DefAnd
+ MP --> DefAnd
+ DefOr --> T6
+ A1 --> T6
+ MP --> T6
+ DefAnd --> T7
+ A1 --> T7
+ A3 --> T7
+ MP --> T7
+ DefAnd --> T8
+ A1 --> T8
+ A3 --> T8
+ MP --> T8
+ A1 --> T9
+ A2 --> T9
+ MP --> T9
+ DefOr --> T13
+ T2 --> T13
+ T3 --> T13
+ MP --> T13
+ DefAnd --> T14
+ T4 --> T14
+ MP --> T14
+ DefAnd --> T15
+ A1 --> T15
+ A2 --> T15
+ MP --> T15
+ DefOr --> T16
+ T4 --> T16
+ MP --> T16
+ DefAnd --> T17
+ T9 --> T17
+ MP --> T17
+ DefAnd --> T18
+ DefOr --> T18
+ T6 --> T18
+ T7 --> T18
+ T8 --> T18
+ T9 --> T18
+ MP --> T18
+ A1 --> T19
+ A2 --> T19
+ MP --> T19
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class A1,A2,A3,MP axiom
+ class DefOr,DefAnd definition
+ class T1,T2,T3,T4,T6,T7,T8,T9,T13,T14,T15,T16,T17,T18,T19 theorem
\ No newline at end of file
diff --git a/data/propositional-logic.json b/data/propositional-logic.json
new file mode 100644
index 0000000000000000000000000000000000000000..7d25b5083410a250ae81624903c41ab9c96d1f00
--- /dev/null
+++ b/data/propositional-logic.json
@@ -0,0 +1,600 @@
+{
+ "schemaVersion": "1.0",
+ "discourse": {
+ "id": "propositional-logic",
+ "name": "Propositional Logic",
+ "subject": "logic",
+ "variant": "classical",
+ "description": "Hilbert-style axiomatic development of classical propositional logic. Three axioms (Łukasiewicz P2), modus ponens, definitions of disjunction, conjunction, biconditional, and key theorems (double negation, De Morgan, excluded middle, deduction theorem).",
+ "structure": {
+ "axioms": 4,
+ "definitions": 3,
+ "theorems": 19
+ }
+ },
+ "metadata": {
+ "created": "2026-03-15",
+ "lastUpdated": "2026-03-15",
+ "version": "1.0.0",
+ "license": "CC BY 4.0",
+ "authors": [
+ "Welz, G."
+ ],
+ "methodology": "Programming Framework",
+ "citation": "Welz, G. (2026). Propositional Logic Dependency Graph. Programming Framework.",
+ "keywords": [
+ "propositional logic",
+ "Hilbert",
+ "Łukasiewicz",
+ "tautology",
+ "modus ponens"
+ ]
+ },
+ "sources": [
+ {
+ "id": "frege",
+ "type": "primary",
+ "authors": "Frege, G.",
+ "title": "Begriffsschrift",
+ "year": "1879",
+ "notes": "First axiomatic propositional logic"
+ },
+ {
+ "id": "lukasiewicz",
+ "type": "primary",
+ "authors": "Łukasiewicz, J.",
+ "title": "Elements of Mathematical Logic",
+ "year": "1929",
+ "notes": "P2: 3 axioms"
+ },
+ {
+ "id": "wikipedia",
+ "type": "digital",
+ "title": "Propositional calculus",
+ "url": "https://en.wikipedia.org/wiki/Propositional_calculus",
+ "notes": "Axioms and theorems"
+ }
+ ],
+ "nodes": [
+ {
+ "id": "A1",
+ "type": "axiom",
+ "label": "φ → (ψ → φ)",
+ "shortLabel": "A1",
+ "short": "Weakening",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "A2",
+ "type": "axiom",
+ "label": "(φ→(ψ→χ)) → ((φ→ψ)→(φ→χ))",
+ "shortLabel": "A2",
+ "short": "Distrib. of impl.",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "A3",
+ "type": "axiom",
+ "label": "(¬φ→¬ψ) → (ψ→φ)",
+ "shortLabel": "A3",
+ "short": "Contraposition",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "MP",
+ "type": "axiom",
+ "label": "Modus Ponens: φ, (φ→ψ) ⊢ ψ",
+ "shortLabel": "MP",
+ "short": "MP",
+ "colorClass": "axiom"
+ },
+ {
+ "id": "T1",
+ "type": "theorem",
+ "label": "φ → φ",
+ "shortLabel": "T1",
+ "short": "Self-implication",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T2",
+ "type": "theorem",
+ "label": "¬¬φ → φ",
+ "shortLabel": "T2",
+ "short": "Double neg. elim",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T3",
+ "type": "theorem",
+ "label": "φ → ¬¬φ",
+ "shortLabel": "T3",
+ "short": "Double neg. intro",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T4",
+ "type": "theorem",
+ "label": "(φ→ψ) → (¬ψ→¬φ)",
+ "shortLabel": "T4",
+ "short": "Transposition",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T5",
+ "type": "theorem",
+ "label": "(φ→ψ)∧(ψ→χ) ⇒ (φ→χ)",
+ "shortLabel": "T5",
+ "short": "Hyp. syllogism",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "DefOr",
+ "type": "definition",
+ "label": "φ ∨ ψ := ¬φ → ψ",
+ "shortLabel": "DefOr",
+ "short": "Def. disjunction",
+ "colorClass": "definition"
+ },
+ {
+ "id": "DefAnd",
+ "type": "definition",
+ "label": "φ ∧ ψ := ¬(φ → ¬ψ)",
+ "shortLabel": "DefAnd",
+ "short": "Def. conjunction",
+ "colorClass": "definition"
+ },
+ {
+ "id": "DefIff",
+ "type": "definition",
+ "label": "φ ↔ ψ := (φ→ψ)∧(ψ→φ)",
+ "shortLabel": "DefIff",
+ "short": "Def. biconditional",
+ "colorClass": "definition"
+ },
+ {
+ "id": "T6",
+ "type": "theorem",
+ "label": "φ → (φ ∨ ψ)",
+ "shortLabel": "T6",
+ "short": "Addition (∨I)",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T7",
+ "type": "theorem",
+ "label": "(φ∧ψ) → φ",
+ "shortLabel": "T7",
+ "short": "Simplification (∧E)",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T8",
+ "type": "theorem",
+ "label": "(φ∧ψ) → ψ",
+ "shortLabel": "T8",
+ "short": "Simplification (∧E)",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T9",
+ "type": "theorem",
+ "label": "φ → (ψ → (φ∧ψ))",
+ "shortLabel": "T9",
+ "short": "Conjunction (∧I)",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T10",
+ "type": "theorem",
+ "label": "(φ→ψ) ↔ (¬φ∨ψ)",
+ "shortLabel": "T10",
+ "short": "Material impl.",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T11",
+ "type": "theorem",
+ "label": "¬(φ∧ψ) ↔ (¬φ∨¬ψ)",
+ "shortLabel": "T11",
+ "short": "De Morgan (1)",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T12",
+ "type": "theorem",
+ "label": "¬(φ∨ψ) ↔ (¬φ∧¬ψ)",
+ "shortLabel": "T12",
+ "short": "De Morgan (2)",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T13",
+ "type": "theorem",
+ "label": "φ ∨ ¬φ",
+ "shortLabel": "T13",
+ "short": "Excluded middle",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T14",
+ "type": "theorem",
+ "label": "¬(φ ∧ ¬φ)",
+ "shortLabel": "T14",
+ "short": "Non-contradiction",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T15",
+ "type": "theorem",
+ "label": "(φ∧¬φ) → ψ",
+ "shortLabel": "T15",
+ "short": "Explosion",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T16",
+ "type": "theorem",
+ "label": "(φ∨ψ) ↔ (ψ∨φ)",
+ "shortLabel": "T16",
+ "short": "Commutation (∨)",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T17",
+ "type": "theorem",
+ "label": "(φ∧ψ) ↔ (ψ∧φ)",
+ "shortLabel": "T17",
+ "short": "Commutation (∧)",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T18",
+ "type": "theorem",
+ "label": "(φ∧(ψ∨χ)) ↔ ((φ∧ψ)∨(φ∧χ))",
+ "shortLabel": "T18",
+ "short": "Distribution",
+ "colorClass": "theorem"
+ },
+ {
+ "id": "T19",
+ "type": "theorem",
+ "label": "Deduction theorem",
+ "shortLabel": "T19",
+ "short": "Deduction thm",
+ "colorClass": "theorem"
+ }
+ ],
+ "edges": [
+ {
+ "from": "A1",
+ "to": "T1"
+ },
+ {
+ "from": "A2",
+ "to": "T1"
+ },
+ {
+ "from": "MP",
+ "to": "T1"
+ },
+ {
+ "from": "A3",
+ "to": "T2"
+ },
+ {
+ "from": "T1",
+ "to": "T2"
+ },
+ {
+ "from": "MP",
+ "to": "T2"
+ },
+ {
+ "from": "A1",
+ "to": "T3"
+ },
+ {
+ "from": "A3",
+ "to": "T3"
+ },
+ {
+ "from": "MP",
+ "to": "T3"
+ },
+ {
+ "from": "A3",
+ "to": "T4"
+ },
+ {
+ "from": "T2",
+ "to": "T4"
+ },
+ {
+ "from": "T3",
+ "to": "T4"
+ },
+ {
+ "from": "MP",
+ "to": "T4"
+ },
+ {
+ "from": "A2",
+ "to": "T5"
+ },
+ {
+ "from": "T1",
+ "to": "T5"
+ },
+ {
+ "from": "MP",
+ "to": "T5"
+ },
+ {
+ "from": "A1",
+ "to": "DefOr"
+ },
+ {
+ "from": "A2",
+ "to": "DefOr"
+ },
+ {
+ "from": "A3",
+ "to": "DefOr"
+ },
+ {
+ "from": "MP",
+ "to": "DefOr"
+ },
+ {
+ "from": "DefOr",
+ "to": "DefAnd"
+ },
+ {
+ "from": "A3",
+ "to": "DefAnd"
+ },
+ {
+ "from": "MP",
+ "to": "DefAnd"
+ },
+ {
+ "from": "DefAnd",
+ "to": "DefIff"
+ },
+ {
+ "from": "T9",
+ "to": "DefIff"
+ },
+ {
+ "from": "MP",
+ "to": "DefIff"
+ },
+ {
+ "from": "DefOr",
+ "to": "T6"
+ },
+ {
+ "from": "A1",
+ "to": "T6"
+ },
+ {
+ "from": "MP",
+ "to": "T6"
+ },
+ {
+ "from": "DefAnd",
+ "to": "T7"
+ },
+ {
+ "from": "A1",
+ "to": "T7"
+ },
+ {
+ "from": "A3",
+ "to": "T7"
+ },
+ {
+ "from": "MP",
+ "to": "T7"
+ },
+ {
+ "from": "DefAnd",
+ "to": "T8"
+ },
+ {
+ "from": "A1",
+ "to": "T8"
+ },
+ {
+ "from": "A3",
+ "to": "T8"
+ },
+ {
+ "from": "MP",
+ "to": "T8"
+ },
+ {
+ "from": "A1",
+ "to": "T9"
+ },
+ {
+ "from": "A2",
+ "to": "T9"
+ },
+ {
+ "from": "MP",
+ "to": "T9"
+ },
+ {
+ "from": "DefOr",
+ "to": "T10"
+ },
+ {
+ "from": "T4",
+ "to": "T10"
+ },
+ {
+ "from": "MP",
+ "to": "T10"
+ },
+ {
+ "from": "DefAnd",
+ "to": "T11"
+ },
+ {
+ "from": "DefOr",
+ "to": "T11"
+ },
+ {
+ "from": "T4",
+ "to": "T11"
+ },
+ {
+ "from": "T6",
+ "to": "T11"
+ },
+ {
+ "from": "MP",
+ "to": "T11"
+ },
+ {
+ "from": "DefAnd",
+ "to": "T12"
+ },
+ {
+ "from": "DefOr",
+ "to": "T12"
+ },
+ {
+ "from": "T4",
+ "to": "T12"
+ },
+ {
+ "from": "MP",
+ "to": "T12"
+ },
+ {
+ "from": "DefOr",
+ "to": "T13"
+ },
+ {
+ "from": "T2",
+ "to": "T13"
+ },
+ {
+ "from": "T3",
+ "to": "T13"
+ },
+ {
+ "from": "MP",
+ "to": "T13"
+ },
+ {
+ "from": "DefAnd",
+ "to": "T14"
+ },
+ {
+ "from": "T4",
+ "to": "T14"
+ },
+ {
+ "from": "MP",
+ "to": "T14"
+ },
+ {
+ "from": "DefAnd",
+ "to": "T15"
+ },
+ {
+ "from": "A1",
+ "to": "T15"
+ },
+ {
+ "from": "A2",
+ "to": "T15"
+ },
+ {
+ "from": "MP",
+ "to": "T15"
+ },
+ {
+ "from": "DefOr",
+ "to": "T16"
+ },
+ {
+ "from": "T4",
+ "to": "T16"
+ },
+ {
+ "from": "MP",
+ "to": "T16"
+ },
+ {
+ "from": "DefAnd",
+ "to": "T17"
+ },
+ {
+ "from": "T9",
+ "to": "T17"
+ },
+ {
+ "from": "MP",
+ "to": "T17"
+ },
+ {
+ "from": "DefAnd",
+ "to": "T18"
+ },
+ {
+ "from": "DefOr",
+ "to": "T18"
+ },
+ {
+ "from": "T6",
+ "to": "T18"
+ },
+ {
+ "from": "T7",
+ "to": "T18"
+ },
+ {
+ "from": "T8",
+ "to": "T18"
+ },
+ {
+ "from": "T9",
+ "to": "T18"
+ },
+ {
+ "from": "MP",
+ "to": "T18"
+ },
+ {
+ "from": "A1",
+ "to": "T19"
+ },
+ {
+ "from": "A2",
+ "to": "T19"
+ },
+ {
+ "from": "MP",
+ "to": "T19"
+ }
+ ],
+ "colorScheme": {
+ "axiom": {
+ "fill": "#e74c3c",
+ "stroke": "#c0392b"
+ },
+ "definition": {
+ "fill": "#3498db",
+ "stroke": "#2980b9"
+ },
+ "theorem": {
+ "fill": "#1abc9c",
+ "stroke": "#16a085"
+ }
+ }
+}
\ No newline at end of file
diff --git a/data/propositional-logic.mmd b/data/propositional-logic.mmd
new file mode 100644
index 0000000000000000000000000000000000000000..32bbaeb2c109c160e2d30e5d200304391054676b
--- /dev/null
+++ b/data/propositional-logic.mmd
@@ -0,0 +1,112 @@
+graph TD
+ A1("A1 Weakening")
+ A2("A2 Distrib. of impl.")
+ A3("A3 Contraposition")
+ MP("MP MP")
+ T1("T1 Self-implication")
+ T2("T2 Double neg. elim")
+ T3("T3 Double neg. intro")
+ T4("T4 Transposition")
+ T5("T5 Hyp. syllogism")
+ DefOr("DefOr Def. disjunction")
+ DefAnd("DefAnd Def. conjunction")
+ DefIff("DefIff Def. biconditional")
+ T6("T6 Addition (orI)")
+ T7("T7 Simplification (andE)")
+ T8("T8 Simplification (andE)")
+ T9("T9 Conjunction (andI)")
+ T10("T10 Material impl.")
+ T11("T11 De Morgan (1)")
+ T12("T12 De Morgan (2)")
+ T13("T13 Excluded middle")
+ T14("T14 Non-contradiction")
+ T15("T15 Explosion")
+ T16("T16 Commutation (or)")
+ T17("T17 Commutation (and)")
+ T18("T18 Distribution")
+ T19("T19 Deduction thm")
+ A1 --> T1
+ A2 --> T1
+ MP --> T1
+ A3 --> T2
+ T1 --> T2
+ MP --> T2
+ A1 --> T3
+ A3 --> T3
+ MP --> T3
+ A3 --> T4
+ T2 --> T4
+ T3 --> T4
+ MP --> T4
+ A2 --> T5
+ T1 --> T5
+ MP --> T5
+ A1 --> DefOr
+ A2 --> DefOr
+ A3 --> DefOr
+ MP --> DefOr
+ DefOr --> DefAnd
+ A3 --> DefAnd
+ MP --> DefAnd
+ DefAnd --> DefIff
+ T9 --> DefIff
+ MP --> DefIff
+ DefOr --> T6
+ A1 --> T6
+ MP --> T6
+ DefAnd --> T7
+ A1 --> T7
+ A3 --> T7
+ MP --> T7
+ DefAnd --> T8
+ A1 --> T8
+ A3 --> T8
+ MP --> T8
+ A1 --> T9
+ A2 --> T9
+ MP --> T9
+ DefOr --> T10
+ T4 --> T10
+ MP --> T10
+ DefAnd --> T11
+ DefOr --> T11
+ T4 --> T11
+ T6 --> T11
+ MP --> T11
+ DefAnd --> T12
+ DefOr --> T12
+ T4 --> T12
+ MP --> T12
+ DefOr --> T13
+ T2 --> T13
+ T3 --> T13
+ MP --> T13
+ DefAnd --> T14
+ T4 --> T14
+ MP --> T14
+ DefAnd --> T15
+ A1 --> T15
+ A2 --> T15
+ MP --> T15
+ DefOr --> T16
+ T4 --> T16
+ MP --> T16
+ DefAnd --> T17
+ T9 --> T17
+ MP --> T17
+ DefAnd --> T18
+ DefOr --> T18
+ T6 --> T18
+ T7 --> T18
+ T8 --> T18
+ T9 --> T18
+ MP --> T18
+ A1 --> T19
+ A2 --> T19
+ MP --> T19
+ classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b
+ classDef definition fill:#3498db,color:#fff,stroke:#2980b9
+ classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085
+ class A1,A2,A3,MP axiom
+ class DefOr,DefAnd,DefIff definition
+ class T1,T2,T3,T4,T5,T6,T7,T8,T9,T10,T11,T12,T13,T14,T15,T16,T17,T18,T19 theorem
\ No newline at end of file
diff --git a/dependency-graphs-design.html b/dependency-graphs-design.html
new file mode 100644
index 0000000000000000000000000000000000000000..313e65b009e3c2edfcc196aa6a4c5a0b7606e8e0
--- /dev/null
+++ b/dependency-graphs-design.html
@@ -0,0 +1,80 @@
+
+
+
+
+
+ Mathematical Dependency Graphs — Design Document
+
+
+
+
+
+ Mathematical Dependency Graphs — Design Document
+
+ Overview
+ A hybrid architecture for representing and visualizing axiomatic dependency structures across multiple mathematical subjects. Supports both static Mermaid subgraphs and interactive full-graph exploration.
+
+ Scope: Target Subjects
+
+ | Subject | Foundations | Derived Items | Notes |
|---|
+ | Euclid's Elements | Postulates, Common Notions, Definitions | 464 Propositions (13 books) | Geometric constructions |
+ | Peano Arithmetic | 5 axioms, definitions | Theorems | Successor, induction |
+ | Other number systems | Axioms (integers, rationals, reals) | Theorems | Construction sequences |
+ | Number theory | Definitions, lemmas | Theorems | Divisibility, primes |
+ | Algebra | Group/ring/field axioms | Theorems | Abstract structures |
+ | Hilbert's geometry | 5 groups of axioms | Theorems | Grundlagen der Geometrie |
+ | Tarski's geometry | Betweenness, congruence relations | Theorems | First-order, decidable |
+ | Analysis | Completeness, continuity axioms | Theorems | Real analysis, limits |
+
+
+ Metadata & Sources
+ Each discourse includes metadata (created, lastUpdated, version, license, authors, methodology, citation) and sources (primary texts, digital editions, commentaries). Nodes can reference sources via sourceRef.
+
+ Hybrid Architecture
+
+ - Canonical JSON — One file per discourse, source of truth
+ - Mermaid generator — Filter by book/chapter, output subgraph
+ - Interactive viewer — Full graph with zoom, search, highlight
+ - Index/Registry — Catalog of all discourses with metadata
+
+
+ Implementation Phases
+
+ | Phase | Deliverable |
|---|
+ | 1 | Schema + Euclid Props 1–6 JSON; Mermaid generator script |
+ | 2 | Euclid Book I full JSON; static pages for Books I–IV |
+ | 3 | Interactive viewer (single discourse) |
+ | 4 | Peano Arithmetic, Hilbert Geometry JSON |
+ | 5 | Multi-discourse index; cross-discourse navigation |
+ | 6 | Tarski, Analysis, other subjects |
+
+
+ References
+
+ - Euclid's Elements: Perseus Digital Library
+ - Heath, T.L. The Thirteen Books of Euclid's Elements (1908, 2nd ed.)
+ - Hilbert: Grundlagen der Geometrie (1899)
+ - Tarski: What is Elementary Geometry? (1959)
+ - Peano: Arithmetices principia (1889)
+
+
+ Programming Framework · Mathematical Dependency Graphs Design · 2026
+
+
diff --git a/discourse-schema.json b/discourse-schema.json
new file mode 100644
index 0000000000000000000000000000000000000000..0e359735b6cd812383f91fd84a79cbc57d521850
--- /dev/null
+++ b/discourse-schema.json
@@ -0,0 +1,102 @@
+{
+ "$schema": "http://json-schema.org/draft-07/schema#",
+ "title": "Mathematical Discourse Dependency Graph",
+ "description": "Schema for axiomatic dependency structures across mathematical subjects",
+ "type": "object",
+ "required": ["schemaVersion", "discourse", "nodes", "edges"],
+ "properties": {
+ "schemaVersion": { "type": "string", "const": "1.0" },
+ "discourse": {
+ "type": "object",
+ "required": ["id", "name", "subject"],
+ "properties": {
+ "id": { "type": "string", "pattern": "^[a-z0-9-]+$" },
+ "name": { "type": "string" },
+ "subject": {
+ "type": "string",
+ "enum": ["geometry", "arithmetic", "algebra", "analysis", "number_theory", "foundations", "other"]
+ },
+ "variant": { "type": "string" },
+ "description": { "type": "string" },
+ "structure": { "type": "object" }
+ }
+ },
+ "metadata": {
+ "type": "object",
+ "properties": {
+ "created": { "type": "string", "format": "date" },
+ "lastUpdated": { "type": "string", "format": "date" },
+ "version": { "type": "string" },
+ "license": { "type": "string" },
+ "authors": { "type": "array", "items": { "type": "string" } },
+ "methodology": { "type": "string" },
+ "citation": { "type": "string" },
+ "keywords": { "type": "array", "items": { "type": "string" } }
+ }
+ },
+ "sources": {
+ "type": "array",
+ "items": {
+ "type": "object",
+ "required": ["id"],
+ "properties": {
+ "id": { "type": "string" },
+ "type": { "type": "string", "enum": ["primary", "secondary", "digital", "commentary"] },
+ "authors": { "type": "string" },
+ "title": { "type": "string" },
+ "year": { "type": ["string", "integer"] },
+ "edition": { "type": "string" },
+ "publisher": { "type": "string" },
+ "url": { "type": "string", "format": "uri" },
+ "doi": { "type": "string" },
+ "notes": { "type": "string" }
+ }
+ }
+ },
+ "nodes": {
+ "type": "array",
+ "items": {
+ "type": "object",
+ "required": ["id", "type", "label"],
+ "properties": {
+ "id": { "type": "string" },
+ "type": {
+ "type": "string",
+ "enum": ["axiom", "postulate", "commonNotion", "definition", "proposition", "theorem", "lemma", "corollary"]
+ },
+ "label": { "type": "string" },
+ "shortLabel": { "type": "string" },
+ "book": { "type": "integer" },
+ "chapter": { "type": "integer" },
+ "number": { "type": "integer" },
+ "colorClass": { "type": "string" },
+ "sourceRef": { "type": "string", "description": "Reference to sources[].id + location (e.g., 'euclid-heath, Book I, Prop 1')" },
+ "notes": { "type": "string" },
+ "keywords": { "type": "array", "items": { "type": "string" } },
+ "relatedNodes": { "type": "array", "items": { "type": "string" }, "description": "IDs of conceptually related nodes" }
+ }
+ }
+ },
+ "edges": {
+ "type": "array",
+ "items": {
+ "type": "object",
+ "required": ["from", "to"],
+ "properties": {
+ "from": { "type": "string" },
+ "to": { "type": "string" }
+ }
+ }
+ },
+ "colorScheme": {
+ "type": "object",
+ "additionalProperties": {
+ "type": "object",
+ "properties": {
+ "fill": { "type": "string" },
+ "stroke": { "type": "string" }
+ }
+ }
+ }
+ }
+}
diff --git a/generator/build-aristotle-syllogistic.js b/generator/build-aristotle-syllogistic.js
new file mode 100644
index 0000000000000000000000000000000000000000..ffdb5d4689ea6929a2fbe6d4e2ce4ce3b5e4c1f4
--- /dev/null
+++ b/generator/build-aristotle-syllogistic.js
@@ -0,0 +1,355 @@
+#!/usr/bin/env node
+/**
+ * Build Aristotle Syllogistic Logic discourse JSON and Mermaid.
+ * Based on Prior Analytics: 4 perfect syllogisms, 3 conversion rules, 10 imperfect syllogisms.
+ * Source: Stanford Encyclopedia of Philosophy, Aristotle's Logic.
+ */
+
+const fs = require('fs');
+const path = require('path');
+
+const NODES = [
+ { id: "DefAEIO", type: "definition", label: "A, E, I, O: All/No/Some/Some-not", short: "Categorical forms", colorClass: "definition" },
+ { id: "DefFig", type: "definition", label: "Three figures: middle term position", short: "Figures 1,2,3", colorClass: "definition" },
+ { id: "Econv", type: "axiom", label: "Eab implies Eba (No M is P implies No P is M)", short: "E-conversion", colorClass: "axiom" },
+ { id: "Iconv", type: "axiom", label: "Iab implies Iba (Some M is P implies Some P is M)", short: "I-conversion", colorClass: "axiom" },
+ { id: "Aconv", type: "axiom", label: "Aab implies Iba (All M is P implies Some P is M)", short: "A-conversion", colorClass: "axiom" },
+ { id: "Barbara", type: "axiom", label: "AAA-1: All M is P, All S is M therefore All S is P", short: "Barbara", colorClass: "axiom" },
+ { id: "Celarent", type: "axiom", label: "EAE-1: No M is P, All S is M therefore No S is P", short: "Celarent", colorClass: "axiom" },
+ { id: "Darii", type: "axiom", label: "AII-1: All M is P, Some S is M therefore Some S is P", short: "Darii", colorClass: "axiom" },
+ { id: "Ferio", type: "axiom", label: "EIO-1: No M is P, Some S is M therefore Some S is not P", short: "Ferio", colorClass: "axiom" },
+ { id: "Cesare", type: "theorem", label: "EAE-2: No P is M, All S is M therefore No S is P", short: "Cesare", colorClass: "theorem" },
+ { id: "Camestres", type: "theorem", label: "AEE-2: All P is M, No S is M therefore No S is P", short: "Camestres", colorClass: "theorem" },
+ { id: "Festino", type: "theorem", label: "EIO-2: No P is M, Some S is M therefore Some S is not P", short: "Festino", colorClass: "theorem" },
+ { id: "Baroco", type: "theorem", label: "AOO-2: All P is M, Some S is not M therefore Some S is not P", short: "Baroco", colorClass: "theorem" },
+ { id: "Darapti", type: "theorem", label: "AAI-3: All M is P, All M is S therefore Some S is P", short: "Darapti", colorClass: "theorem" },
+ { id: "Felapton", type: "theorem", label: "EAO-3: No M is P, All M is S therefore Some S is not P", short: "Felapton", colorClass: "theorem" },
+ { id: "Disamis", type: "theorem", label: "IAI-3: Some M is P, All M is S therefore Some S is P", short: "Disamis", colorClass: "theorem" },
+ { id: "Datisi", type: "theorem", label: "AII-3: All M is P, Some M is S therefore Some S is P", short: "Datisi", colorClass: "theorem" },
+ { id: "Bocardo", type: "theorem", label: "OAO-3: Some M is not P, All M is S therefore Some S is not P", short: "Bocardo", colorClass: "theorem" },
+ { id: "Ferison", type: "theorem", label: "EIO-3: No M is P, Some M is S therefore Some S is not P", short: "Ferison", colorClass: "theorem" }
+];
+
+// Dependencies (from → to). Based on Prior Analytics reduction proofs.
+const DEPS = {
+ Barbara: ["DefAEIO", "DefFig"],
+ Celarent: ["DefAEIO", "DefFig"],
+ Darii: ["DefAEIO", "DefFig"],
+ Ferio: ["DefAEIO", "DefFig"],
+ Cesare: ["Econv", "Celarent"],
+ Camestres: ["Econv", "Celarent"],
+ Festino: ["Econv", "Ferio"],
+ Baroco: ["Barbara"],
+ Darapti: ["Aconv", "Darii"],
+ Felapton: ["Aconv", "Ferio"],
+ Disamis: ["Iconv", "Darii"],
+ Datisi: ["Aconv", "Darii"],
+ Bocardo: ["Barbara"],
+ Ferison: ["Aconv", "Ferio"]
+};
+
+const discourse = {
+ schemaVersion: "1.0",
+ discourse: {
+ id: "aristotle-syllogistic",
+ name: "Aristotle Syllogistic Logic",
+ subject: "logic",
+ variant: "classical",
+ description: "Aristotelian categorical syllogistic. Four perfect syllogisms (Barbara, Celarent, Darii, Ferio), three conversion rules, and ten imperfect syllogisms reduced to the perfect ones. Based on Prior Analytics.",
+ structure: { axioms: 7, definitions: 2, theorems: 10 }
+ },
+ metadata: {
+ created: "2026-03-15",
+ lastUpdated: "2026-03-15",
+ version: "1.0.0",
+ license: "CC BY 4.0",
+ authors: ["Welz, G."],
+ methodology: "Programming Framework",
+ citation: "Welz, G. (2026). Aristotle Syllogistic Dependency Graph. Programming Framework.",
+ keywords: ["Aristotle", "syllogism", "categorical logic", "Prior Analytics", "Barbara", "Celarent"]
+ },
+ sources: [
+ { id: "aristotle", type: "primary", authors: "Aristotle", title: "Prior Analytics", year: "c. 350 BCE", notes: "Syllogistic theory" },
+ { id: "stanford", type: "secondary", authors: "Smith, R.", title: "Aristotle's Logic", url: "https://plato.stanford.edu/entries/aristotle-logic/", notes: "Stanford Encyclopedia" }
+ ],
+ nodes: [],
+ edges: [],
+ colorScheme: {
+ axiom: { fill: "#e74c3c", stroke: "#c0392b" },
+ definition: { fill: "#3498db", stroke: "#2980b9" },
+ theorem: { fill: "#1abc9c", stroke: "#16a085" }
+ }
+};
+
+// Add nodes
+for (const n of NODES) {
+ discourse.nodes.push({
+ id: n.id,
+ type: n.type,
+ label: n.label,
+ shortLabel: n.id,
+ short: n.short,
+ colorClass: n.colorClass
+ });
+ for (const dep of DEPS[n.id] || []) {
+ discourse.edges.push({ from: dep, to: n.id });
+ }
+}
+
+// Write JSON
+const dataDir = path.join(__dirname, "..", "data");
+const outPath = path.join(dataDir, "aristotle-syllogistic.json");
+fs.mkdirSync(dataDir, { recursive: true });
+fs.writeFileSync(outPath, JSON.stringify(discourse, null, 2), "utf8");
+console.log("Wrote", outPath);
+
+// Sanitize label for Mermaid
+function sanitizeMermaidLabel(s) {
+ return String(s)
+ .replace(/→/g, "impl")
+ .replace(/⊢/g, "|-")
+ .replace(/∨/g, "or")
+ .replace(/∧/g, "and")
+ .replace(/↔/g, "iff")
+ .replace(/\n/g, " ");
+}
+
+// Generate Mermaid
+function toMermaid(filter) {
+ const nodes = filter ? discourse.nodes.filter(filter) : discourse.nodes;
+ const nodeIds = new Set(nodes.map(n => n.id));
+ const edges = discourse.edges.filter(e => nodeIds.has(e.from) && nodeIds.has(e.to));
+ const lines = ["graph TD"];
+ for (const n of nodes) {
+ const desc = n.short || n.label;
+ const raw = (n.shortLabel || n.id) + " " + (desc.length > 30 ? desc.slice(0, 27) + "..." : desc);
+ const lbl = sanitizeMermaidLabel(raw).replace(/"/g, '\\"');
+ lines.push(` ${n.id}("${lbl}")`);
+ }
+ for (const e of edges) {
+ lines.push(` ${e.from} --> ${e.to}`);
+ }
+ lines.push(" classDef axiom fill:#e74c3c,color:#fff,stroke:#c0392b");
+ lines.push(" classDef definition fill:#3498db,color:#fff,stroke:#2980b9");
+ lines.push(" classDef theorem fill:#1abc9c,color:#fff,stroke:#16a085");
+ const axiomIds = nodes.filter(n => n.type === "axiom").map(n => n.id).join(",");
+ const defIds = nodes.filter(n => n.type === "definition").map(n => n.id).join(",");
+ const thmIds = nodes.filter(n => n.type === "theorem").map(n => n.id).join(",");
+ if (axiomIds) lines.push(` class ${axiomIds} axiom`);
+ if (defIds) lines.push(` class ${defIds} definition`);
+ if (thmIds) lines.push(` class ${thmIds} theorem`);
+ return lines.join("\n");
+}
+
+function closure(ids) {
+ const needed = new Set(ids);
+ let changed = true;
+ while (changed) {
+ changed = false;
+ for (const e of discourse.edges) {
+ if (needed.has(e.to) && !needed.has(e.from)) { needed.add(e.from); changed = true; }
+ }
+ }
+ return n => needed.has(n.id);
+}
+
+function toMermaidWithCounts(filter) {
+ const nodes = filter ? discourse.nodes.filter(filter) : discourse.nodes;
+ const nodeIds = new Set(nodes.map(n => n.id));
+ const edges = discourse.edges.filter(e => nodeIds.has(e.from) && nodeIds.has(e.to));
+ return { mermaid: toMermaid(filter), nodes: nodes.length, edges: edges.length };
+}
+
+// 3 sections
+const sections = [
+ { name: "foundations-perfect", ids: ["DefAEIO", "DefFig", "Econv", "Iconv", "Aconv", "Barbara", "Celarent", "Darii", "Ferio"], title: "Foundations and Perfect Syllogisms", desc: "Categorical forms, figures, conversion rules, and the four perfect syllogisms (Barbara, Celarent, Darii, Ferio)" },
+ { name: "figure-2", ids: ["Cesare", "Camestres", "Festino", "Baroco"], title: "Figure 2 Syllogisms", desc: "Cesare, Camestres, Festino, Baroco reduced to perfect syllogisms via conversion" },
+ { name: "figure-3", ids: ["Darapti", "Felapton", "Disamis", "Datisi", "Bocardo", "Ferison"], title: "Figure 3 Syllogisms", desc: "Darapti, Felapton, Disamis, Datisi, Bocardo, Ferison reduced to perfect syllogisms" }
+];
+
+const subgraphData = [];
+for (const s of sections) {
+ const filter = closure(s.ids);
+ const { mermaid: sub, nodes: n, edges: e } = toMermaidWithCounts(filter);
+ subgraphData.push({ ...s, mermaid: sub, nodes: n, edges: e });
+ fs.writeFileSync(path.join(dataDir, `aristotle-syllogistic-${s.name}.mmd`), sub, "utf8");
+ console.log("Wrote", path.join(dataDir, `aristotle-syllogistic-${s.name}.mmd`));
+}
+
+fs.writeFileSync(path.join(dataDir, "aristotle-syllogistic.mmd"), toMermaid(), "utf8");
+
+// Generate HTML
+const MATH_DB = process.env.MATH_DB || "/home/gdubs/copernicus-web-public/huggingface-space/mathematics-processes-database";
+const DISC_DIR = path.join(MATH_DB, "processes", "discrete_mathematics");
+
+function htmlTemplate(title, subtitle, mermaid, nodes, edges) {
+ const mermaidEscaped = mermaid.replace(//g, ">");
+ return `
+
+
+
+
+ ${title} - Mathematics Process
+
+
+
+
+
+
+
+
+
+
+
Description
+
${subtitle}
+
Source: Aristotle, Prior Analytics (c. 350 BCE); Stanford Encyclopedia of Philosophy
+
+
+
Dependency Flowchart
+
Note: Arrows mean "depends on" (tail to head).
+
${mermaidEscaped}
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Aristotle
- syllogism
- Barbara
- Celarent
- Prior Analytics
- categorical logic
+
+
+
+
+
+
+
+
Aristotle Syllogistic Logic
+
Categorical syllogistic from Prior Analytics. Four perfect syllogisms (Barbara, Celarent, Darii, Ferio), three conversion rules, and ten imperfect syllogisms reduced to the perfect ones. Split into three views.
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
Note: Arrows mean "depends on" (tail to head).
+
${mermaidEscaped}
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - combinatorics
- permutations
- combinations
- binomial theorem
- counting
+
+
+
+
+
+
+
+
Combinatorics
+
Counting principles: sum and product rules, permutations, combinations, binomial theorem, pigeonhole principle, inclusion-exclusion, derangements. Split into three views.
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
Note: Arrows mean "depends on" (tail → head). Edge crossing can create illusions of connections between adjacent nodes—e.g., I.47 and I.40 have no direct dependency; both depend on I.31 among others.
+
${mermaidEscaped}
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book I
- axioms
- postulates
- propositions
- geometry
- Pythagorean theorem
+
+
+
+
+
+
+
+
Euclid's Elements, Book I
+
Full dependency graph of all 48 propositions. Dependencies from David E. Joyce, Clark University. Split into five views for clarity.
+
+
Propositions 1–10 — Foundations, SAS, SSS, bisections, perpendiculars
+
Propositions 11–20 — Right angles, straight lines, vertical angles, triangle exterior, triangle inequality
+
Propositions 21–30 — Lines within triangle, construct triangle/angle, parallel lines
+
Propositions 31–41 — Parallelograms, triangles, areas
+
Propositions 42–48 — Constructions, Pythagorean theorem
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
${mermaidEscaped}
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book II
- geometric algebra
- rectangles
- squares
- golden section
- quadrature
+
+
+
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
${mermaidEscaped}
+
+
+
Color Scheme
+
+
Gray
Book I, Prop II.5 (foundation)
+
+
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book III
- circles
- chords
- tangents
+
+
+
+
+
+
+
+
Euclid's Elements Book III
+
Theory of circles: 11 definitions, 37 propositions. Chords, tangents, angles at center and circumference. All depend on Book I. Prop III.35 uses Prop II.5. Dependency charts split into three views (~16 nodes each).
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
${mermaidEscaped}
+
+
+
Color Scheme
+
+
Gray
Book I, Book III, Prop II.11 (foundation)
+
+
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book IV
- inscribed
- circumscribed
- pentagon
- hexagon
+
+
+
+
+
+
+
+
Euclid's Elements Book IV
+
Inscribed and circumscribed figures: triangle, square, pentagon, hexagon, 15-gon. All depend on Books I and III. Prop IV.10 uses Prop II.11 (golden section). Dependency charts split into two views.
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
${mermaidEscaped}
+
+
+
Color Scheme
+
+
Gray
Book VII, Book VIII (foundation)
+
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book IX
- prime
- perfect
- odd
- even
+
+
+
+
+
+
+
+
Euclid's Elements Book IX
+
Primes, perfect numbers, odd/even. 36 propositions. Depends on Books VII and VIII. IX.20: infinitely many primes. IX.36: perfect numbers.
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
${mermaidEscaped}
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book V
- proportion
- ratio
- Eudoxus
+
+
+
+
+
+
+
+
Euclid's Elements Book V
+
Theory of ratio and proportion (Eudoxus). 18 definitions, 25 propositions. Does not depend on previous books. Foundation for Book VI and Books X–XIII.
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
${mermaidEscaped}
+
+
+
Color Scheme
+
+
Gray
Book I, Book V (foundation)
+
+
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book VI
- similar
- proportion
- triangles
+
+
+
+
+
+
+
+
Euclid's Elements Book VI
+
Similar figures. 4 definitions, 33 propositions. Depends on Book I and Book V. VI.1 (triangles under same height) is basis for most. VI.33 uses proportion def directly.
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
${mermaidEscaped}
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book VII
- number theory
- GCD
- prime
- LCM
+
+
+
+
+
+
+
+
Euclid's Elements Book VII
+
Number theory: GCD (Euclidean algorithm), proportions of numbers, primes, LCM. 22 definitions, 39 propositions. Does not depend on previous books.
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
${mermaidEscaped}
+
+
+
Color Scheme
+
+
Gray
Book VII (foundation)
+
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book VIII
- continued proportion
- plane
- solid
+
+
+
+
+
+
+
+
Euclid's Elements Book VIII
+
Continued proportions. 27 propositions. Depends on Book VII. Plane numbers, solid numbers, mean proportionals.
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
${mermaidEscaped}
+
+
+
Color Scheme
+
+
Gray
Book I, V, VI (foundation)
+
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book X
- incommensurable
- binomial
- apotome
- medial
+
+
+
+
+
+
+
+
Euclid's Elements Book X
+
Incommensurables, binomials, apotomes. 115 propositions, 16 definitions. Depends on Books I, V, VI. Thirteen irrational straight lines; X.115: infinite irrationals from medial.
+
+
Propositions 1–15 Commensurable, incommensurable, GCM, ratio
+
Propositions 16–30 Medial, rational, parallelogram
+
Propositions 31–45 Binomial, bimedial, major
+
Propositions 46–60 Find binomials, area by rational
+
Propositions 61–75 Square on binomial, apotome
+
Propositions 76–90 Apotome types, find apotomes
+
Propositions 91–105 Area by apotome, commensurable
+
Propositions 106–115 Rational from irrational, order
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
${mermaidEscaped}
+
+
+
Color Scheme
+
+
Gray
Book I, VI (foundation)
+
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book XI
- solid geometry
- plane
- parallelepiped
- prism
+
+
+
+
+
+
+
+
Euclid's Elements Book XI
+
Solid geometry: planes, perpendiculars, parallelepipeds, prisms. 39 propositions, 28 definitions. Depends on Books I and VI.
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
${mermaidEscaped}
+
+
+
Color Scheme
+
+
Gray
Book I, V, VI, XI (foundation)
+
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book XII
- measurement
- pyramid
- cone
- cylinder
- sphere
+
+
+
+
+
+
+
+
Euclid's Elements Book XII
+
Measurement of figures: circles (XII.2), pyramids (XII.5–7), cones and cylinders (XII.10–15), spheres (XII.18). 18 propositions. Depends on Books I, V, VI, XI.
+
+
+
+
+
+
+
+
+
Dependency Flowchart
+
${mermaidEscaped}
+
+
+
Color Scheme
+
+
Gray
Book I, IV, VI, X, XI (foundation)
+
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Euclid
- Elements
- Book XIII
- regular solids
- Platonic
- tetrahedron
- octahedron
- cube
- icosahedron
- dodecahedron
+
+
+
+
+
+
+
+
Euclid's Elements Book XIII
+
Regular solids: tetrahedron, octahedron, cube, icosahedron, dodecahedron. 18 propositions. Depends on Books I, IV, VI, X, XI. XIII.18: no other such figure exists.
+
+
+
+
+
+
+
+
Description
+
${subtitle}
+
Source: Landau, E. Foundations of Analysis (1930); Peano, G. Arithmetices principia (1889)
+
+
+
Dependency Flowchart
+
Note: Arrows mean "depends on" (tail → head).
+
${mermaidEscaped}
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - Peano
- arithmetic
- natural numbers
- induction
- successor
- foundations
+
+
+
+
+
+
+
+
Peano Arithmetic
+
Axiomatic development of natural number arithmetic. Five axioms, definitions of addition and multiplication, and key theorems. Based on Landau, Foundations of Analysis. Split into three views.
+
+
+
+
+
+
+
+
Description
+
${subtitle}
+
Source: Frege, G. Begriffsschrift (1879); Łukasiewicz, J. Elements of Mathematical Logic (1929)
+
+
+
Dependency Flowchart
+
Note: Arrows mean "depends on" (tail → head).
+
${mermaidEscaped}
+
+
+
+
+
Statistics
+
+ - Nodes: ${nodes}
+ - Edges: ${edges}
+
+
+
+
Keywords
+
+ - propositional logic
- Hilbert
- Łukasiewicz
- tautology
- modus ponens
- De Morgan
+
+
+
+
+
+
+
+
Propositional Logic
+
Hilbert-style axiomatic development of classical propositional logic. Three axioms (Łukasiewicz P2), modus ponens, definitions of disjunction, conjunction, biconditional, and key theorems. Split into three views.
+
+
-
-
🛠️
-
The Programming Framework
-
A Universal Method for Process Analysis
-
- Combining Large Language Models with Mermaid visualization to dissect and understand
- complex processes across any discipline—from biology to business, physics to psychology.
-
-
-
-
📋 Summary
-
- The Programming Framework is a universal meta-tool for analyzing complex processes across any discipline by combining Large Language Models (LLMs) with visual flowchart representation. The Framework transforms textual process descriptions into structured, interactive Mermaid flowcharts stored as JSON, enabling systematic analysis, visualization, and integration with knowledge systems.
-
-
- Successfully demonstrated through GLMP (Genome Logic Modeling Project) with 50+ biological processes, and applied across Chemistry, Mathematics, Physics, and Computer Science. The Framework serves as the foundational methodology for the CopernicusAI Knowledge Engine, enabling domain-specific process visualization and analysis.
-
-
-
📚 Prior Work & Research Contributions
-
-
-
Overview
-
- The Programming Framework represents prior work that demonstrates a novel methodology for analyzing complex processes by combining Large Language Models (LLMs) with visual flowchart representation. This research establishes a universal, domain-agnostic approach to process analysis that transforms textual descriptions into structured, interactive visualizations.
-
-
+ .header {
+ text-align: center;
+ color: white;
+ margin-bottom: 40px;
+ }
-
-
-
🔬 Research Contributions
-
- - • Universal Process Analysis: Domain-agnostic methodology applicable across multiple fields
- - • LLM-Powered Extraction: Automated extraction using Google Gemini 2.0 Flash
- - • Structured Visualization: Mermaid.js-based flowchart generation encoded as JSON
- - • Iterative Refinement: Systematic approach enabling continuous improvement
- - • Scale Demonstration: Applied to 313+ processes across 5 disciplines (Biology: 52, Chemistry: 91, Physics: 21, Computer Science: 21, Mathematics: 20, GLMP: 109)
- - • Validation: Successfully processes complex biological, chemical, and computational workflows with high accuracy
-
-
-
-
-
⚙️ Technical Achievements
-
- - • Meta-Tool Architecture: Framework for creating specialized analysis tools
- - • JSON-Based Storage: Structured format enabling version control and API integration
- - • Multi-Domain Application: Successfully applied to biological processes (GLMP)
- - • Integration Framework: Designed for knowledge engines and collaborative platforms
-
-
-
+ .header h1 {
+ font-size: 3rem;
+ margin-bottom: 10px;
+ text-shadow: 2px 2px 4px rgba(0,0,0,0.3);
+ }
-
-
🎯 Position Within CopernicusAI Knowledge Engine
-
- The Programming Framework serves as the foundational meta-tool of the CopernicusAI Knowledge Engine, providing the underlying methodology that enables specialized applications:
-
-
-
- - • GLMP (Genome Logic Modeling Project)
- - • CopernicusAI (main knowledge engine)
- - • Research Papers Metadata Database
-
-
- - • Science Video Database
- - • Multi-domain process analysis
-
-
-
- This work establishes a proof-of-concept for AI-assisted process analysis, demonstrating how LLMs can systematically extract and visualize complex logic from textual sources across diverse domains.
-
-
-
-
-
-
-
-
JSON
-
Structured Data
-
-
-
🎯 What is the Programming Framework?
-
-
- The Programming Framework is a meta-tool—a tool for creating tools. It provides a
- systematic method for analyzing any complex process by combining the analytical power of Large Language
- Models with the clarity of visual flowcharts.
-
-
-
-
-
🔍 The Problem
-
- Complex processes—whether biological, computational, or organizational—are difficult to
- understand because they involve many steps, decision points, and interactions. Traditional
- descriptions in text are hard to follow.
-
-
+ .header p {
+ font-size: 1.2rem;
+ opacity: 0.9;
+ }
-
-
✨ The Solution
-
- Use LLMs to extract process logic from literature, then encode it as Mermaid flowcharts
- stored in JSON. Result: Clear, interactive visualizations that reveal hidden patterns and
- enable systematic analysis.
-
-
-
-
-
-
⚙️ How It Works
-
-
-
-
1️⃣
-
Input Process
-
Provide scientific papers, documentation, or process descriptions
-
+ .overview {
+ margin-bottom: 40px;
+ }
-
-
2️⃣
-
LLM Analysis
-
AI extracts steps, decisions, branches, and logic flow
-
+ .overview h2 {
+ color: #667eea;
+ font-size: 2rem;
+ margin-bottom: 20px;
+ border-bottom: 3px solid #667eea;
+ padding-bottom: 10px;
+ }
-
-
3️⃣
-
Generate Flowchart
-
Create Mermaid diagram encoded as JSON structure
-
+ .overview p {
+ font-size: 1.1rem;
+ margin-bottom: 15px;
+ text-align: justify;
+ }
-
-
4️⃣
-
Visualize & Iterate
-
Interactive flowchart reveals insights and enables refinement
-
-
+ .color-system {
+ margin-bottom: 40px;
+ }
-
-
📝 Concrete Example:
-
-
Input:
-
- "DNA replication begins when the origin recognition complex (ORC) binds to DNA replication origins. This triggers the loading of the MCM2-7 helicase complex, which unwinds the DNA double helix. DNA polymerases then synthesize new strands using the unwound strands as templates..."
-
-
LLM Analysis:
-
- Extracts 15 steps, identifies 3 decision points (origin recognition, helicase loading, polymerase binding), recognizes 4 key enzymes (ORC, MCM2-7, DNA polymerase, ligase), and maps regulatory checkpoints.
-
-
Output:
-
- Mermaid flowchart with 25 nodes, 28 edges, 3 decision gates, properly colored using the 5-color scheme (red for inputs, yellow for structures, green for operations, blue for intermediates, violet for products), stored as structured JSON enabling interactive visualization and programmatic access.
-
-
-
+ .color-system h2 {
+ color: #667eea;
+ font-size: 2rem;
+ margin-bottom: 20px;
+ border-bottom: 3px solid #667eea;
+ padding-bottom: 10px;
+ }
-
-
📊 Live Interactive Example:
-
- graph TD
- A[Complex Process Input] --> B{LLM Analysis}
- B -->|Extract Logic| C[Identify Steps]
- B -->|Extract Decisions| D[Identify Branches]
- C --> E[Create Flowchart Nodes]
- D --> F[Create Decision Points]
- E --> G[Generate Mermaid Syntax]
- F --> G
- G --> H[Store as JSON]
- H --> I[Interactive Visualization]
- I --> J{Insights Gained?}
- J -->|No| K[Refine Analysis]
- J -->|Yes| L[Apply Knowledge]
- K --> B
-
- style A fill:#ff6b6b,color:#fff
- style B fill:#74c0fc,color:#fff
- style C fill:#51cf66,color:#fff
- style D fill:#51cf66,color:#fff
- style E fill:#ffd43b,color:#000
- style F fill:#ffd43b,color:#000
- style G fill:#51cf66,color:#fff
- style H fill:#74c0fc,color:#fff
- style I fill:#74c0fc,color:#fff
- style J fill:#74c0fc,color:#fff
- style K fill:#51cf66,color:#fff
- style L fill:#b197fc,color:#fff
-
-
-
Color Legend:
-
- Red - Triggers & Inputs
- Yellow - Structures & Objects
- Green - Processing & Operations
- Blue - Intermediates & States
- Violet - Products & Outputs
-
-
-
-
💡 Core Principles
-
-
-
-
🌍
-
Domain Agnostic
-
- Works across any field: biology, chemistry, software engineering, business processes,
- legal workflows, manufacturing, and beyond.
-
-
+ .color-grid {
+ display: grid;
+ grid-template-columns: repeat(auto-fit, minmax(300px, 1fr));
+ gap: 20px;
+ margin-bottom: 30px;
+ }
-
-
🔄
-
Iterative Refinement
-
- Start with rough analysis, visualize, identify gaps, refine with LLM, repeat until
- the process logic is crystal clear.
-
-
+ .color-card {
+ border-radius: 10px;
+ padding: 20px;
+ color: white;
+ text-align: center;
+ box-shadow: 0 5px 15px rgba(0,0,0,0.2);
+ transition: transform 0.3s ease;
+ }
-
-
📦
-
Structured Data
-
- JSON storage enables programmatic access, version control, cross-referencing,
- and integration with other tools and databases.
-
-
-
-
📚 Process Diagram Collections
-
- The Programming Framework has been applied across multiple scientific disciplines. Explore interactive flowchart collections organized by domain:
-
-
-
-
-
-
- 🧬 Biology
-
-
- Biological process visualizations: GLMP covers biochemical/molecular processes; Biology Database covers higher-level organismal processes.
-
-
-
- Biology Database: 52 processes (organismal/ecological) | GLMP: 50+ processes (biochemical/molecular)
-
-
+ .color-card p {
+ font-size: 1rem;
+ opacity: 0.9;
+ }
-
-
-
- ⚗️ Chemistry
-
-
- Comprehensive chemistry process diagrams across all major branches.
-
-
- 🗄️ Chemistry Database Table →
-
-
- 56 processes across 14 subcategories
-
-
+ .red { background: linear-gradient(135deg, #ff6b6b, #ee5a52); }
+ .yellow { background: linear-gradient(135deg, #ffd43b, #fcc419); color: #333; }
+ .green { background: linear-gradient(135deg, #51cf66, #40c057); }
+ .blue { background: linear-gradient(135deg, #74c0fc, #4dabf7); }
+ .violet { background: linear-gradient(135deg, #b197fc, #9775fa); }
-
-
-
- 🔢 Mathematics
-
-
- Mathematical algorithms, proof methods, and computational processes.
-
-
- 🗄️ Mathematics Database Table →
-
-
- 20 processes across 7 subcategories
-
-
+ .resources {
+ margin-bottom: 40px;
+ }
-
-
-
- ⚛️ Physics
-
-
- Physical processes including quantum mechanics, thermodynamics, and particle physics.
-
-
- 🗄️ Physics Database Table →
-
-
- 21 processes across 7 subcategories
-
-
+ .resources h2 {
+ color: #667eea;
+ font-size: 2rem;
+ margin-bottom: 20px;
+ border-bottom: 3px solid #667eea;
+ padding-bottom: 10px;
+ }
-
-
-
-
-
⚙️ Technical Architecture
+ .footer {
+ background: rgba(255,255,255,0.1);
+ color: white;
+ text-align: center;
+ padding: 30px;
+ border-radius: 15px;
+ backdrop-filter: blur(10px);
+ }
+
+ .footer h3 {
+ margin-bottom: 15px;
+ font-size: 1.5rem;
+ }
+
+ .footer p {
+ margin-bottom: 5px;
+ opacity: 0.9;
+ }
+
+ .footer a {
+ color: #ffd43b;
+ text-decoration: none;
+ }
+
+ .footer a:hover {
+ text-decoration: underline;
+ }
+
+ @media (max-width: 768px) {
+ .header h1 {
+ font-size: 2rem;
+ }
-
-
-
🤖 LLM Integration
-
- - • Google Gemini 2.0 Flash for analysis
- - • Vertex AI for enterprise deployment
- - • Custom prompts for process extraction
- - • Structured JSON output formatting
-
-
+ .main-content {
+ padding: 20px;
+ }
+
+ .color-grid {
+ grid-template-columns: 1fr;
+ }
+
+ .resource-grid {
+ grid-template-columns: 1fr;
+ }
-
-
📊 Visualization Stack
-
- - • Mermaid.js for flowchart rendering
- - • JSON schema for data validation
- - • Interactive SVG output
- - • Export to PNG/PDF supported
-
-
+ .db-grid {
+ grid-template-columns: 1fr;
+ }
+ }
+
+
+
+
+
-
-
💾 Data Storage
-
- - • Google Cloud Storage for JSON files
- - • Firestore for metadata indexing
- - • Version control with Git
- - • Cross-referencing with papers database
-
-
+
+
+
Project Overview
+
The Programming Framework is a systematic visualization methodology for analyzing complex systems across disciplines using Mermaid Markdown and a universal five-color code.
+
+
Complex systems across biology, chemistry, and physics exhibit remarkable similarities in their organizational principles despite operating at vastly different scales and domains. Traditional analysis methods often remain siloed within specific disciplines, limiting our ability to identify common patterns and computational logic that govern system behavior.
+
+
Here, we present the Programming Framework, a systematic methodology that translates complex system dynamics into standardized computational representations using Mermaid Markdown syntax and LLM processing.
+
-
-
🔗 Integration Points
-
- - • GLMP specialized collections
- - • CopernicusAI knowledge graph
- - • Research papers database
- - • API endpoints for programmatic access
-
-
+
+
Technical Foundation: Mermaid Markdown
+
+
The Invention of Mermaid
+
Knut Sveidqvist invented the Mermaid markdown format. He created Mermaid, a JavaScript-based diagramming and charting tool, to simplify diagram creation in documentation workflows. The project was inspired by his experience trying to update a diagram in a document, which was difficult due to the file format.
+
+
Sveidqvist's innovation revolutionized how diagrams are created and maintained in documentation by providing a text-based syntax that can be version-controlled, easily edited, and automatically rendered into visual diagrams. This approach eliminates the need for external diagramming tools and ensures diagrams stay synchronized with their documentation.
+
+
Mermaid Markdown (.mmd) Format
+
The Programming Framework leverages Mermaid's .mmd file format, which provides:
+
+ - Text-based syntax for creating complex flowcharts and diagrams
+ - Version control compatibility - diagrams can be tracked in Git repositories
+ - LLM-friendly format - AI systems can generate and modify diagram code
+ - Cross-platform compatibility - works in any environment that supports JavaScript
+ - Embeddable rendering - diagrams can be displayed in HTML, Markdown, and other formats
+
+
+
LLM Integration and Workflow
+
Our methodology uses Large Language Models to:
+
+ - Generate .mmd files - LLMs create detailed Mermaid syntax for complex processes
+ - Apply color coding - Systematic application of the 5-category color system
+ - Ensure consistency - Standardized node naming and connection patterns
+ - Embed in HTML - .mmd files are embedded in HTML for web display
+ - Maintain quality - LLMs can validate and optimize diagram structure
+
+
+
This workflow enables rapid creation of sophisticated visualizations that would be impractical to create manually, while maintaining the flexibility and editability of text-based formats.
-
-
-
-
-
-
✅ Validation & Accuracy
-
-
-
-
🔍 Quality Assurance Process
-
- - • Automated Validation: All flowcharts validated for Mermaid syntax correctness before publication
- - • Metadata Quality Checks: JSON schema validation ensures >=85% metadata completeness (NSF standard)
- - • Source Citation Verification: All processes include verified research paper citations with DOI/PubMed links
- - • Cross-Reference Validation: Automated checks ensure discipline links and back-references are correct
- - • Color Scheme Consistency: All processes follow standardized 5-color scheme for visual consistency
-
-
+
+
Universal Color Coding System
+
The framework uses a standardized five-color system across all disciplines:
-
-
📊 Scale & Coverage
-
- - • 314 Processes Validated: Successfully applied across 6 discipline databases (Biology, Chemistry, Physics, CS, Mathematics, GLMP)
- - • Multi-Domain Testing: Framework validated on biological pathways, chemical reactions, computational algorithms, and mathematical proofs
- - • Iterative Refinement: Processes refined through multiple LLM analysis cycles to improve accuracy
- - • User Feedback Integration: Community feedback mechanism enables continuous improvement (see "Improve this process" on each flowchart)
- - • Expert Validation: GLMP processes validated against established biochemical pathway databases
-
+
+
+
🔴 Red (#ff6b6b)
+
Triggers & Inputs
+ Environmental signals, reactants, input data, energy sources, axioms
+
+
+
+
🟡 Yellow (#ffd43b)
+
Structures & Objects
+ Enzymes, catalysts, data structures, fields, theorems
+
+
+
+
🟢 Green (#51cf66)
+
Processing & Operations
+ Metabolic reactions, chemical reactions, algorithms, quantum operations
+
+
+
+
🔵 Blue (#74c0fc)
+
Intermediates & States
+ Metabolites, intermediates, variables, states, measurement results
+
+
+
+
🟣 Violet (#b197fc)
+
Products & Outputs
+ Biomolecules, final products, program outputs, phenomena, proven results
+
-
-
🎯 Accuracy Measures
-
-
-
Syntax Accuracy
-
100% of published flowcharts render without Mermaid syntax errors
+
+
📊 Batch Architecture Status
+
+
+
📐 Mathematics
+
Status: Complete ✅
+
Batches: 7/7 (21 processes)
+
Number Theory, Analysis & Calculus, Abstract Algebra, Geometry & Topology, Applied Mathematics, Discrete Mathematics, Historical & Educational
+
+
+
+
⚗️ Chemistry
+
Status: Completed ✅
+
Batches: 14/14 (70 processes)
+
Organic Chemistry, Physical Chemistry, Analytical Chemistry, Inorganic Chemistry, Biochemistry, Materials Chemistry, Environmental Chemistry, Electrochemical Processes, Surface Chemistry & Catalysis, Thermodynamic Processes, Kinetic Processes, Spectroscopy & Analysis, Quantum Chemistry, Materials Chemistry
-
-
Metadata Completeness
-
>=85% average quality score across all processes (exceeds NSF requirements)
+
+
+
💻 Computer Science
+
Status: In Progress 🔄
+
Batches: 1/7 (3 processes)
+
Algorithms & Data Structures, Software Engineering, Artificial Intelligence, Computer Security, Computer Networks, Database Systems, Computer Graphics
-
-
Source Coverage
-
All processes include 1-3 verified research paper citations with accessible links
+
+
+
⚛️ Physics
+
Status: In Progress 🔄
+
Batches: 1/7 (3 processes)
+
Classical Mechanics, Electromagnetism, Thermodynamics, Quantum Mechanics, Relativity, Astrophysics, Particle Physics
+
+
+
+
🧬 Biology
+
Status: External GLMP ✅
+
Location: Hugging Face Space
+
Comprehensive biological systems analysis with genome logic modeling and metabolic pathway visualization
-
-
⚠️ Known Limitations
-
- - • LLM-Dependent Accuracy: Flowchart accuracy depends on LLM interpretation of source material; complex processes may require multiple refinement cycles
- - • Domain Expertise Required: While the Framework is domain-agnostic, optimal results benefit from domain-specific knowledge for validation
- - • Source Material Quality: Accuracy is limited by the quality and completeness of input source material
- - • Continuous Improvement: Framework is actively refined based on user feedback and validation results
-
-
-
-
-
-
-
- 🔗 Related Projects
-
-
-
-
🧬 GLMP - Genome Logic Modeling
-
- First specialized application of the Programming Framework to biochemical processes.
- 100+ biological pathways visualized as interactive flowcharts.
-
-
- Explore GLMP → (opens in new tab)
-
-
+
+
🗄️ Discipline databases
+
Live searchable tables on cloud storage where deployed; other disciplines use static batch pages while the database spine is built out.
+
+
+
🧬 Biology
+
Biological process visualizations: GLMP covers biochemical and molecular processes; the Biology Database covers higher-level organismal pathways, mechanisms, and lab-style protocols.
+
+
Biology database: 20 processes (3 groups: pathways, mechanisms, assay protocols) · GLMP: 50+ processes (biochemical / molecular)
+
-
-
-
-
-
-
-
-
How to Cite This Work
-
-
-
- Welz, G. (2024–2025). The Programming Framework: A Universal Method for Process Analysis.
- Hugging Face Spaces. https://huggingface.co/spaces/garywelz/programming_framework (opens in new tab)
-
-
-
BibTeX Format:
-
@misc{welz2025programmingframework,
- title={The Programming Framework: A Universal Method for Process Analysis},
- author={Welz, Gary},
- year={2024--2025},
- url={https://huggingface.co/spaces/garywelz/programming_framework},
- note={Hugging Face Spaces}
-}
+
+
⚗️ Chemistry
+
Under development
+
Comprehensive chemistry process diagrams across major branches. A public interactive database table (like Mathematics and Biology) is not live yet; batch HTML pages in this Space are the current entry point.
+
+
+
+
+
🔢 Mathematics
+
Algorithms, proof methods, dependency graphs, and computational processes — main table plus named collections (mathematicians, theorems) and a whole-of-mathematics graph view.
+
+
216 processes across 23 subcategories (see live table for the exact index)
+
+
+
+
⚛️ Physics
+
Under development
+
Physical processes including quantum mechanics, thermodynamics, and particle physics. Interactive cloud table coming; use static physics batches for now.
+
+
+
+
+
💻 Computer Science
+
Under development
+
Algorithms, software-engineering workflows, and computational processes. Interactive cloud table coming; use static CS batches for now.
+
-
-
-
-
- This project serves as a foundational meta-tool for AI-assisted process analysis, enabling systematic extraction and visualization of complex logic from textual sources across diverse scientific and technical domains.
-
-
- The Programming Framework is designed as infrastructure for AI-assisted science, providing a universal methodology that can be specialized for domain-specific applications.
-
+
+
+
Key Resources & Documentation
+
+
+
📚 Complete Documentation
+
Detailed methodology, implementation guidelines, and theoretical foundation
+
View README
+
+
+
+
+
+
⚗️ Chemistry
+
Chemical reaction mechanisms and visualizations
+
Explore Chemistry
+
+
+
+
💻 Computer Science
+
Algorithm and computational analysis
+
Explore CS
+
+
+
+
⚛️ Physics
+
Physical system dynamics and quantum processes
+
Explore Physics
+
+
+
+
📐 Mathematics
+
Mathematical proof and calculation processes
+
Explore Math
+
+
+
+
📖 Academic Article
+
Comprehensive framework documentation and methodology
+
Read Article
+
+
+
+
🔬 GLMP Foundation
+
Original biological systems analysis project
+
Visit GLMP
+
+
+
+
🧪 Experimental Validation
+
Comprehensive validation protocols and experimental design
+
View Validation Paper
+
+
+
+
+
+
📄 Journal Submission Paper
+
Self-contained academic paper suitable for journal submission or arXiv
+
View Journal Paper
+
+
-
-
-
-
-
+
+
 -
-  -
-  -
- 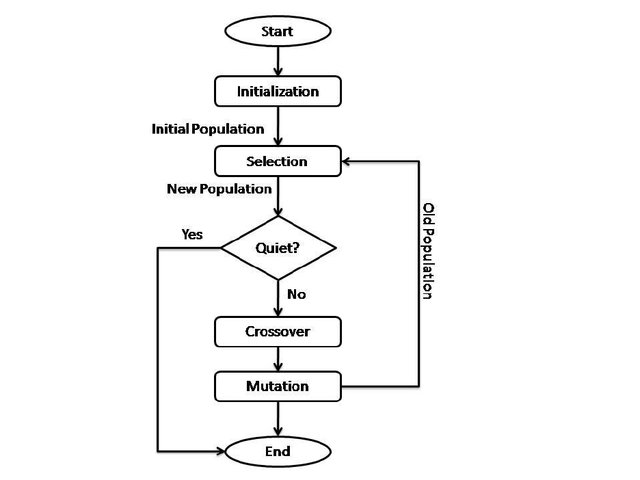 -
-  -
-  -
-  -
-  -
-  -
-